"cancer detection": models, code, and papers
Efficient Lung Cancer Image Classification and Segmentation Algorithm Based on Improved Swin Transformer
Jul 04, 2022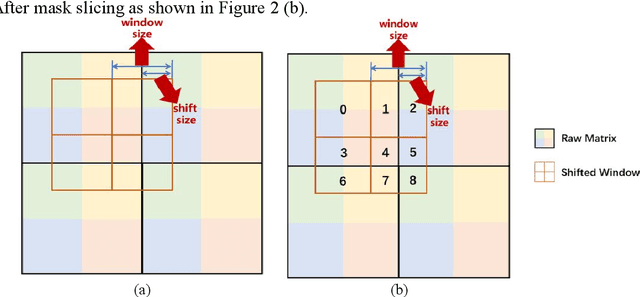
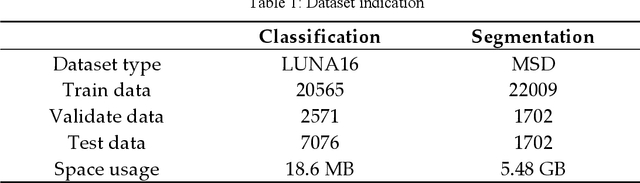
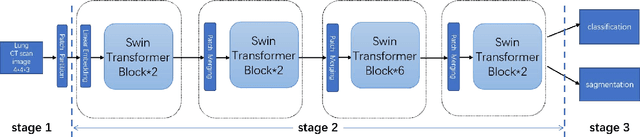
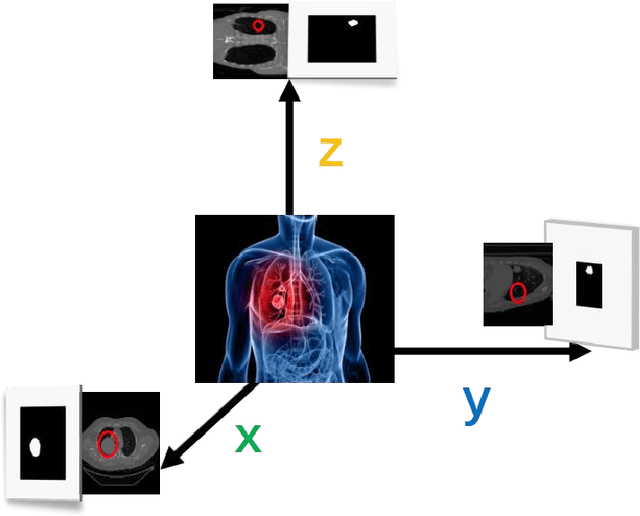
With the development of computer technology, various models have emerged in artificial intelligence. The transformer model has been applied to the field of computer vision (CV) after its success in natural language processing (NLP). Radiologists continue to face multiple challenges in today's rapidly evolving medical field, such as increased workload and increased diagnostic demands. Although there are some conventional methods for lung cancer detection before, their accuracy still needs to be improved, especially in realistic diagnostic scenarios. This paper creatively proposes a segmentation method based on efficient transformer and applies it to medical image analysis. The algorithm completes the task of lung cancer classification and segmentation by analyzing lung cancer data, and aims to provide efficient technical support for medical staff. In addition, we evaluated and compared the results in various aspects. For the classification mission, the max accuracy of Swin-T by regular training and Swin-B in two resolutions by pre-training can be up to 82.3%. For the segmentation mission, we use pre-training to help the model improve the accuracy of our experiments. The accuracy of the three models reaches over 95%. The experiments demonstrate that the algorithm can be well applied to lung cancer classification and segmentation missions.
Multi-Head Feature Pyramid Networks for Breast Mass Detection
Feb 22, 2023
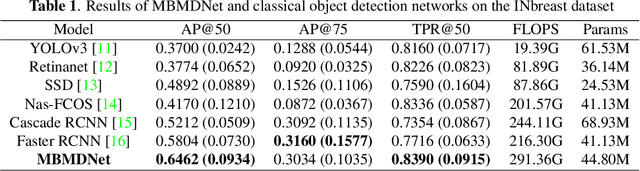
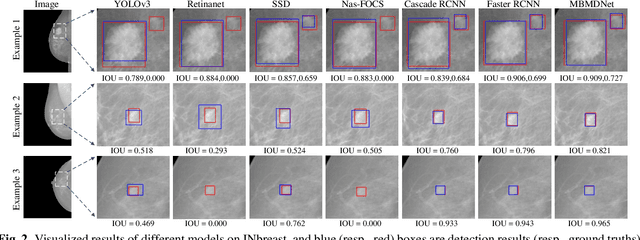
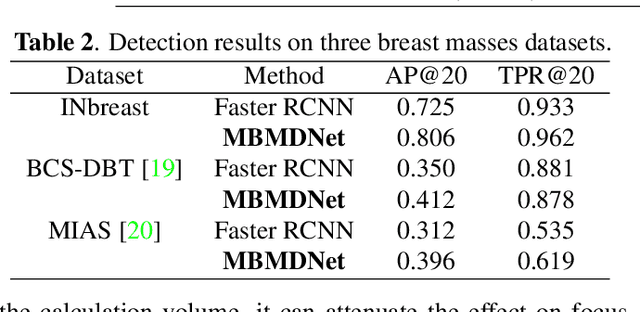
Analysis of X-ray images is one of the main tools to diagnose breast cancer. The ability to quickly and accurately detect the location of masses from the huge amount of image data is the key to reducing the morbidity and mortality of breast cancer. Currently, the main factor limiting the accuracy of breast mass detection is the unequal focus on the mass boxes, leading the network to focus too much on larger masses at the expense of smaller ones. In the paper, we propose the multi-head feature pyramid module (MHFPN) to solve the problem of unbalanced focus of target boxes during feature map fusion and design a multi-head breast mass detection network (MBMDnet). Experimental studies show that, comparing to the SOTA detection baselines, our method improves by 6.58% (in AP@50) and 5.4% (in TPR@50) on the commonly used INbreast dataset, while about 6-8% improvements (in AP@20) are also observed on the public MIAS and BCS-DBT datasets.
A deep learning algorithm for reducing false positives in screening mammography
Apr 13, 2022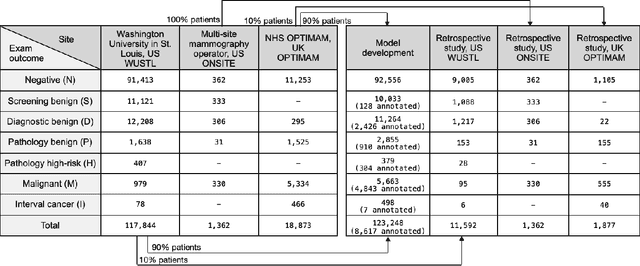
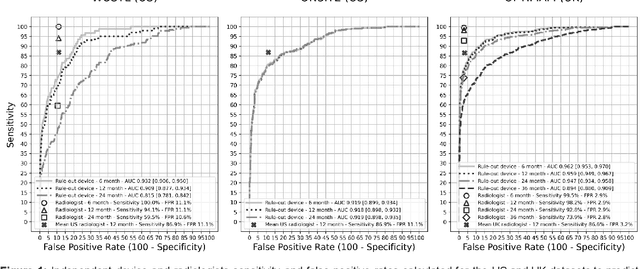
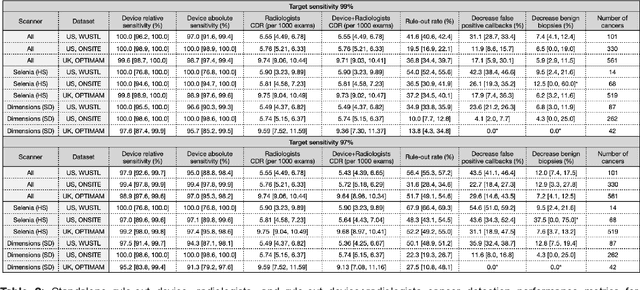
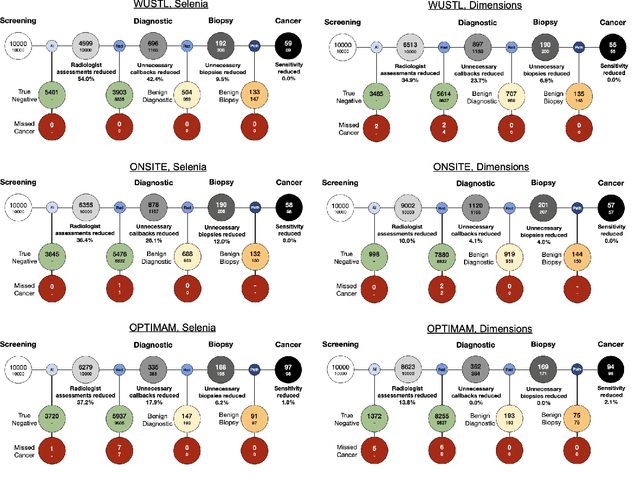
Screening mammography improves breast cancer outcomes by enabling early detection and treatment. However, false positive callbacks for additional imaging from screening exams cause unnecessary procedures, patient anxiety, and financial burden. This work demonstrates an AI algorithm that reduces false positives by identifying mammograms not suspicious for breast cancer. We trained the algorithm to determine the absence of cancer using 123,248 2D digital mammograms (6,161 cancers) and performed a retrospective study on 14,831 screening exams (1,026 cancers) from 15 US and 3 UK sites. Retrospective evaluation of the algorithm on the largest of the US sites (11,592 mammograms, 101 cancers) a) left the cancer detection rate unaffected (p=0.02, non-inferiority margin 0.25 cancers per 1000 exams), b) reduced callbacks for diagnostic exams by 31.1% compared to standard clinical readings, c) reduced benign needle biopsies by 7.4%, and d) reduced screening exams requiring radiologist interpretation by 41.6% in the simulated clinical workflow. This work lays the foundation for semi-autonomous breast cancer screening systems that could benefit patients and healthcare systems by reducing false positives, unnecessary procedures, patient anxiety, and expenses.
Machine Learning Approaches to Predict Breast Cancer: Bangladesh Perspective
Jun 30, 2022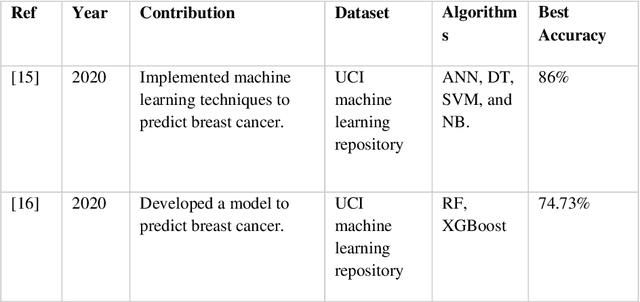
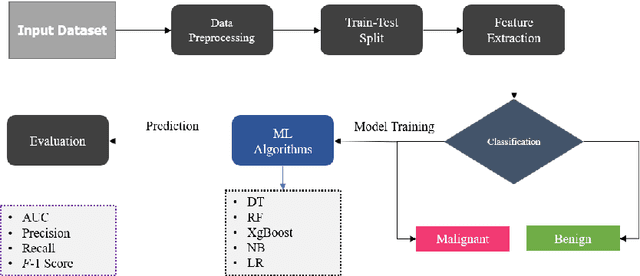
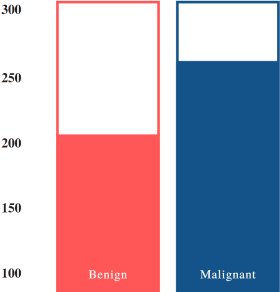
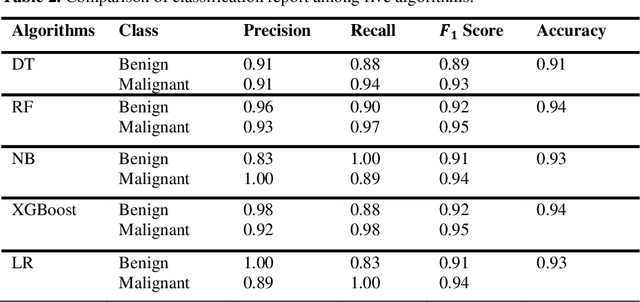
Nowadays, Breast cancer has risen to become one of the most prominent causes of death in recent years. Among all malignancies, this is the most frequent and the major cause of death for women globally. Manually diagnosing this disease requires a good amount of time and expertise. Breast cancer detection is time-consuming, and the spread of the disease can be reduced by developing machine-based breast cancer predictions. In Machine learning, the system can learn from prior instances and find hard-to-detect patterns from noisy or complicated data sets using various statistical, probabilistic, and optimization approaches. This work compares several machine learning algorithm's classification accuracy, precision, sensitivity, and specificity on a newly collected dataset. In this work Decision tree, Random Forest, Logistic Regression, Naive Bayes, and XGBoost, these five machine learning approaches have been implemented to get the best performance on our dataset. This study focuses on finding the best algorithm that can forecast breast cancer with maximum accuracy in terms of its classes. This work evaluated the quality of each algorithm's data classification in terms of efficiency and effectiveness. And also compared with other published work on this domain. After implementing the model, this study achieved the best model accuracy, 94% on Random Forest and XGBoost.
A Radiogenomics Pipeline for Lung Nodules Segmentation and Prediction of EGFR Mutation Status from CT Scans
Nov 12, 2022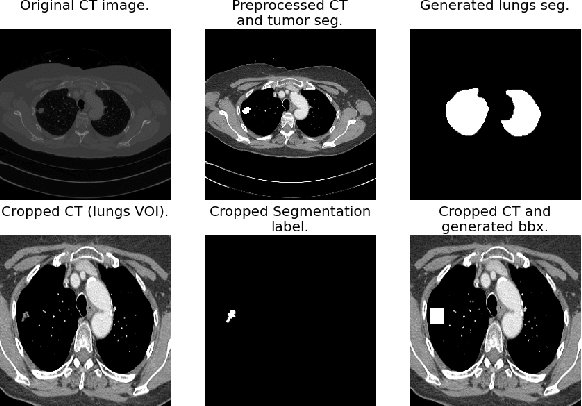
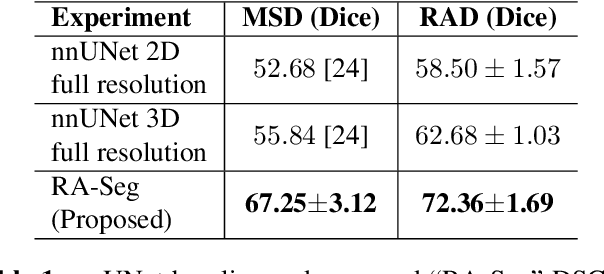
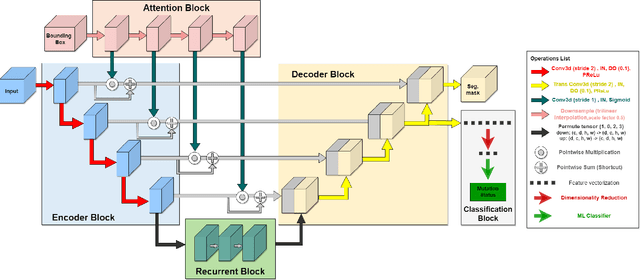
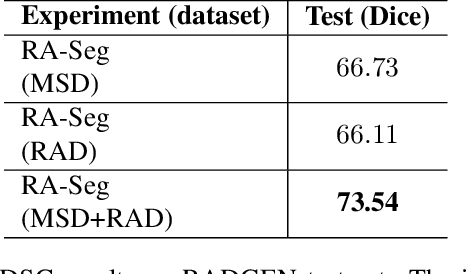
Lung cancer is a leading cause of death worldwide. Early-stage detection of lung cancer is essential for a more favorable prognosis. Radiogenomics is an emerging discipline that combines medical imaging and genomics features for modeling patient outcomes non-invasively. This study presents a radiogenomics pipeline that has: 1) a novel mixed architecture (RA-Seg) to segment lung cancer through attention and recurrent blocks; and 2) deep feature classifiers to distinguish Epidermal Growth Factor Receptor (EGFR) mutation status. We evaluate the proposed algorithm on multiple public datasets to assess its generalizability and robustness. We demonstrate how the proposed segmentation and classification methods outperform existing baseline and SOTA approaches (73.54 Dice and 93 F1 scores).
Detection and Localization of Melanoma Skin Cancer in Histopathological Whole Slide Images
Feb 06, 2023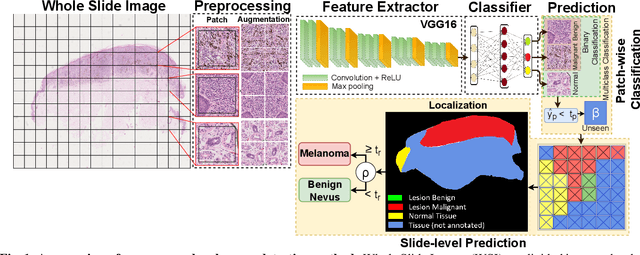
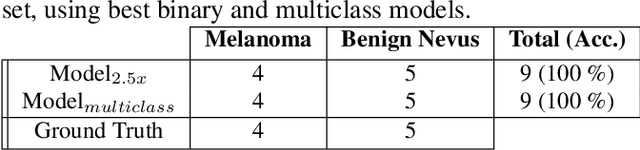
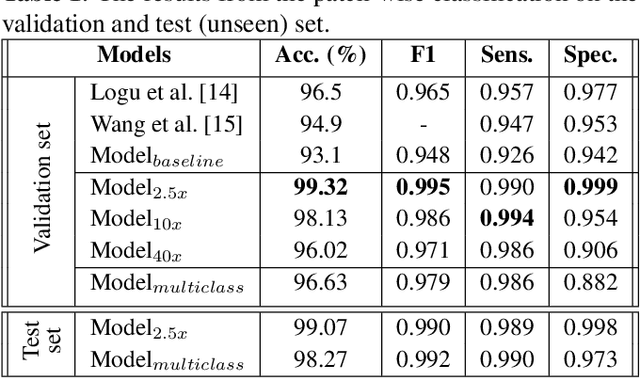
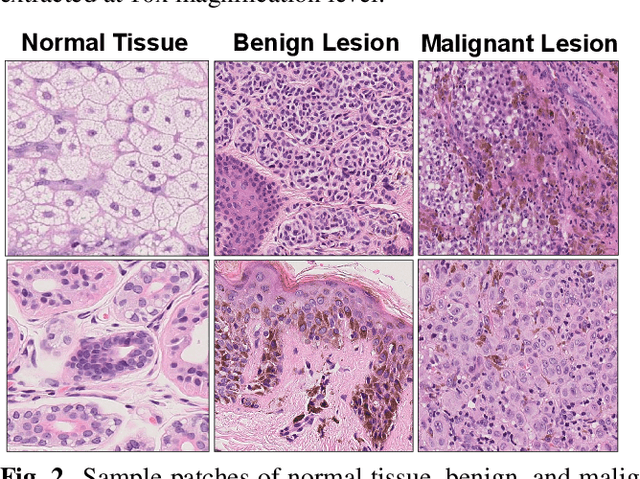

Melanoma diagnosed and treated in its early stages can increase the survival rate. A projected increase in skin cancer incidents and a dearth of dermatopathologists have emphasized the need for computational pathology (CPATH) systems. CPATH systems with deep learning (DL) models have the potential to identify the presence of melanoma by exploiting underlying morphological and cellular features. This paper proposes a DL method to detect melanoma and distinguish between normal skin and benign/malignant melanocytic lesions in Whole Slide Images (WSI). Our method detects lesions with high accuracy and localizes them on a WSI to identify potential regions of interest for pathologists. Interestingly, our DL method relies on using a single CNN network to create localization maps first and use them to perform slide-level predictions to determine patients who have melanoma. Our best model provides favorable patch-wise classification results with a 0.992 F1 score and 0.99 sensitivity on unseen data.
Robust breast cancer detection in mammography and digital breast tomosynthesis using annotation-efficient deep learning approach
Dec 27, 2019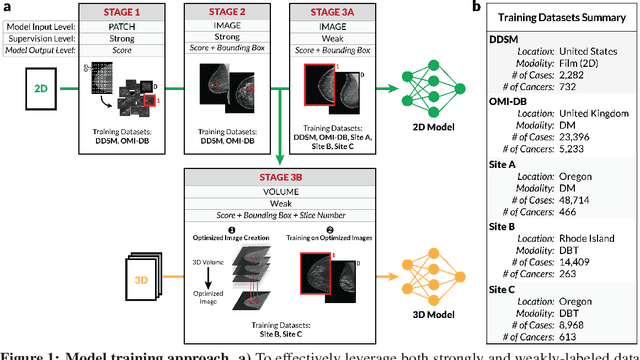
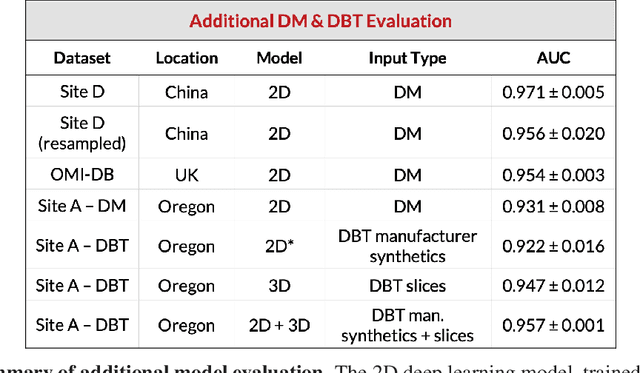
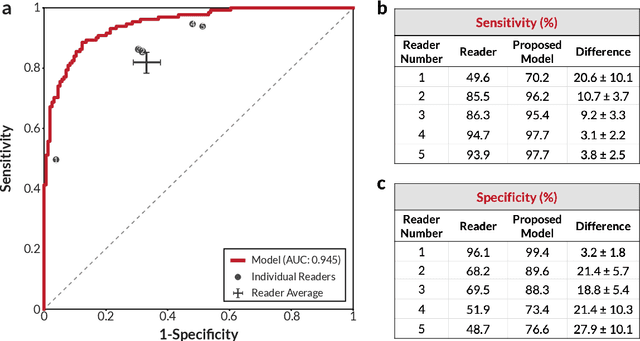
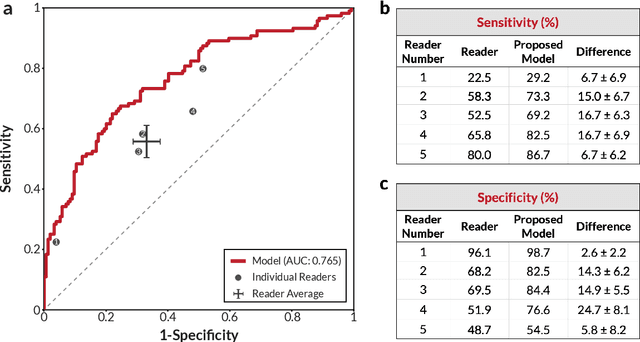
Breast cancer remains a global challenge, causing over 1 million deaths globally in 2018. To achieve earlier breast cancer detection, screening x-ray mammography is recommended by health organizations worldwide and has been estimated to decrease breast cancer mortality by 20-40%. Nevertheless, significant false positive and false negative rates, as well as high interpretation costs, leave opportunities for improving quality and access. To address these limitations, there has been much recent interest in applying deep learning to mammography; however, obtaining large amounts of annotated data poses a challenge for training deep learning models for this purpose, as does ensuring generalization beyond the populations represented in the training dataset. Here, we present an annotation-efficient deep learning approach that 1) achieves state-of-the-art performance in mammogram classification, 2) successfully extends to digital breast tomosynthesis (DBT; "3D mammography"), 3) detects cancers in clinically-negative prior mammograms of cancer patients, 4) generalizes well to a population with low screening rates, and 5) outperforms five-out-of-five full-time breast imaging specialists by improving absolute sensitivity by an average of 14%. Our results demonstrate promise towards software that can improve the accuracy of and access to screening mammography worldwide.
RUPNet: Residual upsampling network for real-time polyp segmentation
Jan 06, 2023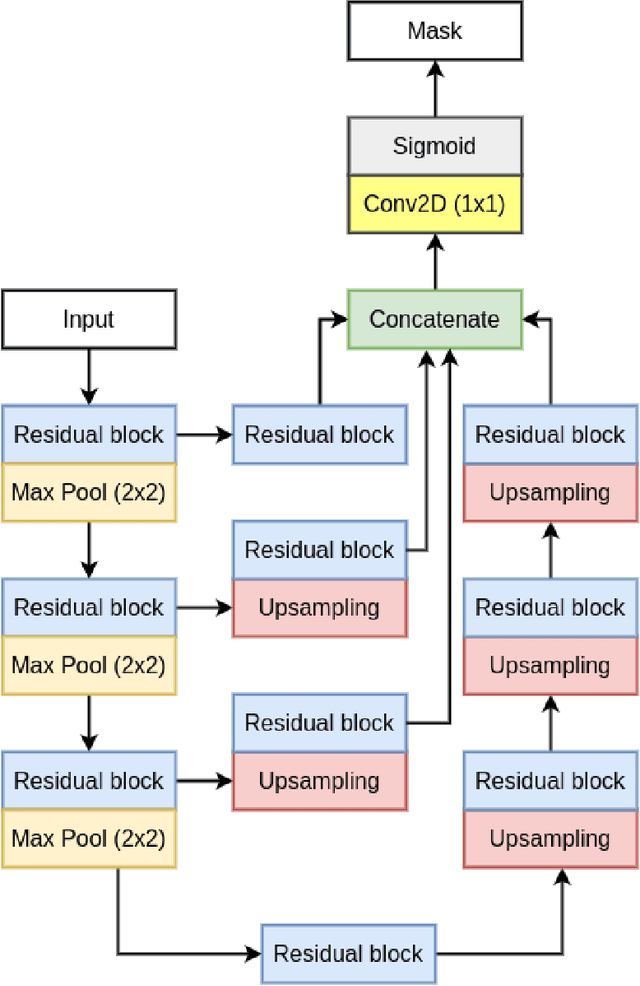
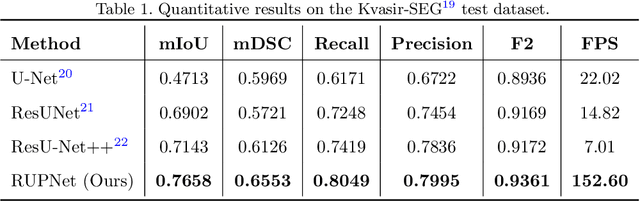
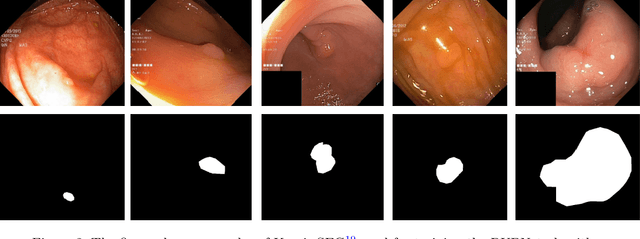
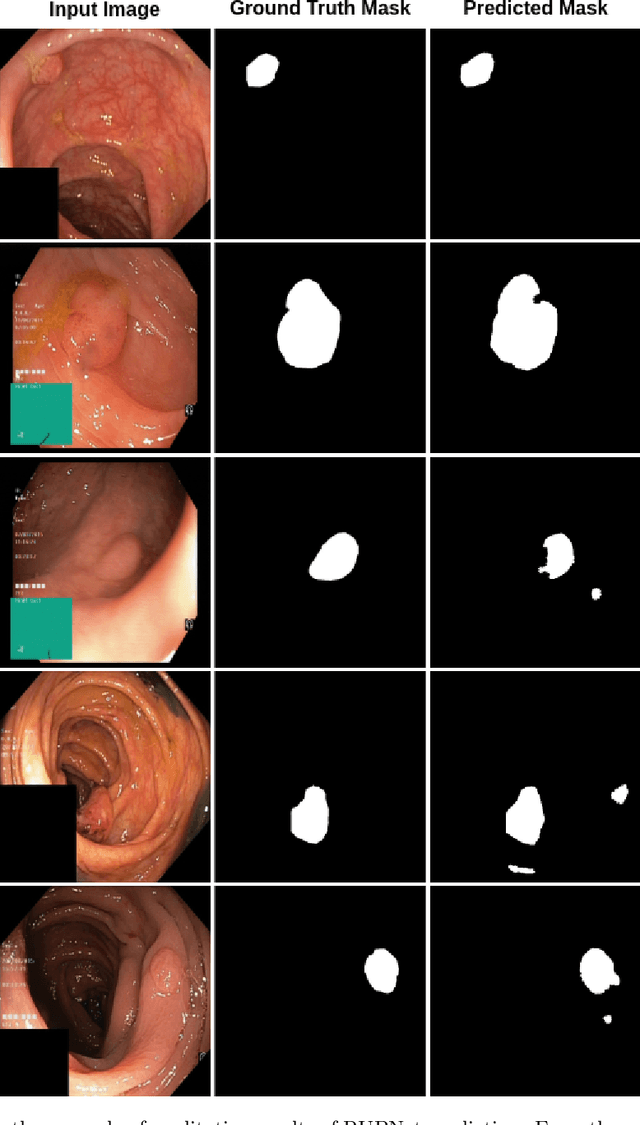
Colorectal cancer is among the most prevalent cause of cancer-related mortality worldwide. Detection and removal of polyps at an early stage can help reduce mortality and even help in spreading over adjacent organs. Early polyp detection could save the lives of millions of patients over the world as well as reduce the clinical burden. However, the detection polyp rate varies significantly among endoscopists. There is numerous deep learning-based method proposed, however, most of the studies improve accuracy. Here, we propose a novel architecture, Residual Upsampling Network (RUPNet) for colon polyp segmentation that can process in real-time and show high recall and precision. The proposed architecture, RUPNet, is an encoder-decoder network that consists of three encoders, three decoder blocks, and some additional upsampling blocks at the end of the network. With an image size of $512 \times 512$, the proposed method achieves an excellent real-time operation speed of 152.60 frames per second with an average dice coefficient of 0.7658, mean intersection of union of 0.6553, sensitivity of 0.8049, precision of 0.7995, and F2-score of 0.9361. The results suggest that RUPNet can give real-time feedback while retaining high accuracy indicating a good benchmark for early polyp detection.
Discovery Radiomics via StochasticNet Sequencers for Cancer Detection
Nov 11, 2015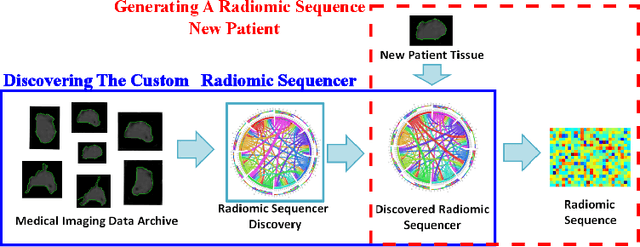
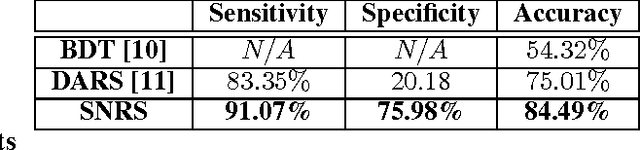
Radiomics has proven to be a powerful prognostic tool for cancer detection, and has previously been applied in lung, breast, prostate, and head-and-neck cancer studies with great success. However, these radiomics-driven methods rely on pre-defined, hand-crafted radiomic feature sets that can limit their ability to characterize unique cancer traits. In this study, we introduce a novel discovery radiomics framework where we directly discover custom radiomic features from the wealth of available medical imaging data. In particular, we leverage novel StochasticNet radiomic sequencers for extracting custom radiomic features tailored for characterizing unique cancer tissue phenotype. Using StochasticNet radiomic sequencers discovered using a wealth of lung CT data, we perform binary classification on 42,340 lung lesions obtained from the CT scans of 93 patients in the LIDC-IDRI dataset. Preliminary results show significant improvement over previous state-of-the-art methods, indicating the potential of the proposed discovery radiomics framework for improving cancer screening and diagnosis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge