"cancer detection": models, code, and papers
Breast Cancer Classification from Histopathological Images with Inception Recurrent Residual Convolutional Neural Network
Nov 10, 2018
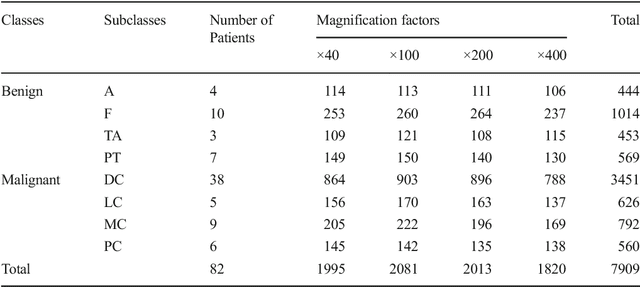
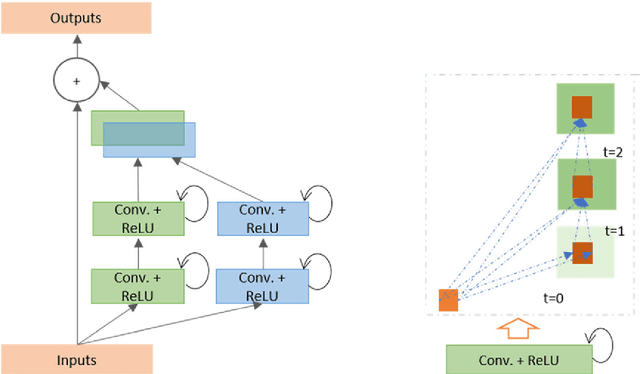
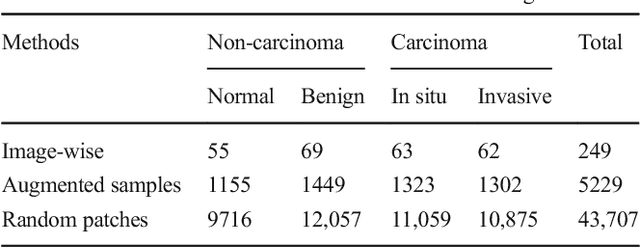
The Deep Convolutional Neural Network (DCNN) is one of the most powerful and successful deep learning approaches. DCNNs have already provided superior performance in different modalities of medical imaging including breast cancer classification, segmentation, and detection. Breast cancer is one of the most common and dangerous cancers impacting women worldwide. In this paper, we have proposed a method for breast cancer classification with the Inception Recurrent Residual Convolutional Neural Network (IRRCNN) model. The IRRCNN is a powerful DCNN model that combines the strength of the Inception Network (Inception-v4), the Residual Network (ResNet), and the Recurrent Convolutional Neural Network (RCNN). The IRRCNN shows superior performance against equivalent Inception Networks, Residual Networks, and RCNNs for object recognition tasks. In this paper, the IRRCNN approach is applied for breast cancer classification on two publicly available datasets including BreakHis and Breast Cancer Classification Challenge 2015. The experimental results are compared against the existing machine learning and deep learning-based approaches with respect to image-based, patch-based, image-level, and patient-level classification. The IRRCNN model provides superior classification performance in terms of sensitivity, Area Under the Curve (AUC), the ROC curve, and global accuracy compared to existing approaches for both datasets.
Segmentation of Infrared Breast Images Using MultiResUnet Neural Network
Oct 31, 2020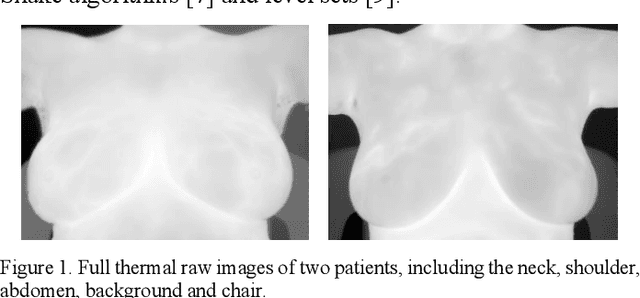


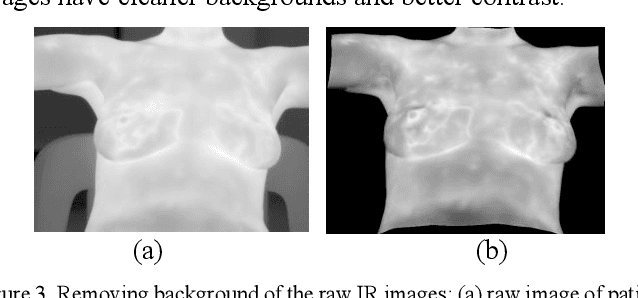
Breast cancer is the second leading cause of death for women in the U.S. Early detection of breast cancer is key to higher survival rates of breast cancer patients. We are investigating infrared (IR) thermography as a noninvasive adjunct to mammography for breast cancer screening. IR imaging is radiation-free, pain-free, and non-contact. Automatic segmentation of the breast area from the acquired full-size breast IR images will help limit the area for tumor search, as well as reduce the time and effort costs of manual segmentation. Autoencoder-like convolutional and deconvolutional neural networks (C-DCNN) had been applied to automatically segment the breast area in IR images in previous studies. In this study, we applied a state-of-the-art deep-learning segmentation model, MultiResUnet, which consists of an encoder part to capture features and a decoder part for precise localization. It was used to segment the breast area by using a set of breast IR images, collected in our pilot study by imaging breast cancer patients and normal volunteers with a thermal infrared camera (N2 Imager). The database we used has 450 images, acquired from 14 patients and 16 volunteers. We used a thresholding method to remove interference in the raw images and remapped them from the original 16-bit to 8-bit, and then cropped and segmented the 8-bit images manually. Experiments using leave-one-out cross-validation (LOOCV) and comparison with the ground-truth images by using Tanimoto similarity show that the average accuracy of MultiResUnet is 91.47%, which is about 2% higher than that of the autoencoder. MultiResUnet offers a better approach to segment breast IR images than our previous model.
Gleason Grading of Histology Prostate Images through Semantic Segmentation via Residual U-Net
May 22, 2020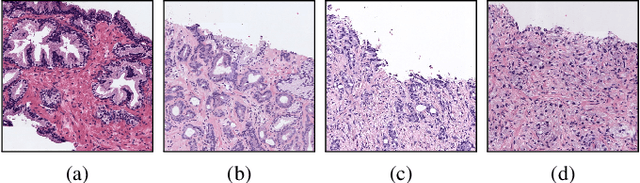
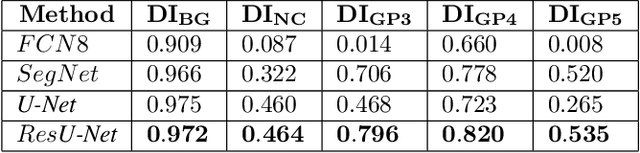
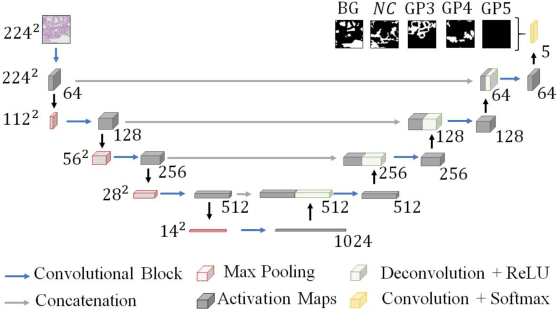
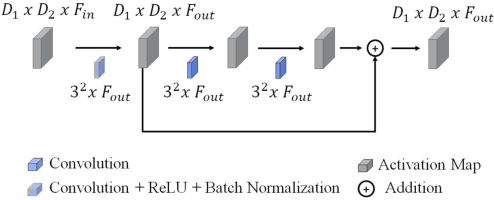
Worldwide, prostate cancer is one of the main cancers affecting men. The final diagnosis of prostate cancer is based on the visual detection of Gleason patterns in prostate biopsy by pathologists. Computer-aided-diagnosis systems allow to delineate and classify the cancerous patterns in the tissue via computer-vision algorithms in order to support the physicians' task. The methodological core of this work is a U-Net convolutional neural network for image segmentation modified with residual blocks able to segment cancerous tissue according to the full Gleason system. This model outperforms other well-known architectures, and reaches a pixel-level Cohen's quadratic Kappa of 0.52, at the level of previous image-level works in the literature, but providing also a detailed localisation of the patterns.
Deep Transfer Learning Methods for Colon Cancer Classification in Confocal Laser Microscopy Images
May 20, 2019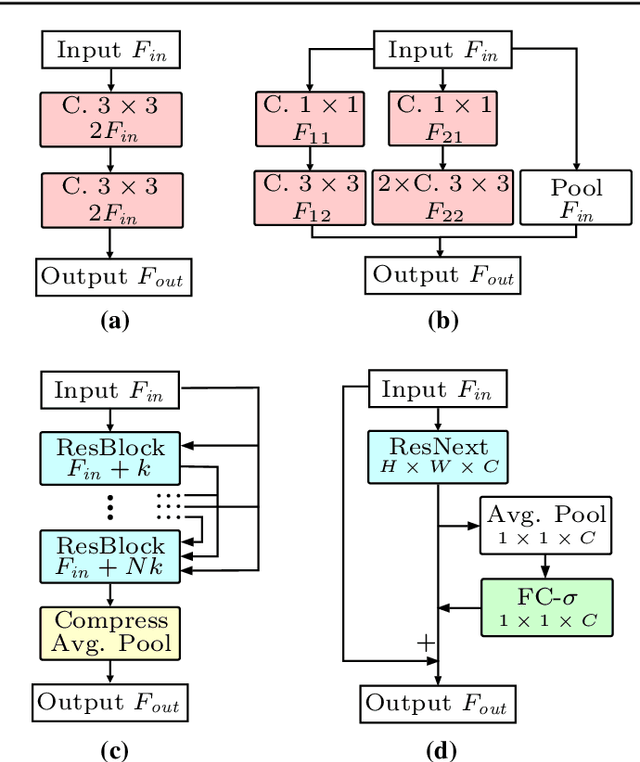
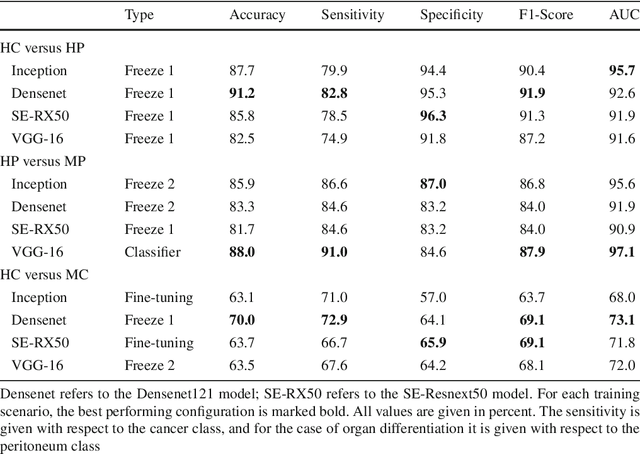
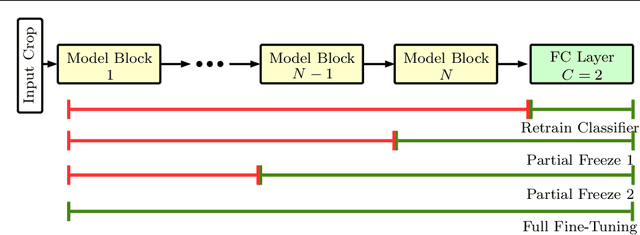
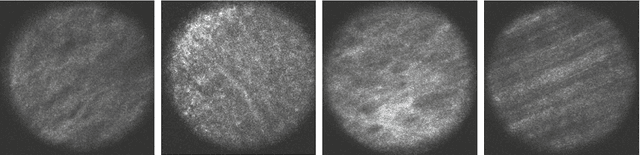
Purpose: The gold standard for colorectal cancer metastases detection in the peritoneum is histological evaluation of a removed tissue sample. For feedback during interventions, real-time in-vivo imaging with confocal laser microscopy has been proposed for differentiation of benign and malignant tissue by manual expert evaluation. Automatic image classification could improve the surgical workflow further by providing immediate feedback. Methods: We analyze the feasibility of classifying tissue from confocal laser microscopy in the colon and peritoneum. For this purpose, we adopt both classical and state-of-the-art convolutional neural networks to directly learn from the images. As the available dataset is small, we investigate several transfer learning strategies including partial freezing variants and full fine-tuning. We address the distinction of different tissue types, as well as benign and malignant tissue. Results: We present a thorough analysis of transfer learning strategies for colorectal cancer with confocal laser microscopy. In the peritoneum, metastases are classified with an AUC of 97.1 and in the colon, the primarius is classified with an AUC of 73.1. In general, transfer learning substantially improves performance over training from scratch. We find that the optimal transfer learning strategy differs for models and classification tasks. Conclusions: We demonstrate that convolutional neural networks and transfer learning can be used to identify cancer tissue with confocal laser microscopy. We show that there is no generally optimal transfer learning strategy and model as well as task-specific engineering is required. Given the high performance for the peritoneum, even with a small dataset, application for intraoperative decision support could be feasible.
Identifying Cancer Patients at Risk for Heart Failure Using Machine Learning Methods
Oct 01, 2019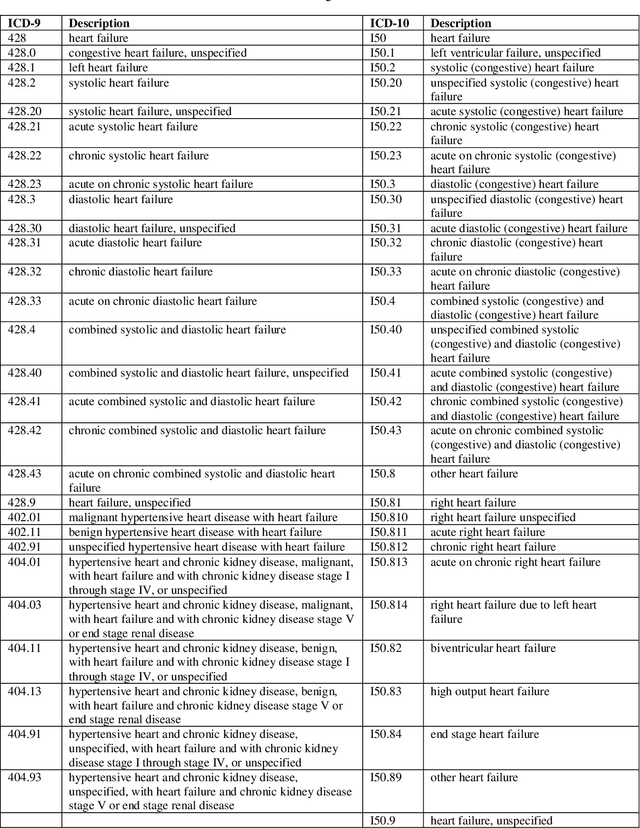
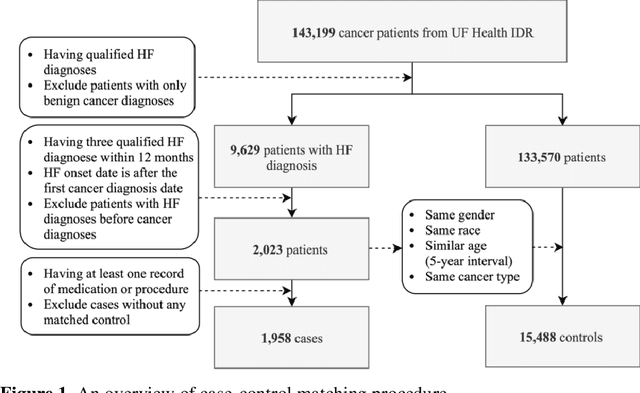
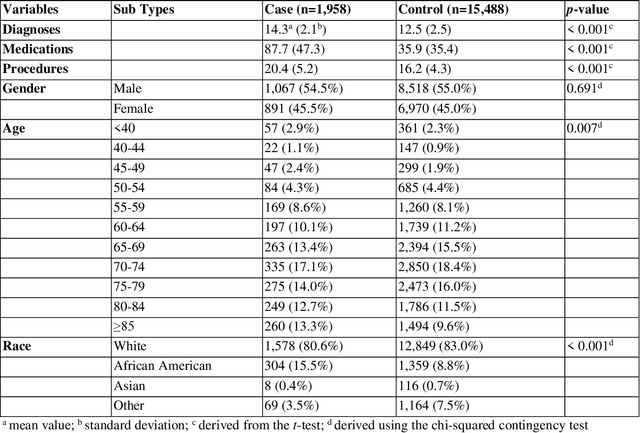
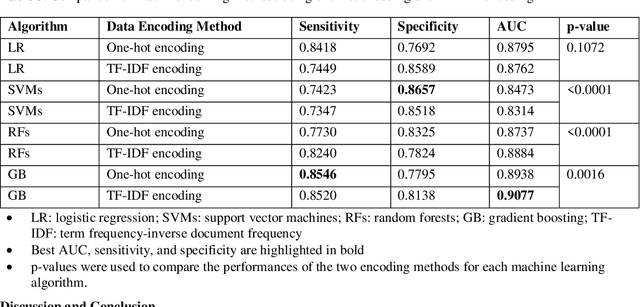
Cardiotoxicity related to cancer therapies has become a serious issue, diminishing cancer treatment outcomes and quality of life. Early detection of cancer patients at risk for cardiotoxicity before cardiotoxic treatments and providing preventive measures are potential solutions to improve cancer patients's quality of life. This study focuses on predicting the development of heart failure in cancer patients after cancer diagnoses using historical electronic health record (EHR) data. We examined four machine learning algorithms using 143,199 cancer patients from the University of Florida Health (UF Health) Integrated Data Repository (IDR). We identified a total number of 1,958 qualified cases and matched them to 15,488 controls by gender, age, race, and major cancer type. Two feature encoding strategies were compared to encode variables as machine learning features. The gradient boosting (GB) based model achieved the best AUC score of 0.9077 (with a sensitivity of 0.8520 and a specificity of 0.8138), outperforming other machine learning methods. We also looked into the subgroup of cancer patients with exposure to chemotherapy drugs and observed a lower specificity score (0.7089). The experimental results show that machine learning methods are able to capture clinical factors that are known to be associated with heart failure and that it is feasible to use machine learning methods to identify cancer patients at risk for cancer therapy-related heart failure.
Disease Detection from Lung X-ray Images based on Hybrid Deep Learning
Mar 02, 2020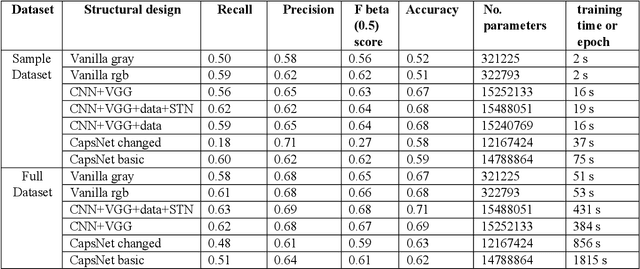
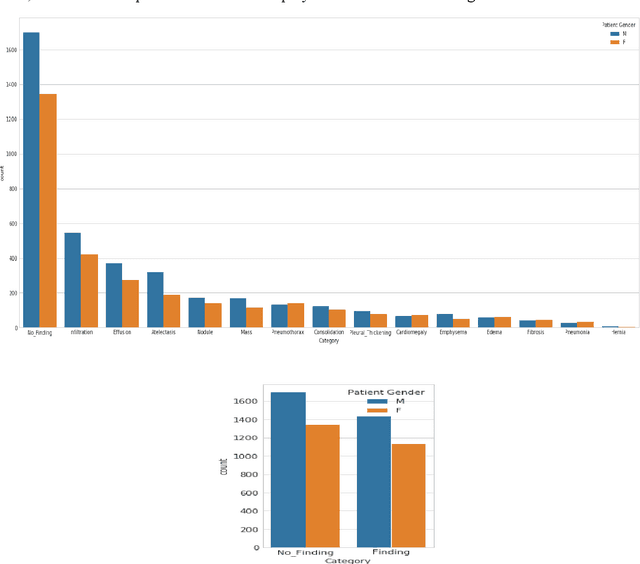
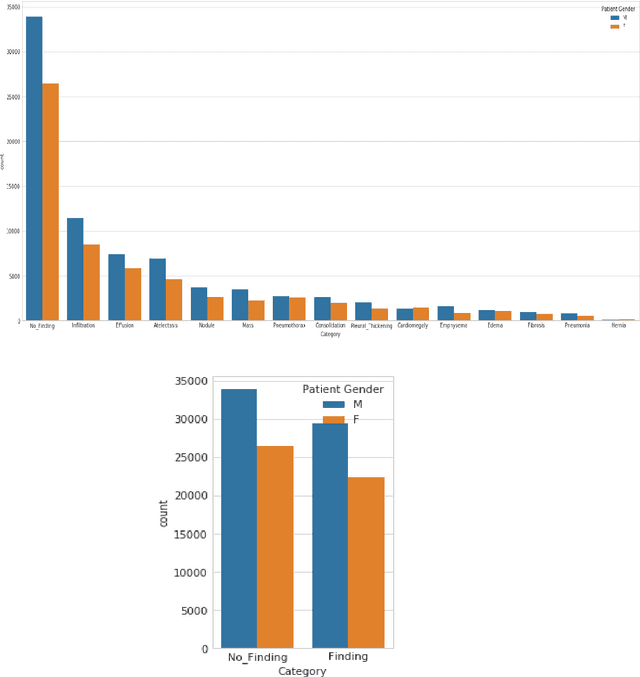
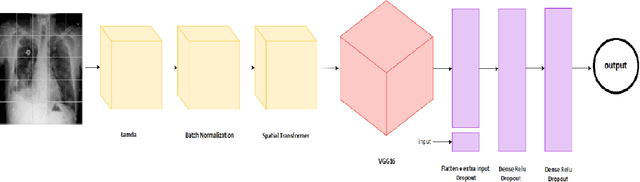
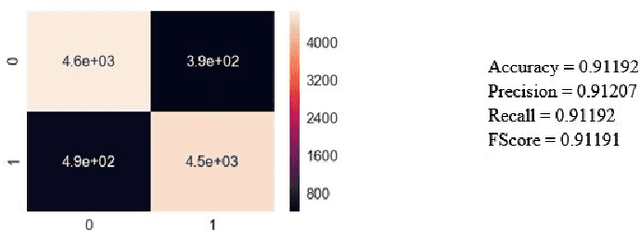
Lung Disease can be considered as the second most common type of disease for men and women. Many people die of lung disease such as lung cancer, Asthma, CPD (Chronic pulmonary disease) etc. in every year. Early detection of lung cancer can lessen the probability of deaths. In this paper, a chest X ray image dataset has been used in order to diagnosis properly and analysis the lung disease. For binary classification, some important is selected. The criteria include precision, recall, F beta score and accuracy. The fusion of AI and cancer diagnosis are acquiring huge interest as a cancer diagnostic tool. In recent days, deep learning based AI for example Convolutional neural network (CNN) can be successfully applied for disease classification and prediction. This paper mainly focuses the performance of Vanilla neural network, CNN, fusion of CNN and Visual Geometry group based neural network (VGG), fusion of CNN, VGG, STN and finally Capsule network. Normally basic CNN has poor performance for rotated, tilted or other abnormal image orientation. As a result, hybrid systems have been exhibited in order to enhance the accuracy with the maintenance of less training time. All models have been implemented in two groups of data sets: full dataset and sample dataset. Therefore, a comparative analysis has been developed in this paper. Some visualization of the attributes of the dataset has also been showed in this paper
Classification and Segmentation of Pulmonary Lesions in CT images using a combined VGG-XGBoost method, and an integrated Fuzzy Clustering-Level Set technique
Jan 04, 2021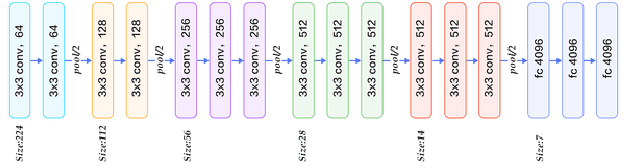
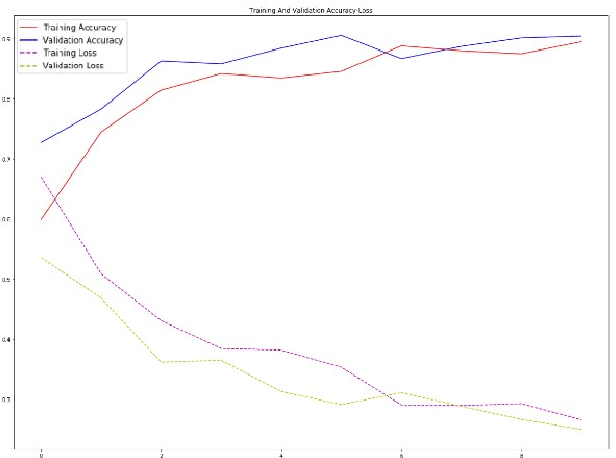
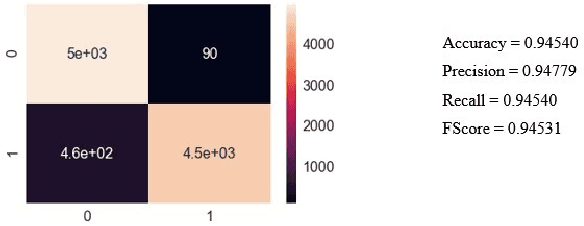
Given that lung cancer is one of the deadliest diseases, and many die from the disease every year, early detection and diagnosis of this disease are valuable, preventing cancer from growing and spreading. So if cancer is diagnosed in the early stage, the patient's life will be saved. However, the current pulmonary disease diagnosis is made by human resources, which is time-consuming and requires a specialist in this field. Also, there is a high level of errors in human diagnosis. Our goal is to develop a system that can detect and classify lung lesions with high accuracy and segment them in CT-scan images. In the proposed method, first, features are extracted automatically from the CT-scan image; then, the extracted features are classified by Ensemble Gradient Boosting methods. Finally, if there is a lesion in the CT-scan image, using a hybrid method based on [1], including Fuzzy Clustering and Level Set, the lesion is segmented. We collected a dataset, including CT-scan images of pulmonary lesions. The target community was the patients in Mashhad. The collected samples were then tagged by a specialist. We used this dataset for training and testing our models. Finally, we were able to achieve an accuracy of 96% for this dataset. This system can help physicians to diagnose pulmonary lesions and prevent possible mistakes.
Computed Tomography Image Enhancement using 3D Convolutional Neural Network
Jul 18, 2018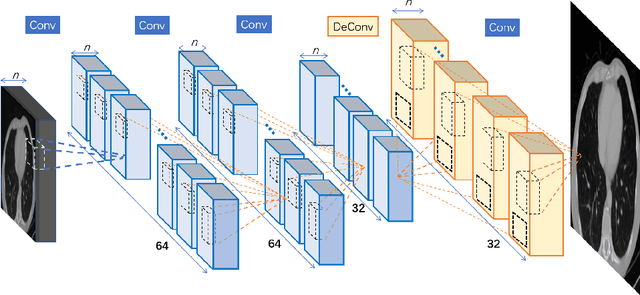
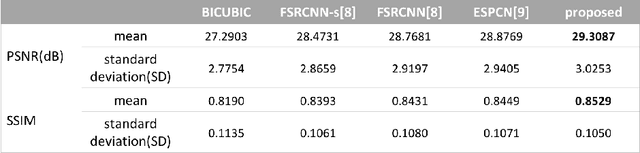
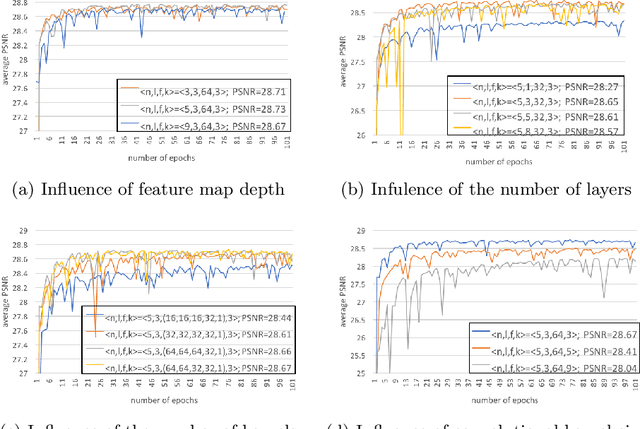
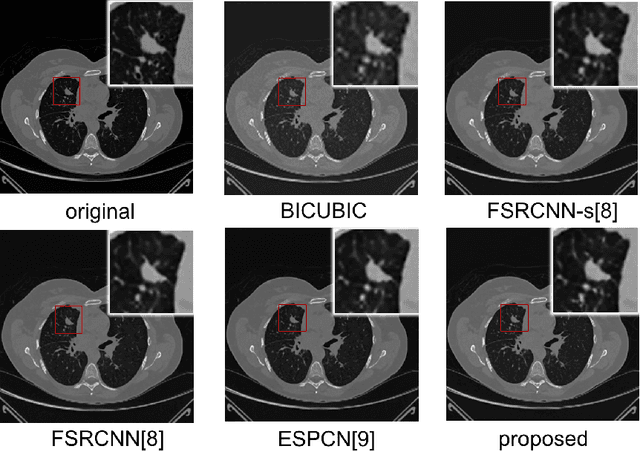
Computed tomography (CT) is increasingly being used for cancer screening, such as early detection of lung cancer. However, CT studies have varying pixel spacing due to differences in acquisition parameters. Thick slice CTs have lower resolution, hindering tasks such as nodule characterization during computer-aided detection due to partial volume effect. In this study, we propose a novel 3D enhancement convolutional neural network (3DECNN) to improve the spatial resolution of CT studies that were acquired using lower resolution/slice thicknesses to higher resolutions. Using a subset of the LIDC dataset consisting of 20,672 CT slices from 100 scans, we simulated lower resolution/thick section scans then attempted to reconstruct the original images using our 3DECNN network. A significant improvement in PSNR (29.3087dB vs. 28.8769dB, p-value < 2.2e-16) and SSIM (0.8529dB vs. 0.8449dB, p-value < 2.2e-16) compared to other state-of-art deep learning methods is observed.
LNDb: A Lung Nodule Database on Computed Tomography
Dec 19, 2019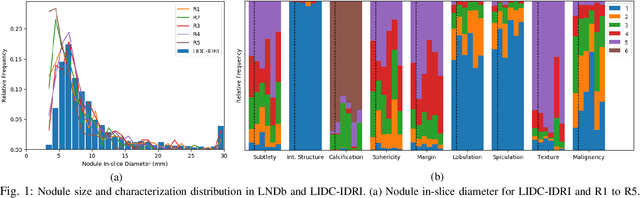
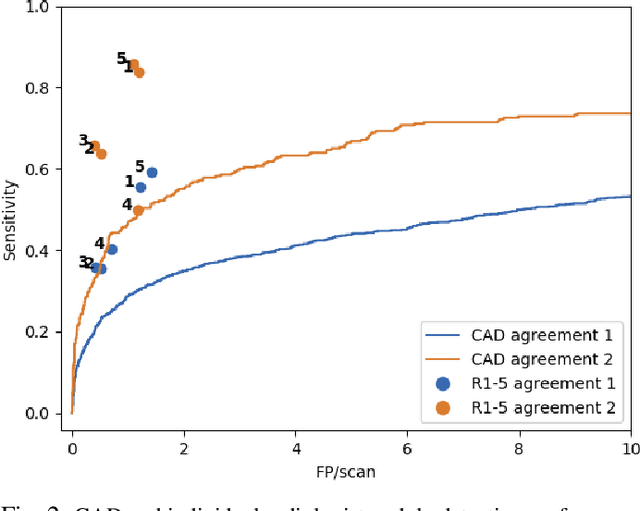
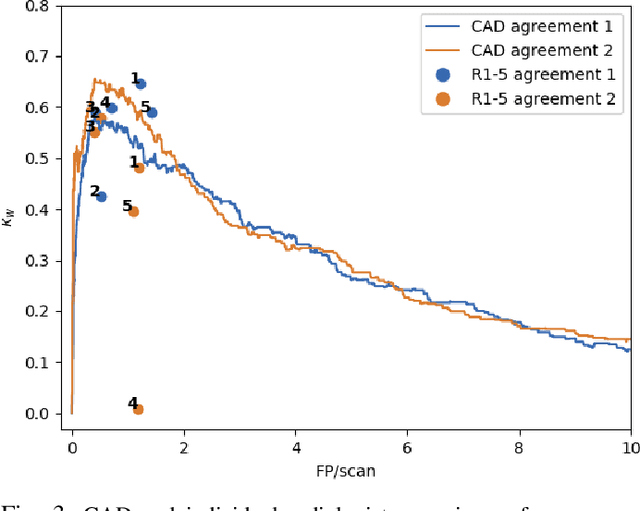
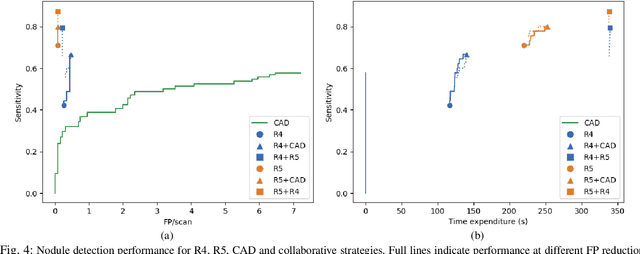
Lung cancer is the deadliest type of cancer worldwide and late detection is the major factor for the low survival rate of patients. Low dose computed tomography has been suggested as a potential screening tool but manual screening is costly, time-consuming and prone to variability. This has fueled the development of automatic methods for the detection, segmentation and characterisation of pulmonary nodules but its application to clinical routine is challenging. In this study, a new database for the development and testing of pulmonary nodule computer-aided strategies is presented which intends to complement current databases by giving additional focus to radiologist variability and local clinical reality. State-of-the-art nodule detection, segmentation and characterization methods are tested and compared to manual annotations as well as collaborative strategies combining multiple radiologists and radiologists and computer-aided systems. It is shown that state-of-the-art methodologies can determine a patient's follow-up recommendation as accurately as a radiologist, though the nodule detection method used shows decreased performance in this database.
RUN:Residual U-Net for Computer-Aided Detection of Pulmonary Nodules without Candidate Selection
May 30, 2018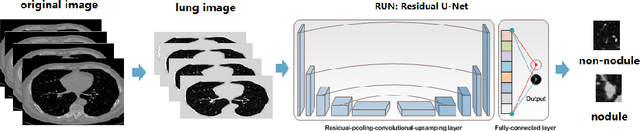
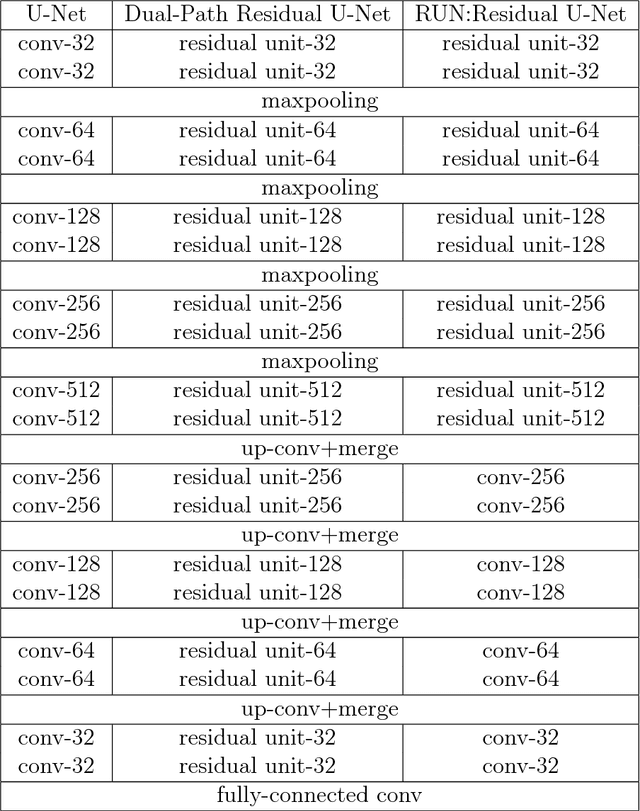
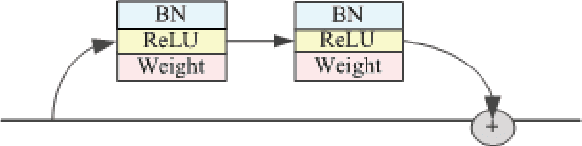
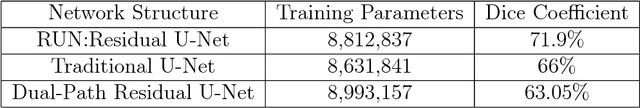
The early detection and early diagnosis of lung cancer are crucial to improve the survival rate of lung cancer patients. Pulmonary nodules detection results have a significant impact on the later diagnosis. In this work, we propose a new network named RUN to complete nodule detection in a single step by bypassing the candidate selection. The system introduces the shortcut of the residual network to improve the traditional U-Net, thereby solving the disadvantage of poor results due to its lack of depth. Furthermore, we compare the experimental results with the traditional U-Net. We validate our method in LUng Nodule Analysis 2016 (LUNA16) Nodule Detection Challenge. We acquire a sensitivity of 90.90% at 2 false positives per scan and therefore achieve better performance than the current state-of-the-art approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge