Andrew McCallum
Revisiting the Architectures like Pointer Networks to Efficiently Improve the Next Word Distribution, Summarization Factuality, and Beyond
May 20, 2023



Abstract:Is the output softmax layer, which is adopted by most language models (LMs), always the best way to compute the next word probability? Given so many attention layers in a modern transformer-based LM, are the pointer networks redundant nowadays? In this study, we discover that the answers to both questions are no. This is because the softmax bottleneck sometimes prevents the LMs from predicting the desired distribution and the pointer networks can be used to break the bottleneck efficiently. Based on the finding, we propose several softmax alternatives by simplifying the pointer networks and accelerating the word-by-word rerankers. In GPT-2, our proposals are significantly better and more efficient than mixture of softmax, a state-of-the-art softmax alternative. In summarization experiments, without significantly decreasing its training/testing speed, our best method based on T5-Small improves factCC score by 2 points in CNN/DM and XSUM dataset, and improves MAUVE scores by 30% in BookSum paragraph-level dataset.
Adaptive Selection of Anchor Items for CUR-based k-NN search with Cross-Encoders
May 04, 2023



Abstract:Cross-encoder models, which jointly encode and score a query-item pair, are typically prohibitively expensive for k-nearest neighbor search. Consequently, k-NN search is performed not with a cross-encoder, but with a heuristic retrieve (e.g., using BM25 or dual-encoder) and re-rank approach. Recent work proposes ANNCUR (Yadav et al., 2022) which uses CUR matrix factorization to produce an embedding space for efficient vector-based search that directly approximates the cross-encoder without the need for dual-encoders. ANNCUR defines this shared query-item embedding space by scoring the test query against anchor items which are sampled uniformly at random. While this minimizes average approximation error over all items, unsuitably high approximation error on top-k items remains and leads to poor recall of top-k (and especially top-1) items. Increasing the number of anchor items is a straightforward way of improving the approximation error and hence k-NN recall of ANNCUR but at the cost of increased inference latency. In this paper, we propose a new method for adaptively choosing anchor items that minimizes the approximation error for the practically important top-k neighbors for a query with minimal computational overhead. Our proposed method incrementally selects a suitable set of anchor items for a given test query over several rounds, using anchors chosen in previous rounds to inform selection of more anchor items. Empirically, our method consistently improves k-NN recall as compared to both ANNCUR and the widely-used dual-encoder-based retrieve-and-rerank approach.
Editable User Profiles for Controllable Text Recommendation
Apr 09, 2023



Abstract:Methods for making high-quality recommendations often rely on learning latent representations from interaction data. These methods, while performant, do not provide ready mechanisms for users to control the recommendation they receive. Our work tackles this problem by proposing LACE, a novel concept value bottleneck model for controllable text recommendations. LACE represents each user with a succinct set of human-readable concepts through retrieval given user-interacted documents and learns personalized representations of the concepts based on user documents. This concept based user profile is then leveraged to make recommendations. The design of our model affords control over the recommendations through a number of intuitive interactions with a transparent user profile. We first establish the quality of recommendations obtained from LACE in an offline evaluation on three recommendation tasks spanning six datasets in warm-start, cold-start, and zero-shot setups. Next, we validate the controllability of LACE under simulated user interactions. Finally, we implement LACE in an interactive controllable recommender system and conduct a user study to demonstrate that users are able to improve the quality of recommendations they receive through interactions with an editable user profile.
Improving Dual-Encoder Training through Dynamic Indexes for Negative Mining
Mar 27, 2023



Abstract:Dual encoder models are ubiquitous in modern classification and retrieval. Crucial for training such dual encoders is an accurate estimation of gradients from the partition function of the softmax over the large output space; this requires finding negative targets that contribute most significantly ("hard negatives"). Since dual encoder model parameters change during training, the use of traditional static nearest neighbor indexes can be sub-optimal. These static indexes (1) periodically require expensive re-building of the index, which in turn requires (2) expensive re-encoding of all targets using updated model parameters. This paper addresses both of these challenges. First, we introduce an algorithm that uses a tree structure to approximate the softmax with provable bounds and that dynamically maintains the tree. Second, we approximate the effect of a gradient update on target encodings with an efficient Nystrom low-rank approximation. In our empirical study on datasets with over twenty million targets, our approach cuts error by half in relation to oracle brute-force negative mining. Furthermore, our method surpasses prior state-of-the-art while using 150x less accelerator memory.
Low-Resource Compositional Semantic Parsing with Concept Pretraining
Jan 30, 2023Abstract:Semantic parsing plays a key role in digital voice assistants such as Alexa, Siri, and Google Assistant by mapping natural language to structured meaning representations. When we want to improve the capabilities of a voice assistant by adding a new domain, the underlying semantic parsing model needs to be retrained using thousands of annotated examples from the new domain, which is time-consuming and expensive. In this work, we present an architecture to perform such domain adaptation automatically, with only a small amount of metadata about the new domain and without any new training data (zero-shot) or with very few examples (few-shot). We use a base seq2seq (sequence-to-sequence) architecture and augment it with a concept encoder that encodes intent and slot tags from the new domain. We also introduce a novel decoder-focused approach to pretrain seq2seq models to be concept aware using Wikidata and use it to help our model learn important concepts and perform well in low-resource settings. We report few-shot and zero-shot results for compositional semantic parsing on the TOPv2 dataset and show that our model outperforms prior approaches in few-shot settings for the TOPv2 and SNIPS datasets.
You can't pick your neighbors, or can you? When and how to rely on retrieval in the $k$NN-LM
Oct 28, 2022Abstract:Retrieval-enhanced language models (LMs), which condition their predictions on text retrieved from large external datastores, have recently shown significant perplexity improvements compared to standard LMs. One such approach, the $k$NN-LM, interpolates any existing LM's predictions with the output of a $k$-nearest neighbors model and requires no additional training. In this paper, we explore the importance of lexical and semantic matching in the context of items retrieved by $k$NN-LM. We find two trends: (1) the presence of large overlapping $n$-grams between the datastore and evaluation set plays an important factor in strong performance, even when the datastore is derived from the training data; and (2) the $k$NN-LM is most beneficial when retrieved items have high semantic similarity with the query. Based on our analysis, we define a new formulation of the $k$NN-LM that uses retrieval quality to assign the interpolation coefficient. We empirically measure the effectiveness of our approach on two English language modeling datasets, Wikitext-103 and PG-19. Our re-formulation of the $k$NN-LM is beneficial in both cases, and leads to nearly 4% improvement in perplexity on the Wikitext-103 test set.
Efficient Nearest Neighbor Search for Cross-Encoder Models using Matrix Factorization
Oct 23, 2022Abstract:Efficient k-nearest neighbor search is a fundamental task, foundational for many problems in NLP. When the similarity is measured by dot-product between dual-encoder vectors or $\ell_2$-distance, there already exist many scalable and efficient search methods. But not so when similarity is measured by more accurate and expensive black-box neural similarity models, such as cross-encoders, which jointly encode the query and candidate neighbor. The cross-encoders' high computational cost typically limits their use to reranking candidates retrieved by a cheaper model, such as dual encoder or TF-IDF. However, the accuracy of such a two-stage approach is upper-bounded by the recall of the initial candidate set, and potentially requires additional training to align the auxiliary retrieval model with the cross-encoder model. In this paper, we present an approach that avoids the use of a dual-encoder for retrieval, relying solely on the cross-encoder. Retrieval is made efficient with CUR decomposition, a matrix decomposition approach that approximates all pairwise cross-encoder distances from a small subset of rows and columns of the distance matrix. Indexing items using our approach is computationally cheaper than training an auxiliary dual-encoder model through distillation. Empirically, for k > 10, our approach provides test-time recall-vs-computational cost trade-offs superior to the current widely-used methods that re-rank items retrieved using a dual-encoder or TF-IDF.
Multi-CLS BERT: An Efficient Alternative to Traditional Ensembling
Oct 10, 2022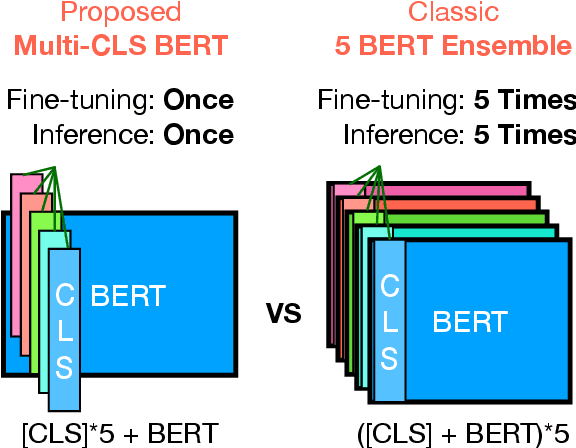
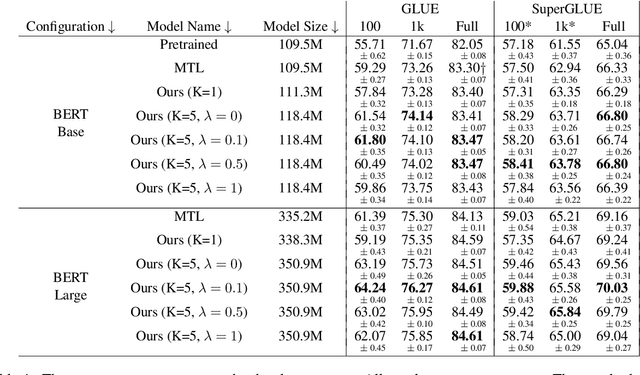
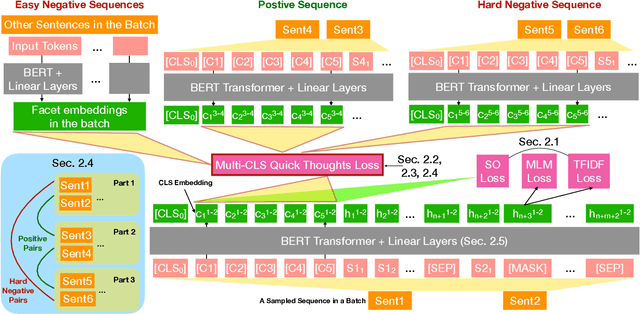
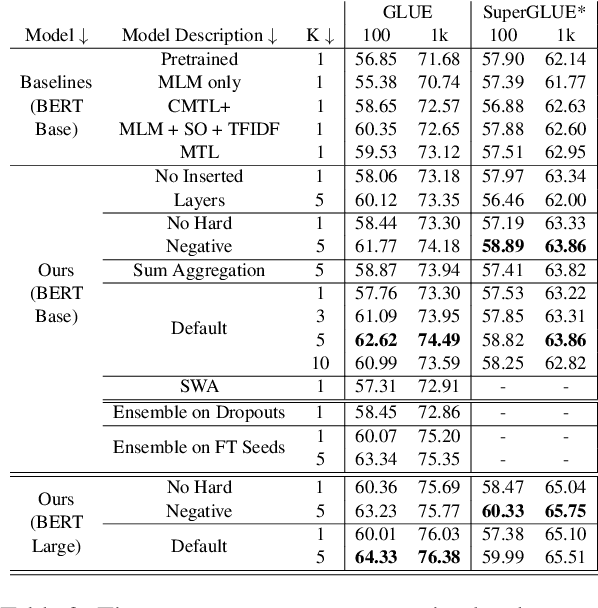
Abstract:Ensembling BERT models often significantly improves accuracy, but at the cost of significantly more computation and memory footprint. In this work, we propose Multi-CLS BERT, a novel ensembling method for CLS-based prediction tasks that is almost as efficient as a single BERT model. Multi-CLS BERT uses multiple CLS tokens with a parameterization and objective that encourages their diversity. Thus instead of fine-tuning each BERT model in an ensemble (and running them all at test time), we need only fine-tune our single Multi-CLS BERT model (and run the one model at test time, ensembling just the multiple final CLS embeddings). To test its effectiveness, we build Multi-CLS BERT on top of a state-of-the-art pretraining method for BERT (Aroca-Ouellette and Rudzicz, 2020). In experiments on GLUE and SuperGLUE we show that our Multi-CLS BERT reliably improves both overall accuracy and confidence estimation. When only 100 training samples are available in GLUE, the Multi-CLS BERT_Base model can even outperform the corresponding BERT_Large model. We analyze the behavior of our Multi-CLS BERT, showing that it has many of the same characteristics and behavior as a typical BERT 5-way ensemble, but with nearly 4-times less computation and memory.
Longtonotes: OntoNotes with Longer Coreference Chains
Oct 07, 2022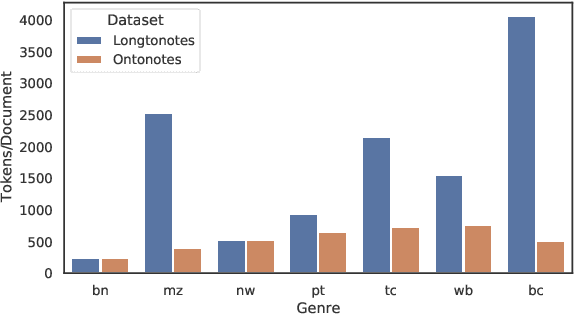
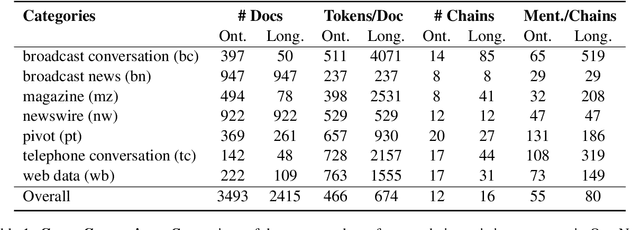
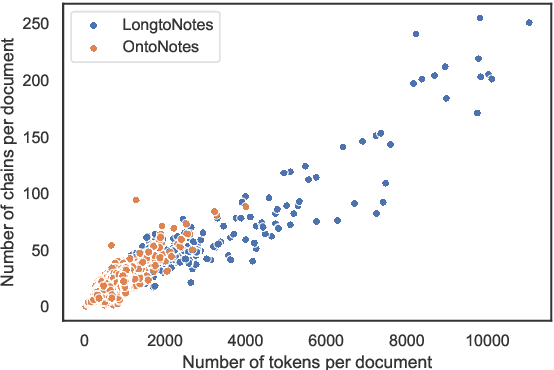
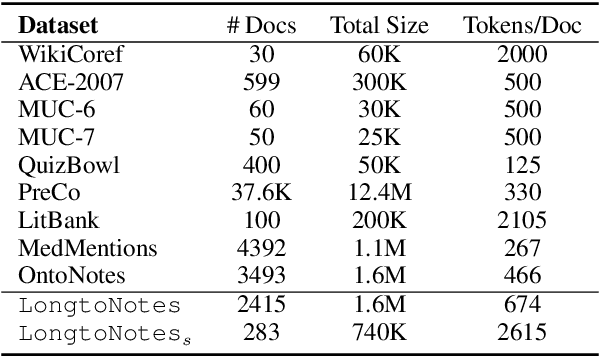
Abstract:Ontonotes has served as the most important benchmark for coreference resolution. However, for ease of annotation, several long documents in Ontonotes were split into smaller parts. In this work, we build a corpus of coreference-annotated documents of significantly longer length than what is currently available. We do so by providing an accurate, manually-curated, merging of annotations from documents that were split into multiple parts in the original Ontonotes annotation process. The resulting corpus, which we call LongtoNotes contains documents in multiple genres of the English language with varying lengths, the longest of which are up to 8x the length of documents in Ontonotes, and 2x those in Litbank. We evaluate state-of-the-art neural coreference systems on this new corpus, analyze the relationships between model architectures/hyperparameters and document length on performance and efficiency of the models, and demonstrate areas of improvement in long-document coreference modeling revealed by our new corpus. Our data and code is available at: https://github.com/kumar-shridhar/LongtoNotes.
Augmenting Scientific Creativity with Retrieval across Knowledge Domains
Jun 02, 2022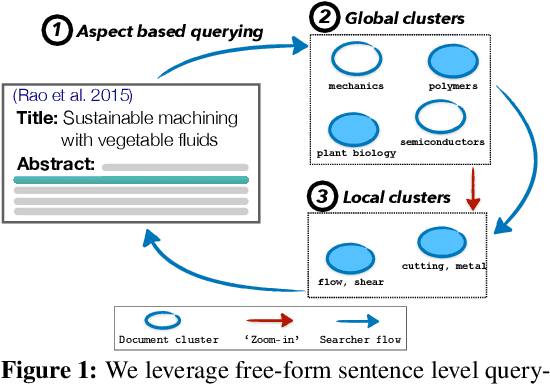
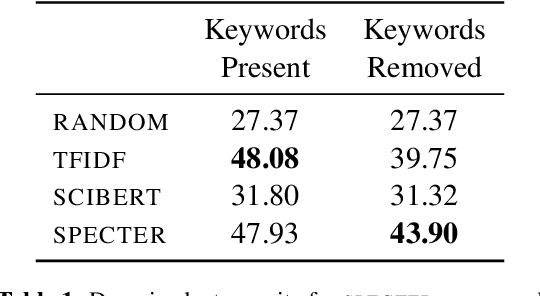
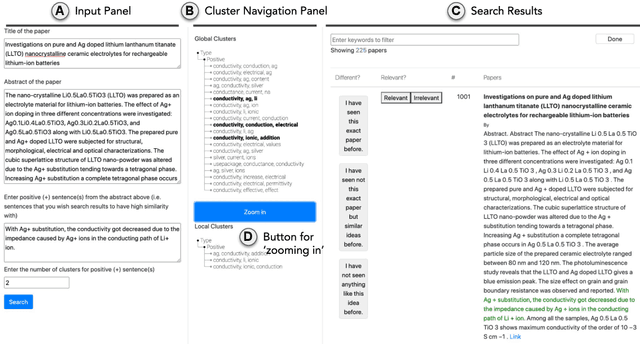

Abstract:Exposure to ideas in domains outside a scientist's own may benefit her in reformulating existing research problems in novel ways and discovering new application domains for existing solution ideas. While improved performance in scholarly search engines can help scientists efficiently identify relevant advances in domains they may already be familiar with, it may fall short of helping them explore diverse ideas \textit{outside} such domains. In this paper we explore the design of systems aimed at augmenting the end-user ability in cross-domain exploration with flexible query specification. To this end, we develop an exploratory search system in which end-users can select a portion of text core to their interest from a paper abstract and retrieve papers that have a high similarity to the user-selected core aspect but differ in terms of domains. Furthermore, end-users can `zoom in' to specific domain clusters to retrieve more papers from them and understand nuanced differences within the clusters. Our case studies with scientists uncover opportunities and design implications for systems aimed at facilitating cross-domain exploration and inspiration.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge