Aparna Elangovan
Adaptive Multi-Agent Response Refinement in Conversational Systems
Nov 11, 2025Abstract:Large Language Models (LLMs) have demonstrated remarkable success in conversational systems by generating human-like responses. However, they can fall short, especially when required to account for personalization or specific knowledge. In real-life settings, it is impractical to rely on users to detect these errors and request a new response. One way to address this problem is to refine the response before returning it to the user. While existing approaches focus on refining responses within a single LLM, this method struggles to consider diverse aspects needed for effective conversations. In this work, we propose refining responses through a multi-agent framework, where each agent is assigned a specific role for each aspect. We focus on three key aspects crucial to conversational quality: factuality, personalization, and coherence. Each agent is responsible for reviewing and refining one of these aspects, and their feedback is then merged to improve the overall response. To enhance collaboration among them, we introduce a dynamic communication strategy. Instead of following a fixed sequence of agents, our approach adaptively selects and coordinates the most relevant agents based on the specific requirements of each query. We validate our framework on challenging conversational datasets, demonstrating that ours significantly outperforms relevant baselines, particularly in tasks involving knowledge or user's persona, or both.
Salient Information Prompting to Steer Content in Prompt-based Abstractive Summarization
Oct 03, 2024



Abstract:Large language models (LLMs) can generate fluent summaries across domains using prompting techniques, reducing the need to train models for summarization applications. However, crafting effective prompts that guide LLMs to generate summaries with the appropriate level of detail and writing style remains a challenge. In this paper, we explore the use of salient information extracted from the source document to enhance summarization prompts. We show that adding keyphrases in prompts can improve ROUGE F1 and recall, making the generated summaries more similar to the reference and more complete. The number of keyphrases can control the precision-recall trade-off. Furthermore, our analysis reveals that incorporating phrase-level salient information is superior to word- or sentence-level. However, the impact on hallucination is not universally positive across LLMs. To conduct this analysis, we introduce Keyphrase Signal Extractor (SigExt), a lightweight model that can be finetuned to extract salient keyphrases. By using SigExt, we achieve consistent ROUGE improvements across datasets and open-weight and proprietary LLMs without any LLM customization. Our findings provide insights into leveraging salient information in building prompt-based summarization systems.
Beyond correlation: The impact of human uncertainty in measuring the effectiveness of automatic evaluation and LLM-as-a-judge
Oct 03, 2024



Abstract:The effectiveness of automatic evaluation of generative models is typically measured by comparing it to human evaluation using correlation metrics. However, metrics like Krippendorff's $\alpha$ and Randolph's $\kappa$, originally designed to measure the reliability of human labeling, make assumptions about human behavior and the labeling process. In this paper, we show how *relying on a single aggregate correlation score* can obscure fundamental differences between human behavior and automatic evaluation methods, including LLM-as-a-Judge. Specifically, we demonstrate that when the proportion of samples with variation or uncertainty in human labels (gathered during human evaluation) is relatively high, machine labels (generated by automatic evaluation methods) may superficially appear to have similar or better correlation with the human majority label compared to human-to-human (HH) correlation. This can create the misleading impression that automatic evaluation is accurate enough to approximate the human majority label. However, as the proportion of samples with consistent human labels increases, the correlation between machine labels and human majority labels declines, falling below HH correlation. Based on these findings, we first propose stratifying results by human label uncertainty to provide a more robust analysis of automatic evaluation performance. Second, recognizing that uncertainty and variation are inherent in perception-based human evaluations, such as those involving attitudes or preferences, we introduce a new metric - *binned Jensen-Shannon Divergence for perception* for such scenarios to better measure the effectiveness of automatic evaluations. Third, we present visualization techniques -- *perception charts*, to compare the strengths and limitations of automatic evaluation and to contextualize correlation measures appropriately
ConSiDERS-The-Human Evaluation Framework: Rethinking Human Evaluation for Generative Large Language Models
May 28, 2024Abstract:In this position paper, we argue that human evaluation of generative large language models (LLMs) should be a multidisciplinary undertaking that draws upon insights from disciplines such as user experience research and human behavioral psychology to ensure that the experimental design and results are reliable. The conclusions from these evaluations, thus, must consider factors such as usability, aesthetics, and cognitive biases. We highlight how cognitive biases can conflate fluent information and truthfulness, and how cognitive uncertainty affects the reliability of rating scores such as Likert. Furthermore, the evaluation should differentiate the capabilities and weaknesses of increasingly powerful large language models -- which requires effective test sets. The scalability of human evaluation is also crucial to wider adoption. Hence, to design an effective human evaluation system in the age of generative NLP, we propose the ConSiDERS-The-Human evaluation framework consisting of 6 pillars --Consistency, Scoring Critera, Differentiating, User Experience, Responsible, and Scalability.
Principles from Clinical Research for NLP Model Generalization
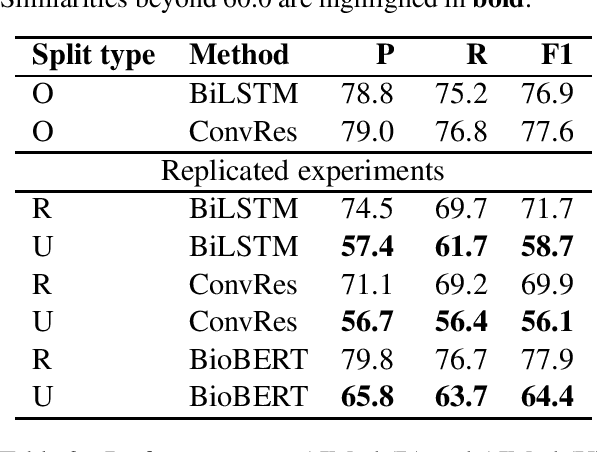
Nov 09, 2023Abstract:The NLP community typically relies on performance of a model on a held-out test set to assess generalization. Performance drops observed in datasets outside of official test sets are generally attributed to "out-of-distribution'' effects. Here, we explore the foundations of generalizability and study the various factors that affect it, articulating generalizability lessons from clinical studies. In clinical research generalizability depends on (a) internal validity of experiments to ensure controlled measurement of cause and effect, and (b) external validity or transportability of the results to the wider population. We present the need to ensure internal validity when building machine learning models in natural language processing, especially where results may be impacted by spurious correlations in the data. We demonstrate how spurious factors, such as the distance between entities in relation extraction tasks, can affect model internal validity and in turn adversely impact generalization. We also offer guidance on how to analyze generalization failures.
Effects of Human Adversarial and Affable Samples on BERT Generalizability
Oct 17, 2023



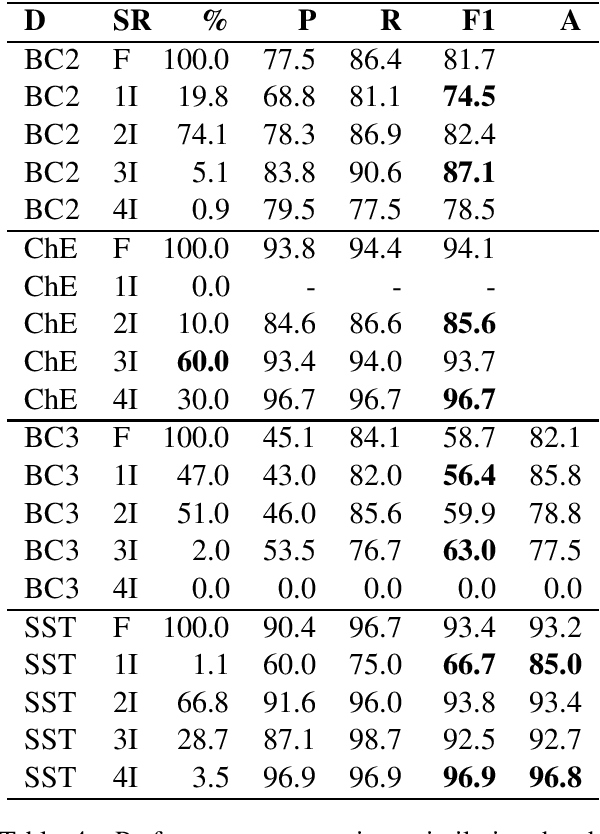
Abstract:BERT-based models have had strong performance on leaderboards, yet have been demonstrably worse in real-world settings requiring generalization. Limited quantities of training data is considered a key impediment to achieving generalizability in machine learning. In this paper, we examine the impact of training data quality, not quantity, on a model's generalizability. We consider two characteristics of training data: the portion of human-adversarial (h-adversarial), i.e., sample pairs with seemingly minor differences but different ground-truth labels, and human-affable (h-affable) training samples, i.e., sample pairs with minor differences but the same ground-truth label. We find that for a fixed size of training samples, as a rule of thumb, having 10-30% h-adversarial instances improves the precision, and therefore F1, by up to 20 points in the tasks of text classification and relation extraction. Increasing h-adversarials beyond this range can result in performance plateaus or even degradation. In contrast, h-affables may not contribute to a model's generalizability and may even degrade generalization performance.
Large-scale protein-protein post-translational modification extraction with distant supervision and confidence calibrated BioBERT
Jan 06, 2022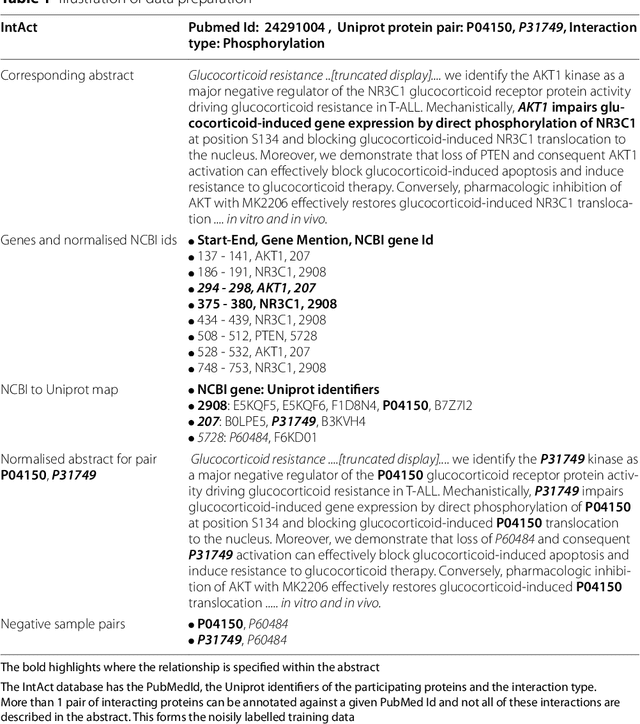
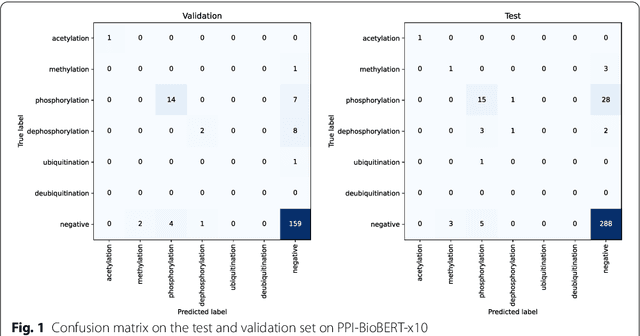
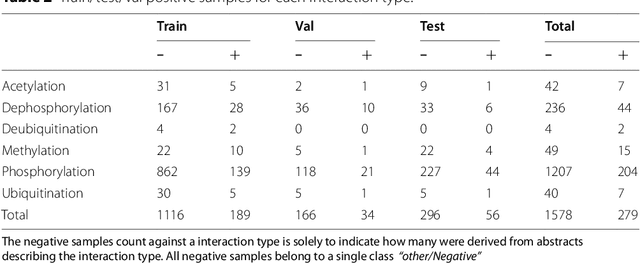
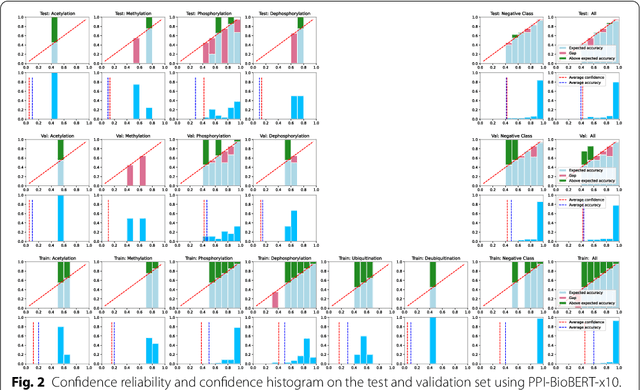
Abstract:Protein-protein interactions (PPIs) are critical to normal cellular function and are related to many disease pathways. However, only 4% of PPIs are annotated with PTMs in biological knowledge databases such as IntAct, mainly performed through manual curation, which is neither time nor cost-effective. We use the IntAct PPI database to create a distant supervised dataset annotated with interacting protein pairs, their corresponding PTM type, and associated abstracts from the PubMed database. We train an ensemble of BioBERT models - dubbed PPI-BioBERT-x10 to improve confidence calibration. We extend the use of ensemble average confidence approach with confidence variation to counteract the effects of class imbalance to extract high confidence predictions. The PPI-BioBERT-x10 model evaluated on the test set resulted in a modest F1-micro 41.3 (P =5 8.1, R = 32.1). However, by combining high confidence and low variation to identify high quality predictions, tuning the predictions for precision, we retained 19% of the test predictions with 100% precision. We evaluated PPI-BioBERT-x10 on 18 million PubMed abstracts and extracted 1.6 million (546507 unique PTM-PPI triplets) PTM-PPI predictions, and filter ~ 5700 (4584 unique) high confidence predictions. Of the 5700, human evaluation on a small randomly sampled subset shows that the precision drops to 33.7% despite confidence calibration and highlights the challenges of generalisability beyond the test set even with confidence calibration. We circumvent the problem by only including predictions associated with multiple papers, improving the precision to 58.8%. In this work, we highlight the benefits and challenges of deep learning-based text mining in practice, and the need for increased emphasis on confidence calibration to facilitate human curation efforts.
* BMC BioInformatics
Memorization vs. Generalization: Quantifying Data Leakage in NLP Performance Evaluation
Feb 03, 2021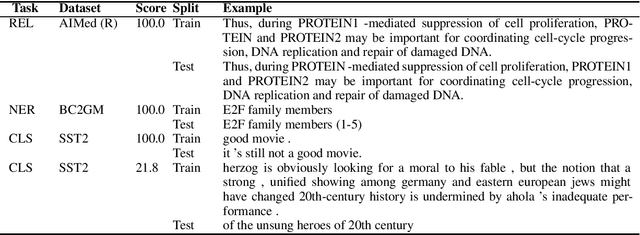
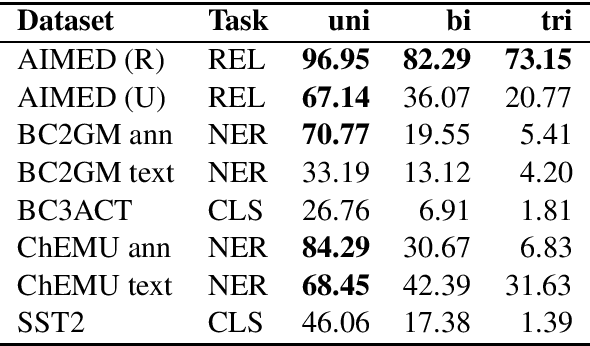
Abstract:Public datasets are often used to evaluate the efficacy and generalizability of state-of-the-art methods for many tasks in natural language processing (NLP). However, the presence of overlap between the train and test datasets can lead to inflated results, inadvertently evaluating the model's ability to memorize and interpreting it as the ability to generalize. In addition, such data sets may not provide an effective indicator of the performance of these methods in real world scenarios. We identify leakage of training data into test data on several publicly available datasets used to evaluate NLP tasks, including named entity recognition and relation extraction, and study them to assess the impact of that leakage on the model's ability to memorize versus generalize.
Assigning function to protein-protein interactions: a weakly supervised BioBERT based approach using PubMed abstracts
Sep 29, 2020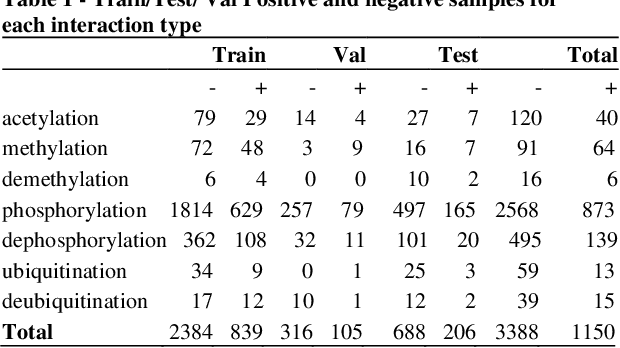
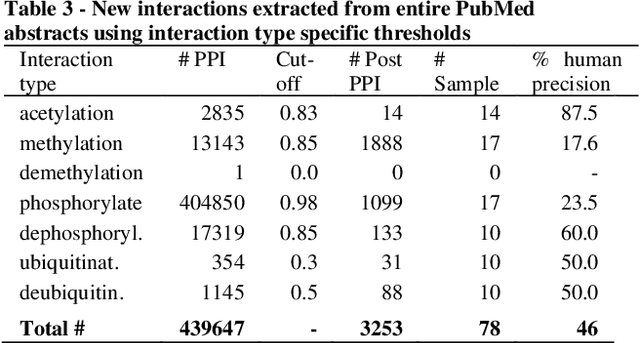
Abstract:Motivation: Protein-protein interactions (PPI) are critical to the function of proteins in both normal and diseased cells, and many critical protein functions are mediated by interactions.Knowledge of the nature of these interactions is important for the construction of networks to analyse biological data. However, only a small percentage of PPIs captured in protein interaction databases have annotations of function available, e.g. only 4% of PPI are functionally annotated in the IntAct database. Here, we aim to label the function type of PPIs by extracting relationships described in PubMed abstracts. Method: We create a weakly supervised dataset from the IntAct PPI database containing interacting protein pairs with annotated function and associated abstracts from the PubMed database. We apply a state-of-the-art deep learning technique for biomedical natural language processing tasks, BioBERT, to build a model - dubbed PPI-BioBERT - for identifying the function of PPIs. In order to extract high quality PPI functions at large scale, we use an ensemble of PPI-BioBERT models to improve uncertainty estimation and apply an interaction type-specific threshold to counteract the effects of variations in the number of training samples per interaction type. Results: We scan 18 million PubMed abstracts to automatically identify 3253 new typed PPIs, including phosphorylation and acetylation interactions, with an overall precision of 46% (87% for acetylation) based on a human-reviewed sample. This work demonstrates that analysis of biomedical abstracts for PPI function extraction is a feasible approach to substantially increasing the number of interactions annotated with function captured in online databases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge