Berkin Bilgic
Harvard-MIT Health Sciences and Technology, Athinoula A. Martinos Center for Biomedical Imaging, Harvard Medical School
Accurate parameter estimation using scan-specific unsupervised deep learning for relaxometry and MR fingerprinting
Dec 13, 2021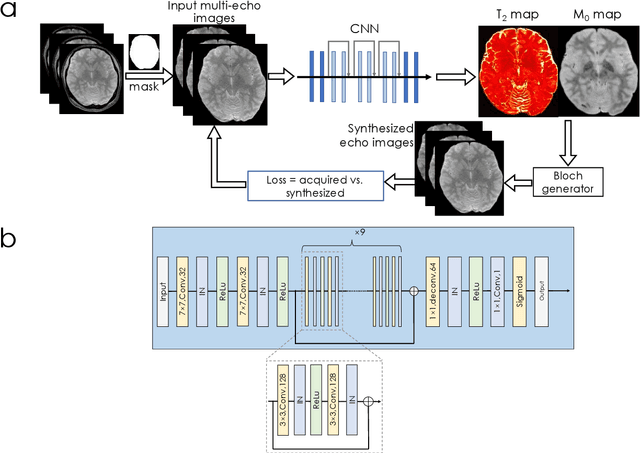
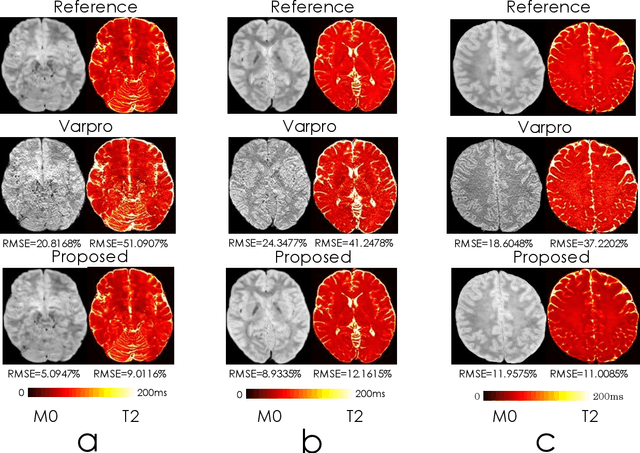
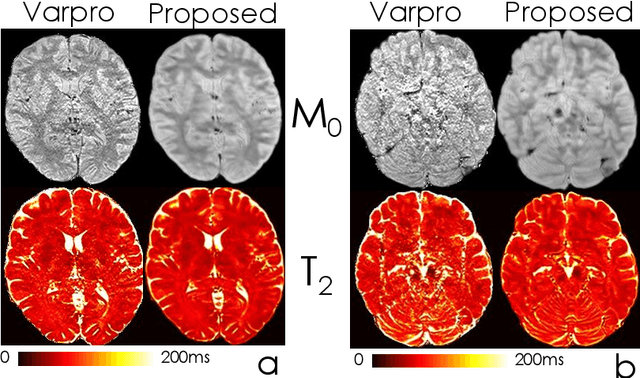
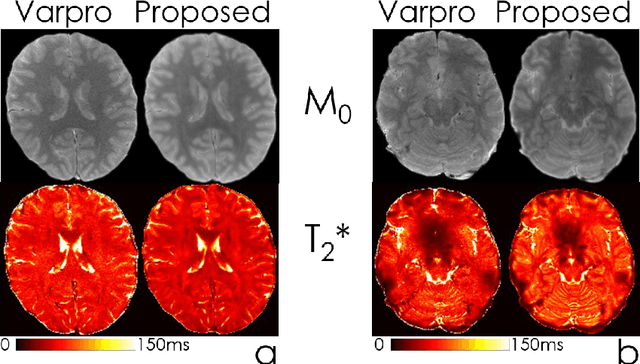
Abstract:We propose an unsupervised convolutional neural network (CNN) for relaxation parameter estimation. This network incorporates signal relaxation and Bloch simulations while taking advantage of residual learning and spatial relations across neighboring voxels. Quantification accuracy and robustness to noise is shown to be significantly improved compared to standard parameter estimation methods in numerical simulations and in vivo data for multi-echo T2 and T2* mapping. The combination of the proposed network with subspace modeling and MR fingerprinting (MRF) from highly undersampled data permits high quality T1 and T2 mapping.
Quantifying the uncertainty of neural networks using Monte Carlo dropout for deep learning based quantitative MRI
Dec 02, 2021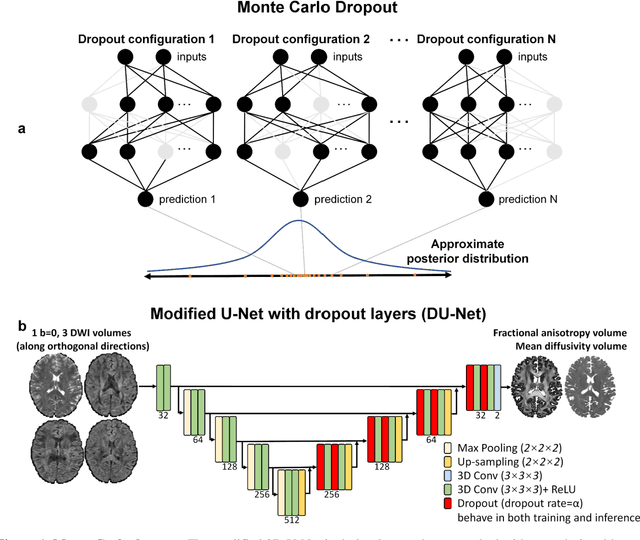
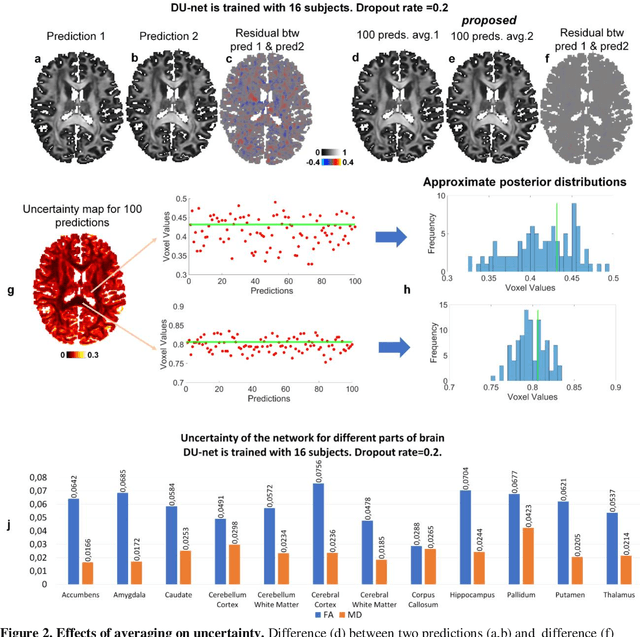

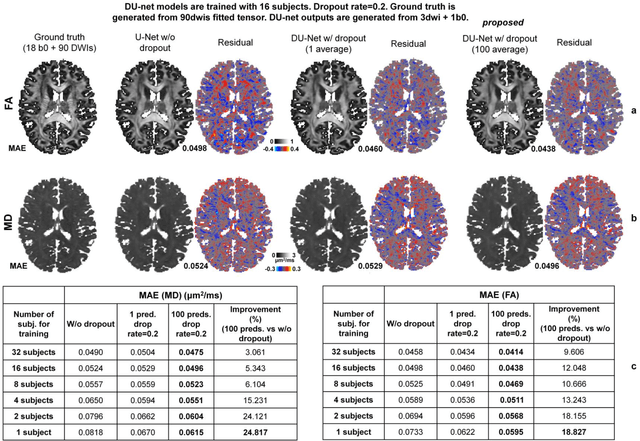
Abstract:Dropout is conventionally used during the training phase as regularization method and for quantifying uncertainty in deep learning. We propose to use dropout during training as well as inference steps, and average multiple predictions to improve the accuracy, while reducing and quantifying the uncertainty. The results are evaluated for fractional anisotropy (FA) and mean diffusivity (MD) maps which are obtained from only 3 direction scans. With our method, accuracy can be improved significantly compared to network outputs without dropout, especially when the training dataset is small. Moreover, confidence maps are generated which may aid in diagnosis of unseen pathology or artifacts.
SDnDTI: Self-supervised deep learning-based denoising for diffusion tensor MRI
Nov 14, 2021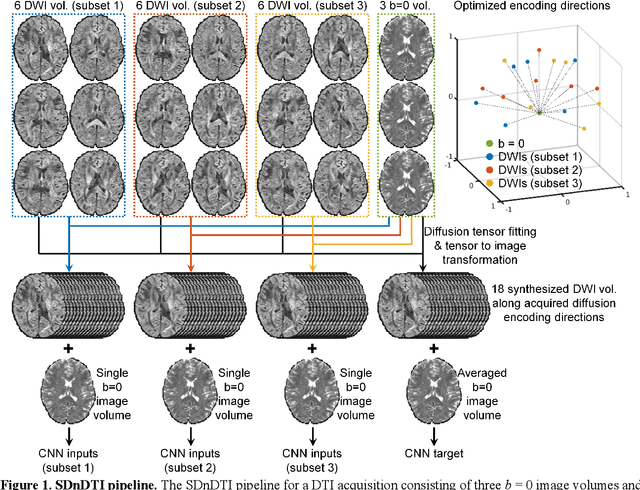
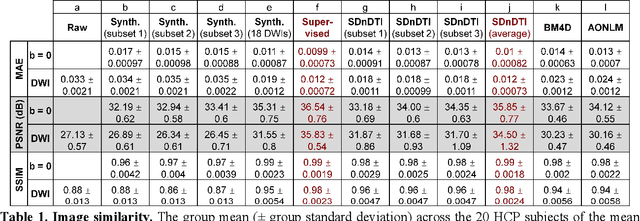
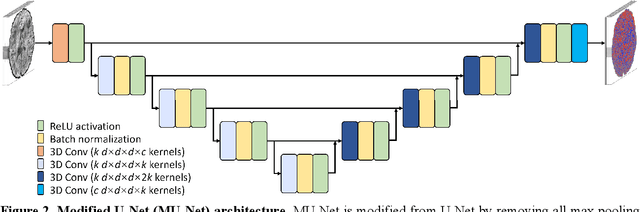
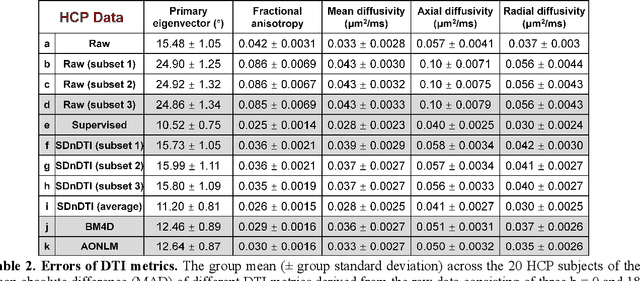
Abstract:The noise in diffusion-weighted images (DWIs) decreases the accuracy and precision of diffusion tensor magnetic resonance imaging (DTI) derived microstructural parameters and leads to prolonged acquisition time for achieving improved signal-to-noise ratio (SNR). Deep learning-based image denoising using convolutional neural networks (CNNs) has superior performance but often requires additional high-SNR data for supervising the training of CNNs, which reduces the practical feasibility. We develop a self-supervised deep learning-based method entitled "SDnDTI" for denoising DTI data, which does not require additional high-SNR data for training. Specifically, SDnDTI divides multi-directional DTI data into many subsets, each consisting of six DWI volumes along optimally chosen diffusion-encoding directions that are robust to noise for the tensor fitting, and then synthesizes DWI volumes along all acquired directions from the diffusion tensors fitted using each subset of the data as the input data of CNNs. On the other hand, SDnDTI synthesizes DWI volumes along acquired diffusion-encoding directions with higher SNR from the diffusion tensors fitted using all acquired data as the training target. SDnDTI removes noise from each subset of synthesized DWI volumes using a deep 3-dimensional CNN to match the quality of the cleaner target DWI volumes and achieves even higher SNR by averaging all subsets of denoised data. The denoising efficacy of SDnDTI is demonstrated on two datasets provided by the Human Connectome Project (HCP) and the Lifespan HCP in Aging. The SDnDTI results preserve image sharpness and textural details and substantially improve upon those from the raw data. The results of SDnDTI are comparable to those from supervised learning-based denoising and outperform those from state-of-the-art conventional denoising algorithms including BM4D, AONLM and MPPCA.
BUDA-SAGE with self-supervised denoising enables fast, distortion-free, high-resolution T2, T2*, para- and dia-magnetic susceptibility mapping
Sep 09, 2021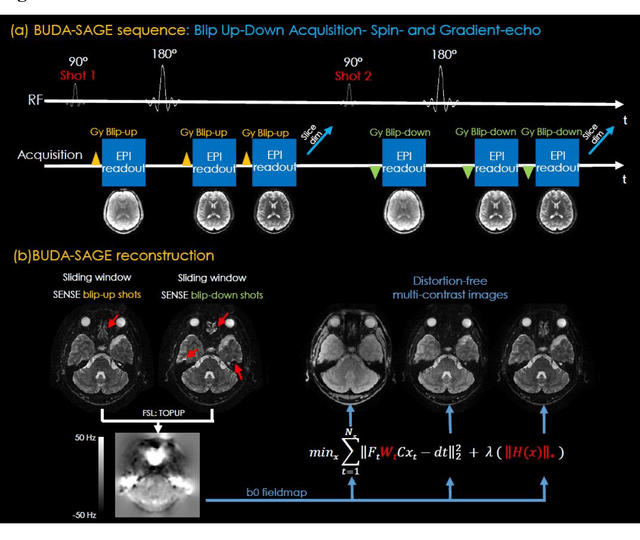
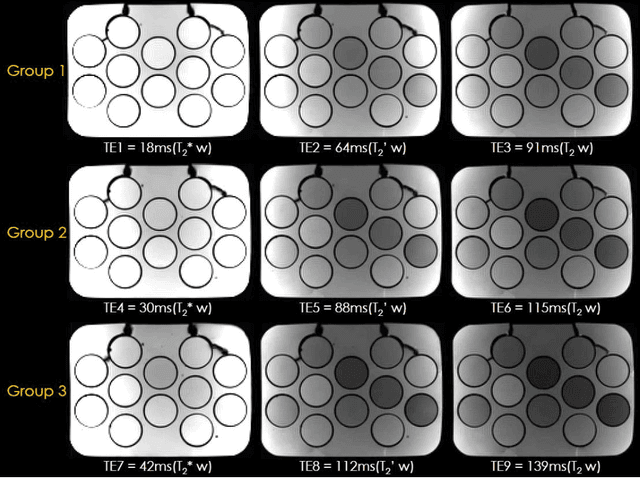
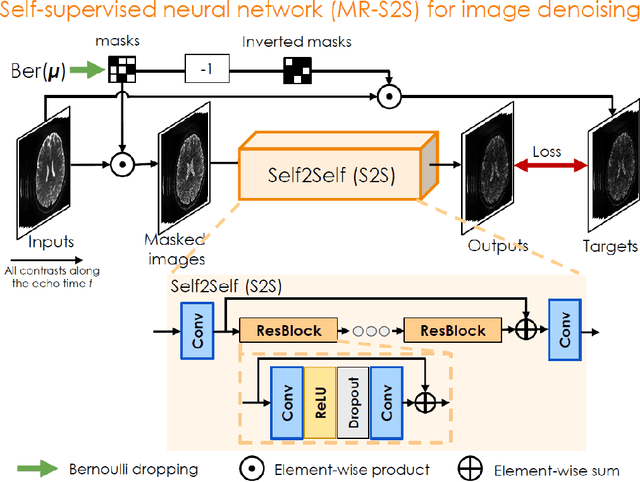

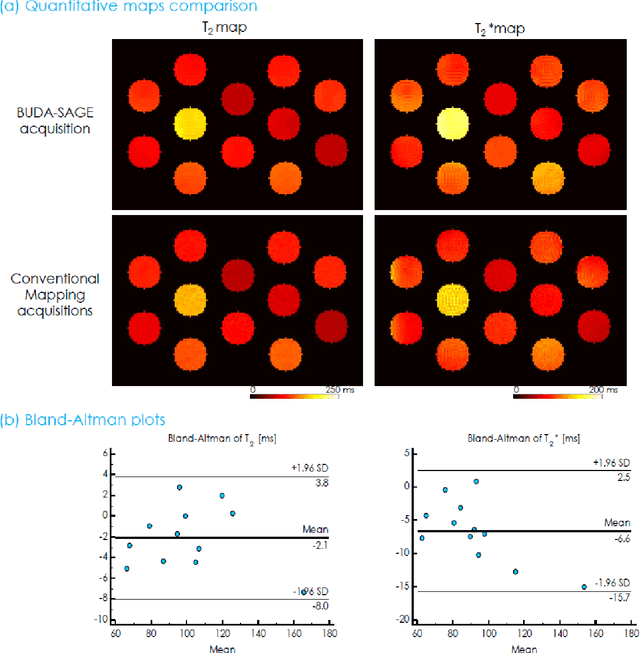
Abstract:To rapidly obtain high resolution T2, T2* and quantitative susceptibility mapping (QSM) source separation maps with whole-brain coverage and high geometric fidelity. We propose Blip Up-Down Acquisition for Spin And Gradient Echo imaging (BUDA-SAGE), an efficient echo-planar imaging (EPI) sequence for quantitative mapping. The acquisition includes multiple T2*-, T2'- and T2-weighted contrasts. We alternate the phase-encoding polarities across the interleaved shots in this multi-shot navigator-free acquisition. A field map estimated from interim reconstructions was incorporated into the joint multi-shot EPI reconstruction with a structured low rank constraint to eliminate geometric distortion. A self-supervised MR-Self2Self (MR-S2S) neural network (NN) was utilized to perform denoising after BUDA reconstruction to boost SNR. Employing Slider encoding allowed us to reach 1 mm isotropic resolution by performing super-resolution reconstruction on BUDA-SAGE volumes acquired with 2 mm slice thickness. Quantitative T2 and T2* maps were obtained using Bloch dictionary matching on the reconstructed echoes. QSM was estimated using nonlinear dipole inversion (NDI) on the gradient echoes. Starting from the estimated R2 and R2* maps, R2' information was derived and used in source separation QSM reconstruction, which provided additional para- and dia-magnetic susceptibility maps. In vivo results demonstrate the ability of BUDA-SAGE to provide whole-brain, distortion-free, high-resolution multi-contrast images and quantitative T2 and T2* maps, as well as yielding para- and dia-magnetic susceptibility maps. Derived quantitative maps showed comparable values to conventional mapping methods in phantom and in vivo measurements. BUDA-SAGE acquisition with self-supervised denoising and Slider encoding enabled rapid, distortion-free, whole-brain T2, T2* mapping at 1 mm3 isotropic resolution in 90 seconds.
Optimized multi-axis spiral projection MR fingerprinting with subspace reconstruction for rapid whole-brain high-isotropic-resolution quantitative imaging
Aug 12, 2021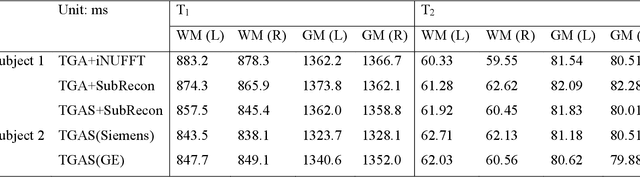

Abstract:Purpose: To improve image quality and accelerate the acquisition of 3D MRF. Methods: Building on the multi-axis spiral-projection MRF technique, a subspace reconstruction with locally low rank (LLR) constraint and a modified spiral-projection spatiotemporal encoding scheme termed tiny-golden-angle-shuffling (TGAS) were implemented for rapid whole-brain high-resolution quantitative mapping. The LLR regularization parameter and the number of subspace bases were tuned using retrospective in-vivo data and simulated examinations, respectively. B0 inhomogeneity correction using multi-frequency interpolation was incorporated into the subspace reconstruction to further improve the image quality by mitigating blurring caused by off-resonance effect. Results: The proposed MRF acquisition and reconstruction framework can produce provide high quality 1-mm isotropic whole-brain quantitative maps in a total acquisition time of 1 minute 55 seconds, with higher-quality results than ones obtained from the previous approach in 6 minutes. The comparison of quantitative results indicates that neither the subspace reconstruction nor the TGAS trajectory induce bias for T1 and T2 mapping. High quality whole-brain MRF data were also obtained at 0.66-mm isotropic resolution in 4 minutes using the proposed technique, where the increased resolution was shown to improve visualization of subtle brain structures. Conclusion: The proposed TGAS-SPI-MRF with optimized spiral-projection trajectory and subspace reconstruction can enable high-resolution quantitative mapping with faster acquisition speed.
eRAKI: Fast Robust Artificial neural networks for K-space Interpolation (RAKI) with Coil Combination and Joint Reconstruction
Jul 07, 2021Abstract:RAKI can perform database-free MRI reconstruction by training models using only auto-calibration signal (ACS) from each specific scan. As it trains a separate model for each individual coil, learning and inference with RAKI can be computationally prohibitive, particularly for large 3D datasets. In this abstract, we accelerate RAKI more than 200 times by directly learning a coil-combined target and further improve the reconstruction performance using joint reconstruction across multiple echoes together with an elliptical-CAIPI sampling approach. We further deploy these improvements in quantitative imaging and rapidly obtain T2 and T2* parameter maps from a fast EPTI scan.
Highly Accelerated EPI with Wave Encoding and Multi-shot Simultaneous Multi-Slice Imaging
Jun 03, 2021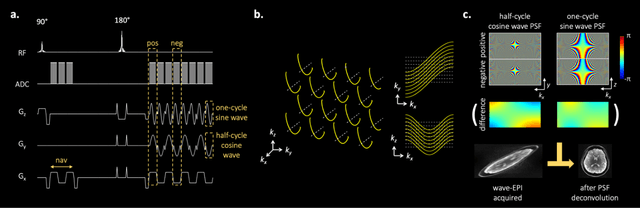
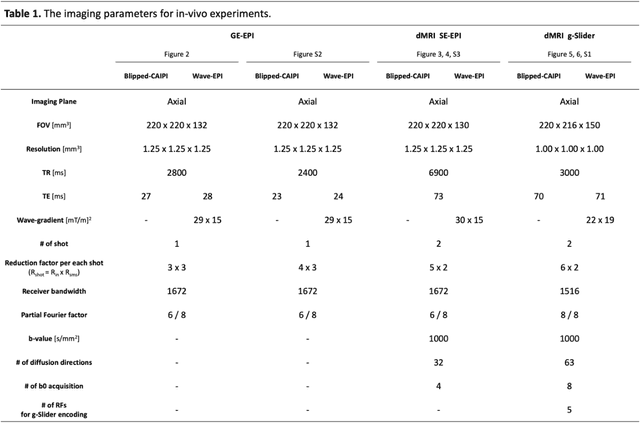
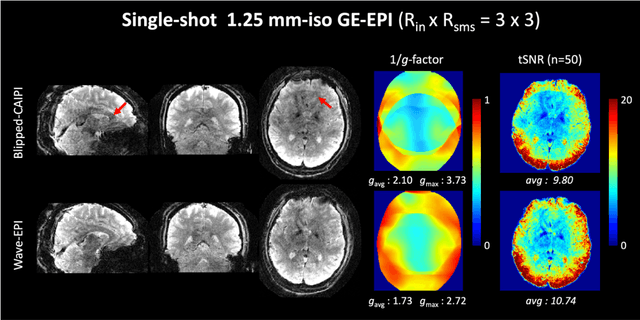
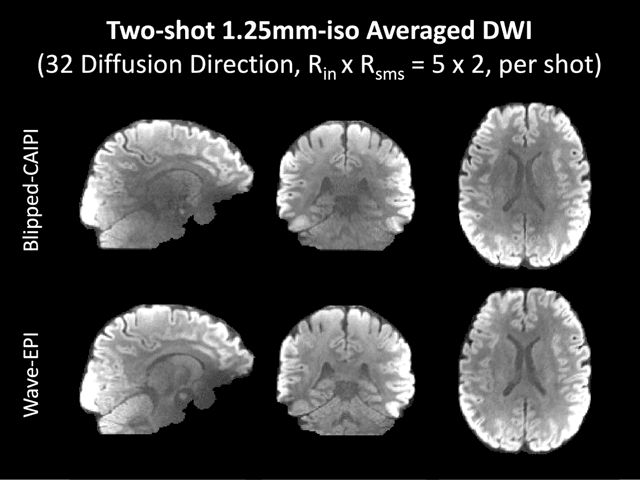
Abstract:We introduce wave encoded acquisition and reconstruction techniques for highly accelerated echo planar imaging (EPI) with reduced g-factor penalty and image artifacts. Wave-EPI involves playing sinusoidal gradients during the EPI readout while employing interslice shifts as in blipped-CAIPI acquisitions. This spreads the aliasing in all spatial directions, thereby taking better advantage of 3D coil sensitivity profiles. The amount of voxel spreading that can be achieved by the wave gradients during the short EPI readout period is constrained by the slew rate of the gradient coils and peripheral nerve stimulation (PNS) monitor. We propose to use a half-cycle sinusoidal gradient to increase the amount of voxel spreading that can be achieved while respecting the slew and stimulation constraints. Extending wave-EPI to multi-shot acquisition minimizes geometric distortion and voxel blurring at high in-plane resolution, while structured low-rank regularization mitigates shot-to-shot phase variations without additional navigators. We propose to use different point spread functions (PSFs) for the k-space lines with positive and negative polarities, which are calibrated with a FLEET-based reference scan and allow for addressing gradient imperfections. Wave-EPI provided whole-brain single-shot gradient echo (GE) and multi-shot spin echo (SE) EPI acquisitions at high acceleration factors and was combined with g-Slider slab encoding to boost the SNR level in 1mm isotropic diffusion imaging. Relative to blipped-CAIPI, wave-EPI reduced average and maximum g-factors by up to 1.21- and 1.37-fold, respectively. In conclusion, wave-EPI allows highly accelerated single- and multi-shot EPI with reduced g-factor and artifacts and may facilitate clinical and neuroscientific applications of EPI by improving the spatial and temporal resolution in functional and diffusion imaging.
Covariance-Free Sparse Bayesian Learning
May 21, 2021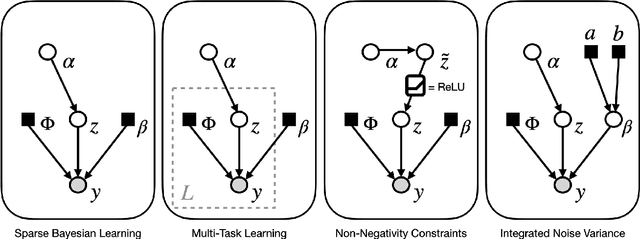
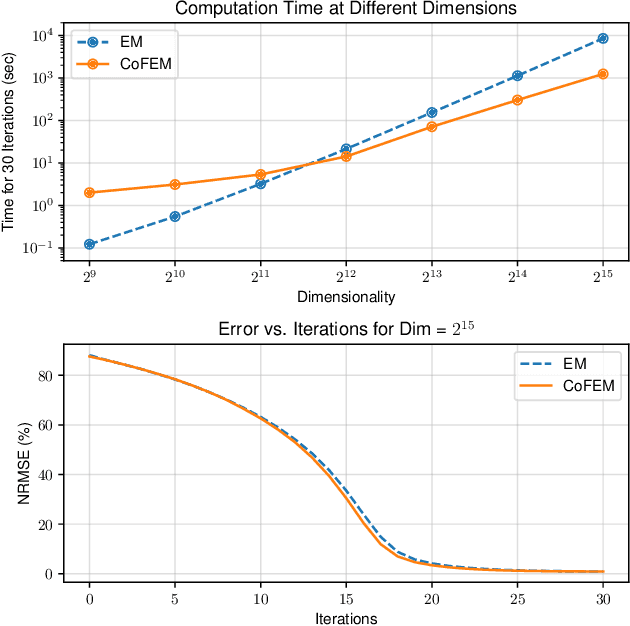
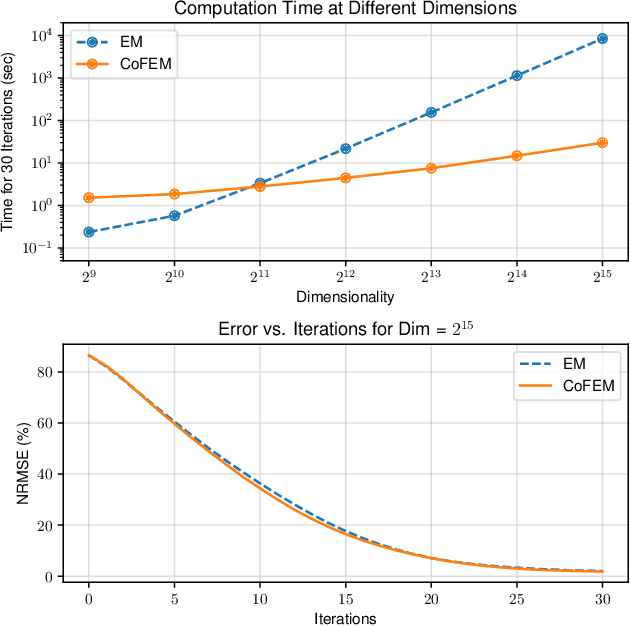
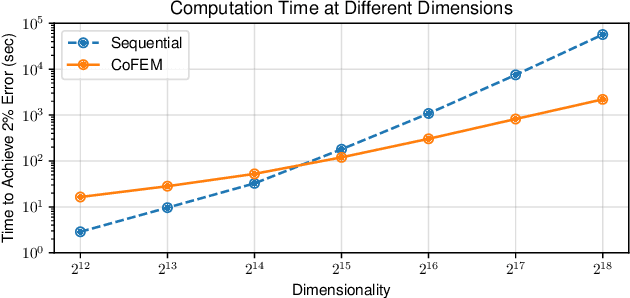
Abstract:Sparse Bayesian learning (SBL) is a powerful framework for tackling the sparse coding problem while also providing uncertainty quantification. However, the most popular inference algorithms for SBL become too expensive for high-dimensional problems due to the need to maintain a large covariance matrix. To resolve this issue, we introduce a new SBL inference algorithm that avoids explicit computation of the covariance matrix, thereby saving significant time and space. Instead of performing costly matrix inversions, our covariance-free method solves multiple linear systems to obtain provably unbiased estimates of the posterior statistics needed by SBL. These systems can be solved in parallel, enabling further acceleration of the algorithm via graphics processing units. In practice, our method can be up to thousands of times faster than existing baselines, reducing hours of computation time to seconds. We showcase how our new algorithm enables SBL to tractably tackle high-dimensional signal recovery problems, such as deconvolution of calcium imaging data and multi-contrast reconstruction of magnetic resonance images. Finally, we open-source a toolbox containing all of our implementations to drive future research in SBL.
Scan Specific Artifact Reduction in K-space Neural Networks Synergize with Physics-based Reconstruction to Accelerate MRI
Apr 02, 2021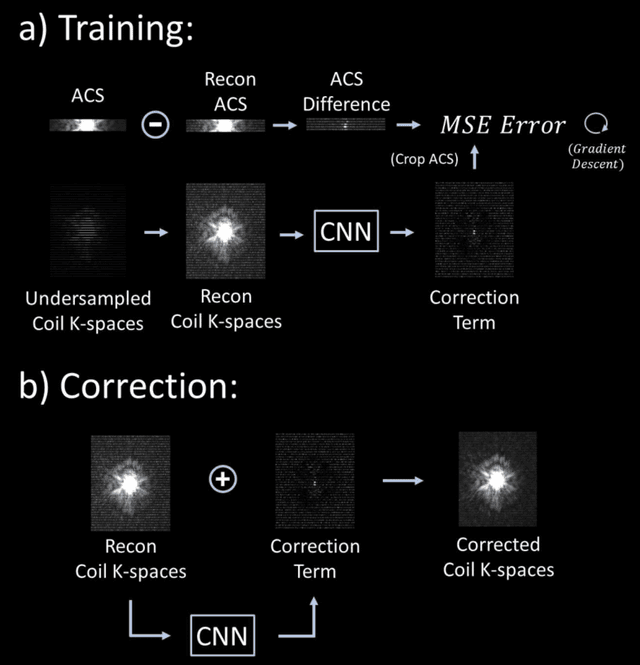
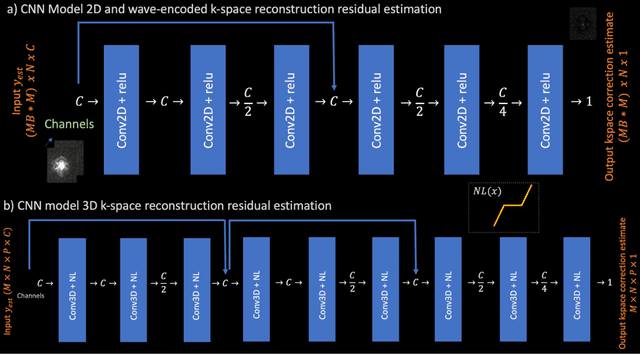
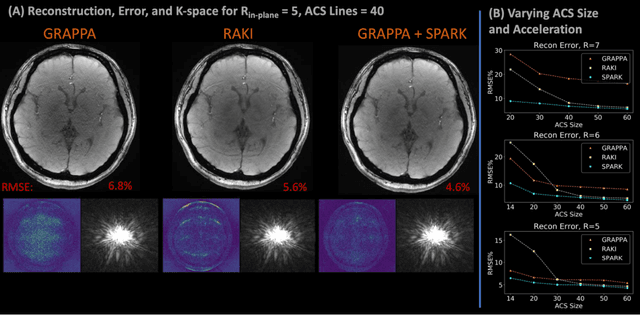
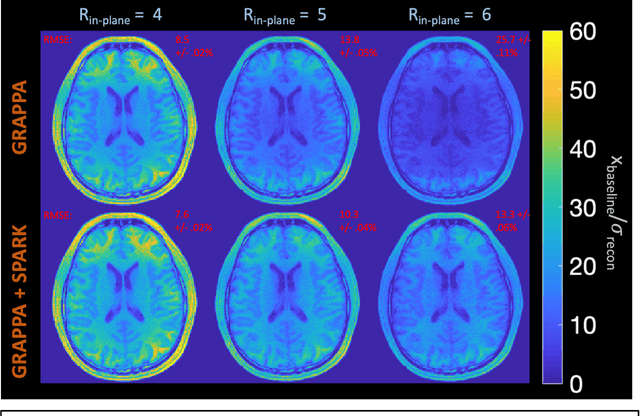
Abstract:Purpose: To develop a scan-specific model that estimates and corrects k-space errors made when reconstructing accelerated Magnetic Resonance Imaging (MRI) data. Methods: Scan-Specific Artifact Reduction in k-space (SPARK) trains a convolutional neural network to estimate k-space errors made by an input reconstruction technique by back-propagating from the mean-squared-error loss between an auto-calibration signal (ACS) and the input technique's reconstructed ACS. First, SPARK is applied to GRAPPA and demonstrates improved robustness over other scan-specific models. Then, SPARK is shown to synergize with advanced reconstruction techniques by improving image quality when applied to 2D virtual coil (VC-) GRAPPA, 2D LORAKS, 3D GRAPPA without an integrated ACS region, and 2D/3D wave-encoded imaging. Results: SPARK yields 1.5 - 2x RMSE reduction when applied to GRAPPA and improves robustness to ACS size for various acceleration rates in comparison to other scan-specific techniques. When applied to advanced parallel imaging techniques such as 2D VC-GRAPPA and LORAKS, SPARK achieves up to 20% RMSE improvement. SPARK with 3D GRAPPA also improves RMSE performance and perceived image quality without a fully sampled ACS region. Finally, SPARK synergizes with non-cartesian, 2D and 3D wave-encoding imaging by reducing RMSE between 20 - 25% and providing qualitative improvements. Conclusion: SPARK synergizes with physics-based reconstruction techniques to improve accelerated MRI by training scan-specific models to estimate and correct reconstruction errors in k-space.
SRDTI: Deep learning-based super-resolution for diffusion tensor MRI
Feb 17, 2021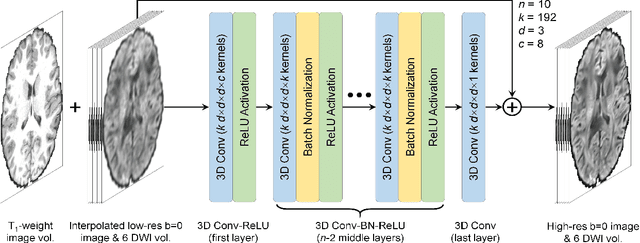
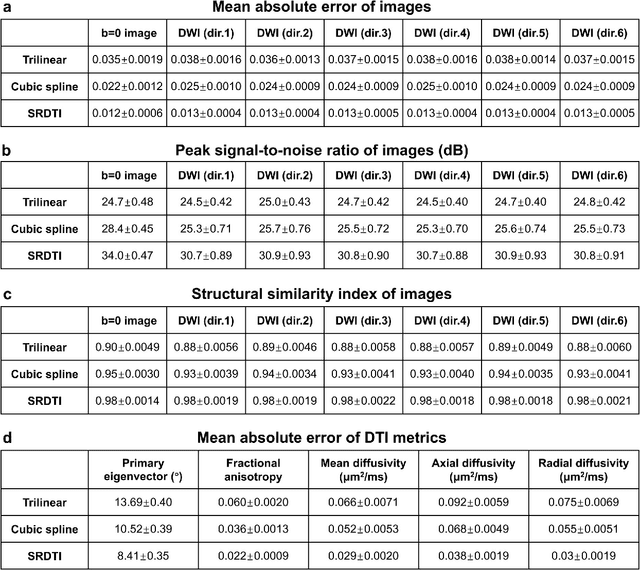
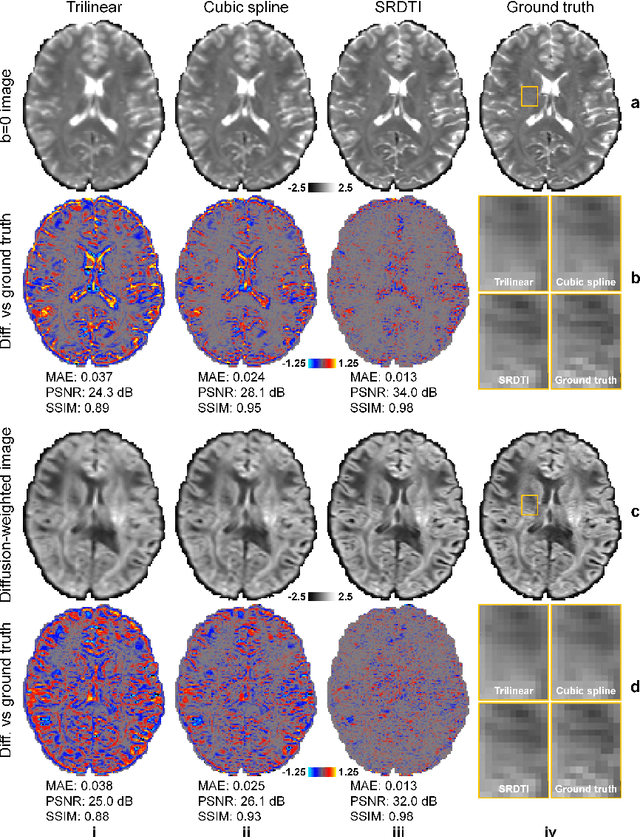
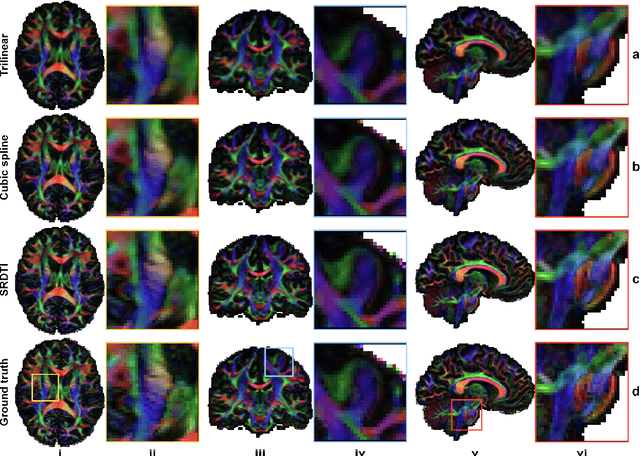
Abstract:High-resolution diffusion tensor imaging (DTI) is beneficial for probing tissue microstructure in fine neuroanatomical structures, but long scan times and limited signal-to-noise ratio pose significant barriers to acquiring DTI at sub-millimeter resolution. To address this challenge, we propose a deep learning-based super-resolution method entitled "SRDTI" to synthesize high-resolution diffusion-weighted images (DWIs) from low-resolution DWIs. SRDTI employs a deep convolutional neural network (CNN), residual learning and multi-contrast imaging, and generates high-quality results with rich textural details and microstructural information, which are more similar to high-resolution ground truth than those from trilinear and cubic spline interpolation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge