"Information": models, code, and papers
Local-Global Information Interaction Debiasing for Dynamic Scene Graph Generation
Aug 10, 2023
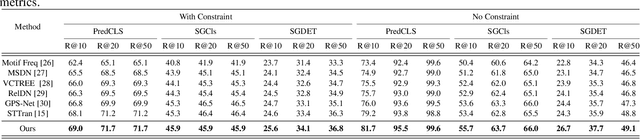
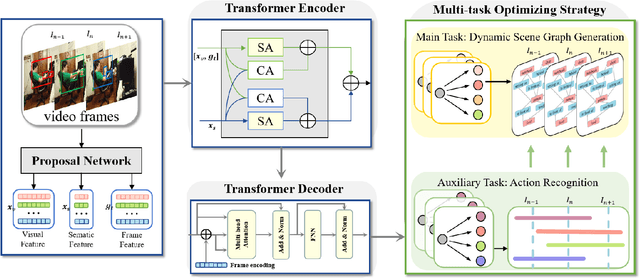
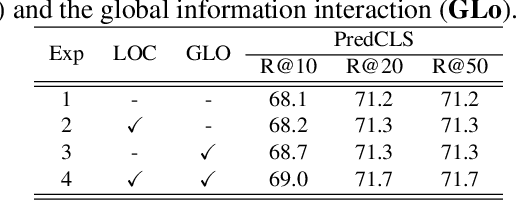
The task of dynamic scene graph generation (DynSGG) aims to generate scene graphs for given videos, which involves modeling the spatial-temporal information in the video. However, due to the long-tailed distribution of samples in the dataset, previous DynSGG models fail to predict the tail predicates. We argue that this phenomenon is due to previous methods that only pay attention to the local spatial-temporal information and neglect the consistency of multiple frames. To solve this problem, we propose a novel DynSGG model based on multi-task learning, DynSGG-MTL, which introduces the local interaction information and global human-action interaction information. The interaction between objects and frame features makes the model more fully understand the visual context of the single image. Long-temporal human actions supervise the model to generate multiple scene graphs that conform to the global constraints and avoid the model being unable to learn the tail predicates. Extensive experiments on Action Genome dataset demonstrate the efficacy of our proposed framework, which not only improves the dynamic scene graph generation but also alleviates the long-tail problem.
Augmenting End-to-End Steering Angle Prediction with CAN Bus Data
Oct 22, 2023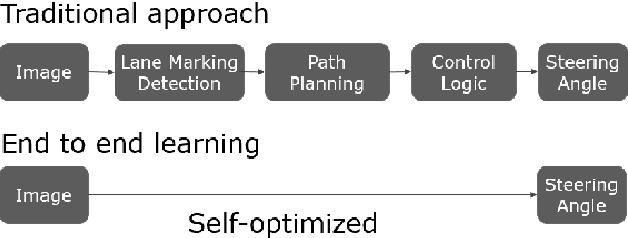
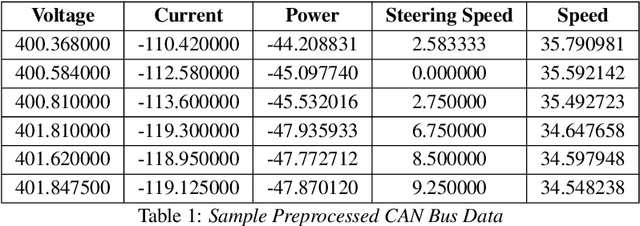
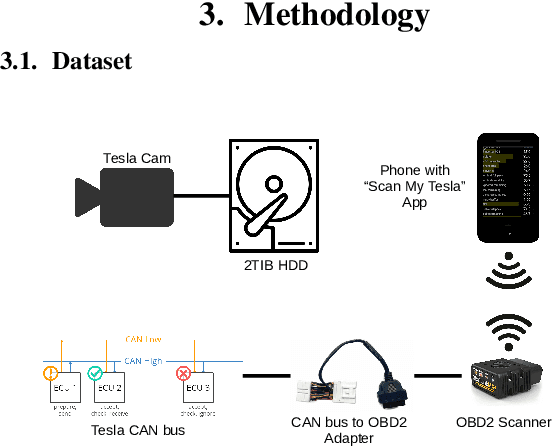
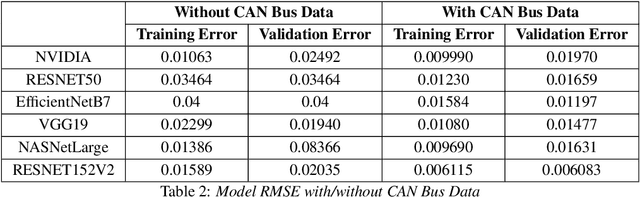
In recent years, end to end steering prediction for autonomous vehicles has become a major area of research. The primary method for achieving end to end steering was to use computer vision models on a live feed of video data. However, to further increase accuracy, many companies have added data from light detection and ranging (LiDAR) and or radar sensors through sensor fusion. However, the addition of lasers and sensors comes at a high financial cost. In this paper, I address both of these issues by increasing the accuracy of the computer vision models without the increased cost of using LiDAR and or sensors. I achieved this by improving the accuracy of computer vision models by sensor fusing CAN bus data, a vehicle protocol, with video data. CAN bus data is a rich source of information about the vehicle's state, including its speed, steering angle, and acceleration. By fusing this data with video data, the accuracy of the computer vision model's predictions can be improved. When I trained the model without CAN bus data, I obtained an RMSE of 0.02492, while the model trained with the CAN bus data achieved an RMSE of 0.01970. This finding indicates that fusing CAN Bus data with video data can reduce the computer vision model's prediction error by 20% with some models decreasing the error by 80%.
Visual-Attribute Prompt Learning for Progressive Mild Cognitive Impairment Prediction
Oct 22, 2023Deep learning (DL) has been used in the automatic diagnosis of Mild Cognitive Impairment (MCI) and Alzheimer's Disease (AD) with brain imaging data. However, previous methods have not fully exploited the relation between brain image and clinical information that is widely adopted by experts in practice. To exploit the heterogeneous features from imaging and tabular data simultaneously, we propose the Visual-Attribute Prompt Learning-based Transformer (VAP-Former), a transformer-based network that efficiently extracts and fuses the multi-modal features with prompt fine-tuning. Furthermore, we propose a Prompt fine-Tuning (PT) scheme to transfer the knowledge from AD prediction task for progressive MCI (pMCI) diagnosis. In details, we first pre-train the VAP-Former without prompts on the AD diagnosis task and then fine-tune the model on the pMCI detection task with PT, which only needs to optimize a small amount of parameters while keeping the backbone frozen. Next, we propose a novel global prompt token for the visual prompts to provide global guidance to the multi-modal representations. Extensive experiments not only show the superiority of our method compared with the state-of-the-art methods in pMCI prediction but also demonstrate that the global prompt can make the prompt learning process more effective and stable. Interestingly, the proposed prompt learning model even outperforms the fully fine-tuning baseline on transferring the knowledge from AD to pMCI.
$Λ$-Split: A Privacy-Preserving Split Computing Framework for Cloud-Powered Generative AI
Oct 23, 2023In the wake of the burgeoning expansion of generative artificial intelligence (AI) services, the computational demands inherent to these technologies frequently necessitate cloud-powered computational offloading, particularly for resource-constrained mobile devices. These services commonly employ prompts to steer the generative process, and both the prompts and the resultant content, such as text and images, may harbor privacy-sensitive or confidential information, thereby elevating security and privacy risks. To mitigate these concerns, we introduce $\Lambda$-Split, a split computing framework to facilitate computational offloading while simultaneously fortifying data privacy against risks such as eavesdropping and unauthorized access. In $\Lambda$-Split, a generative model, usually a deep neural network (DNN), is partitioned into three sub-models and distributed across the user's local device and a cloud server: the input-side and output-side sub-models are allocated to the local, while the intermediate, computationally-intensive sub-model resides on the cloud server. This architecture ensures that only the hidden layer outputs are transmitted, thereby preventing the external transmission of privacy-sensitive raw input and output data. Given the black-box nature of DNNs, estimating the original input or output from intercepted hidden layer outputs poses a significant challenge for malicious eavesdroppers. Moreover, $\Lambda$-Split is orthogonal to traditional encryption-based security mechanisms, offering enhanced security when deployed in conjunction. We empirically validate the efficacy of the $\Lambda$-Split framework using Llama 2 and Stable Diffusion XL, representative large language and diffusion models developed by Meta and Stability AI, respectively. Our $\Lambda$-Split implementation is publicly accessible at https://github.com/nishio-laboratory/lambda_split.
One-dimensional convolutional neural network model for breast cancer subtypes classification and biochemical content evaluation using micro-FTIR hyperspectral images
Oct 23, 2023Breast cancer treatment still remains a challenge, where molecular subtypes classification plays a crucial role in selecting appropriate and specific therapy. The four subtypes are Luminal A (LA), Luminal B (LB), HER2 subtype, and Triple-Negative Breast Cancer (TNBC). Immunohistochemistry is the gold-standard evaluation, although interobserver variations are reported and molecular signatures identification is time-consuming. Fourier transform infrared micro-spectroscopy with machine learning approaches have been used to evaluate cancer samples, presenting biochemical-related explainability. However, this explainability is harder when using deep learning. This study created a 1D deep learning tool for breast cancer subtype evaluation and biochemical contribution. Sixty hyperspectral images were acquired from a human breast cancer microarray. K-Means clustering was applied to select tissue and paraffin spectra. CaReNet-V1, a novel 1D convolutional neural network, was developed to classify breast cancer (CA) and adjacent tissue (AT), and molecular subtypes. A 1D adaptation of Grad-CAM was applied to assess the biochemical impact to the classifications. CaReNet-V1 effectively classified CA and AT (test accuracy of 0.89), as well as HER2 and TNBC subtypes (0.83 and 0.86), with greater difficulty for LA and LB (0.74 and 0.68). The model enabled the evaluation of the most contributing wavenumbers to the predictions, providing a direct relationship with the biochemical content. Therefore, CaReNet-V1 and hyperspectral images is a potential approach for breast cancer biopsies assessment, providing additional information to the pathology report. Biochemical content impact feature may be used for other studies, such as treatment efficacy evaluation and development new diagnostics and therapeutic methods.
Deep Autoencoder-based Z-Interference Channels with Perfect and Imperfect CSI
Oct 23, 2023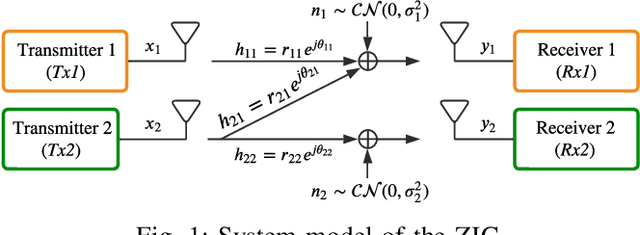
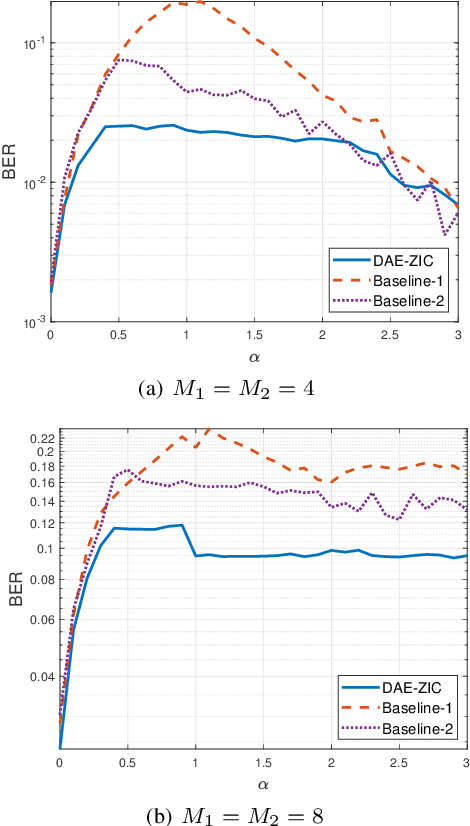
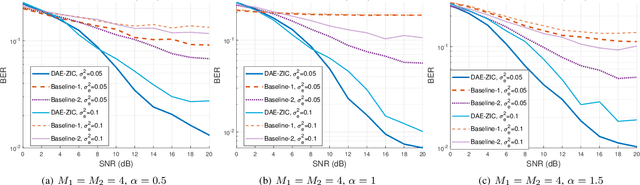
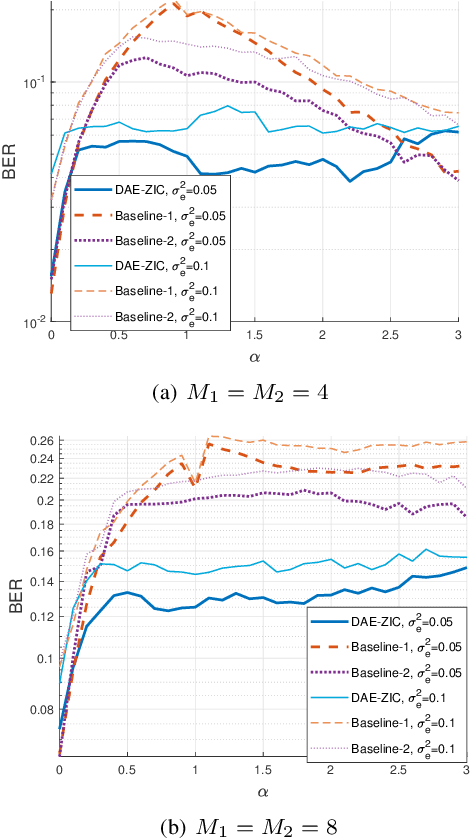
A deep autoencoder (DAE)-based structure for endto-end communication over the two-user Z-interference channel (ZIC) with finite-alphabet inputs is designed in this paper. The proposed structure jointly optimizes the two encoder/decoder pairs and generates interference-aware constellations that dynamically adapt their shape based on interference intensity to minimize the bit error rate (BER). An in-phase/quadrature-phase (I/Q) power allocation layer is introduced in the DAE to guarantee an average power constraint and enable the architecture to generate constellations with nonuniform shapes. This brings further gain compared to standard uniform constellations such as quadrature amplitude modulation. The proposed structure is then extended to work with imperfect channel state information (CSI). The CSI imperfection due to both the estimation and quantization errors are examined. The performance of the DAEZIC is compared with two baseline methods, i.e., standard and rotated constellations. The proposed structure significantly enhances the performance of the ZIC both for the perfect and imperfect CSI. Simulation results show that the improvement is achieved in all interference regimes (weak, moderate, and strong) and consistently increases with the signal-to-noise ratio (SNR). For example, more than an order of magnitude BER reduction is obtained with respect to the most competitive conventional method at weak interference when SNR>15dB and two bits per symbol are transmitted. The improvements reach about two orders of magnitude when quantization error exists, indicating that the DAE-ZIC is more robust to the interference compared to the conventional methods.
Calibration of Time-Series Forecasting Transformers: Detecting and Adapting Context-Driven Distribution Shift
Oct 23, 2023Recent years have witnessed the success of introducing Transformers to time series forecasting. From a data generation perspective, we illustrate that existing Transformers are susceptible to distribution shifts driven by temporal contexts, whether observed or unobserved. Such context-driven distribution shift (CDS) introduces biases in predictions within specific contexts and poses challenges for conventional training paradigm. In this paper, we introduce a universal calibration methodology for the detection and adaptation of CDS with a trained Transformer model. To this end, we propose a novel CDS detector, termed the "residual-based CDS detector" or "Reconditionor", which quantifies the model's vulnerability to CDS by evaluating the mutual information between prediction residuals and their corresponding contexts. A high Reconditionor score indicates a severe susceptibility, thereby necessitating model adaptation. In this circumstance, we put forth a straightforward yet potent adapter framework for model calibration, termed the "sample-level contextualized adapter" or "SOLID". This framework involves the curation of a contextually similar dataset to the provided test sample and the subsequent fine-tuning of the model's prediction layer with a limited number of steps. Our theoretical analysis demonstrates that this adaptation strategy is able to achieve an optimal equilibrium between bias and variance. Notably, our proposed Reconditionor and SOLID are model-agnostic and readily adaptable to a wide range of Transformers. Extensive experiments show that SOLID consistently enhances the performance of current SOTA Transformers on real-world datasets, especially on cases with substantial CDS detected by the proposed Reconditionor, thus validate the effectiveness of the calibration approach.
The Safety Challenges of Deep Learning in Real-World Type 1 Diabetes Management
Oct 23, 2023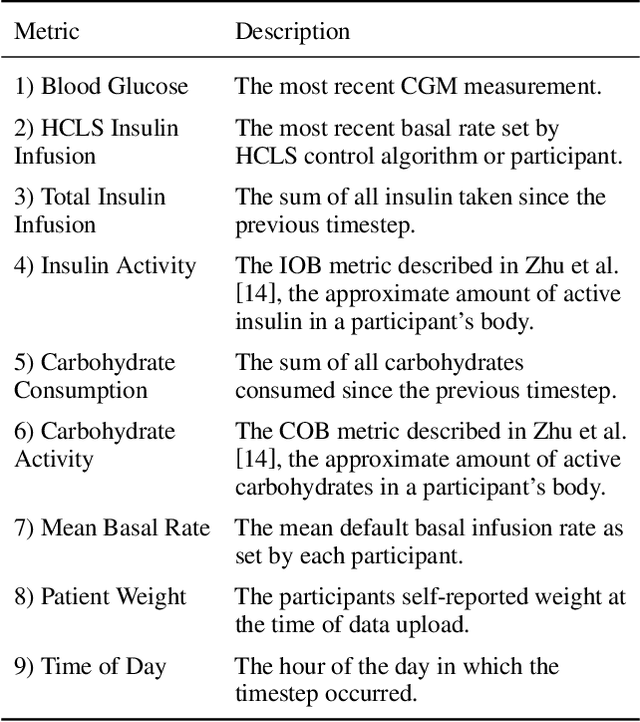
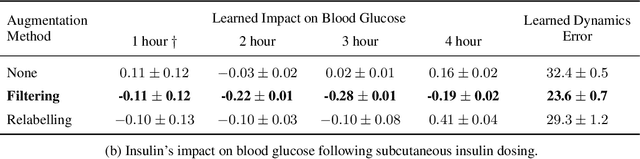
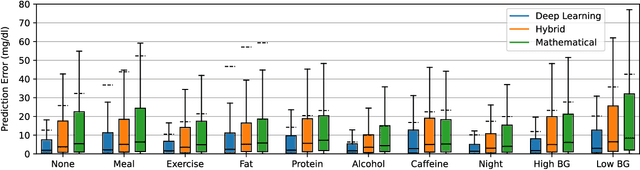
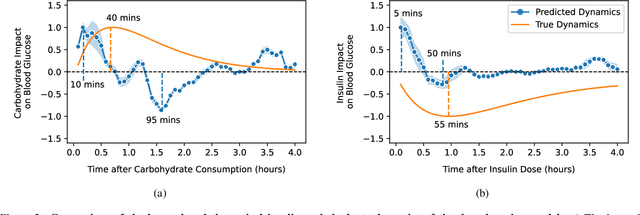

Blood glucose simulation allows the effectiveness of type 1 diabetes (T1D) management strategies to be evaluated without patient harm. Deep learning algorithms provide a promising avenue for extending simulator capabilities; however, these algorithms are limited in that they do not necessarily learn physiologically correct glucose dynamics and can learn incorrect and potentially dangerous relationships from confounders in training data. This is likely to be more important in real-world scenarios, as data is not collected under strict research protocol. This work explores the implications of using deep learning algorithms trained on real-world data to model glucose dynamics. Free-living data was processed from the OpenAPS Data Commons and supplemented with patient-reported tags of challenging diabetes events, constituting one of the most detailed real-world T1D datasets. This dataset was used to train and evaluate state-of-the-art glucose simulators, comparing their prediction error across safety critical scenarios and assessing the physiological appropriateness of the learned dynamics using Shapley Additive Explanations (SHAP). While deep learning prediction accuracy surpassed the widely-used mathematical simulator approach, the model deteriorated in safety critical scenarios and struggled to leverage self-reported meal and exercise information. SHAP value analysis also indicated the model had fundamentally confused the roles of insulin and carbohydrates, which is one of the most basic T1D management principles. This work highlights the importance of considering physiological appropriateness when using deep learning to model real-world systems in T1D and healthcare more broadly, and provides recommendations for building models that are robust to real-world data constraints.
Serverless Federated Learning with flwr-serverless
Oct 23, 2023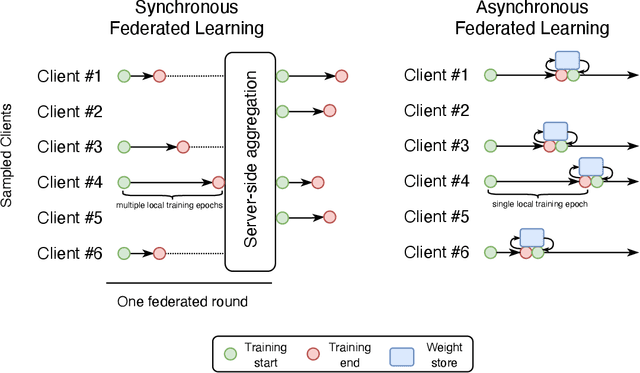
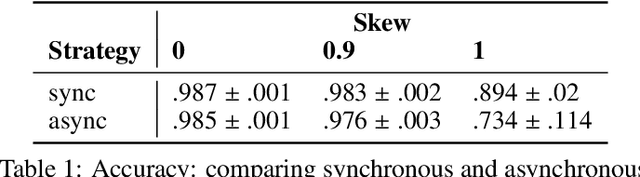
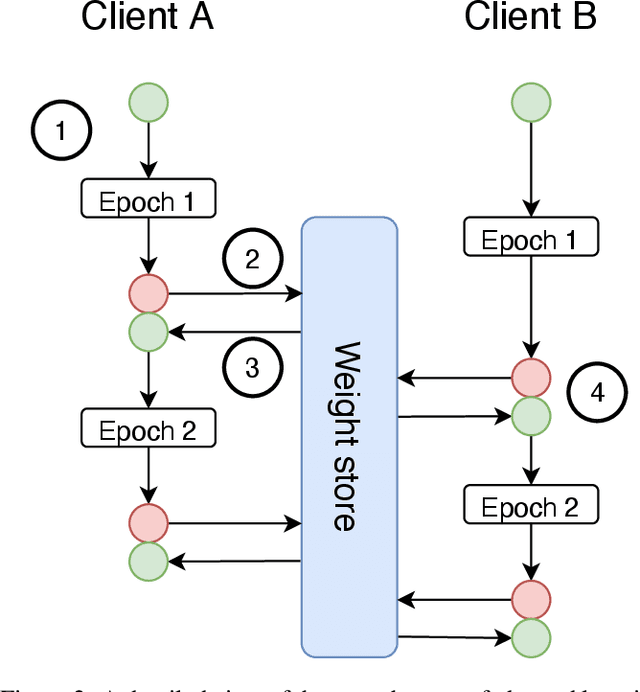
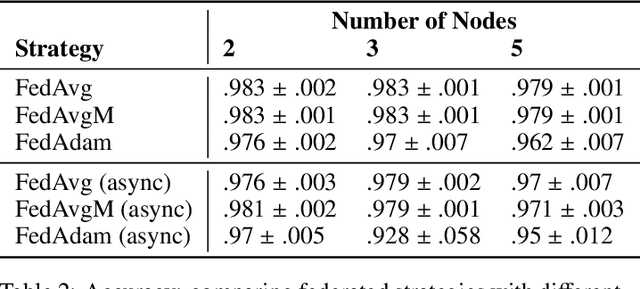
Federated learning is becoming increasingly relevant and popular as we witness a surge in data collection and storage of personally identifiable information. Alongside these developments there have been many proposals from governments around the world to provide more protections for individuals' data and a heightened interest in data privacy measures. As deep learning continues to become more relevant in new and existing domains, it is vital to develop strategies like federated learning that can effectively train data from different sources, such as edge devices, without compromising security and privacy. Recently, the Flower (\texttt{Flwr}) Python package was introduced to provide a scalable, flexible, and easy-to-use framework for implementing federated learning. However, to date, Flower is only able to run synchronous federated learning which can be costly and time-consuming to run because the process is bottlenecked by client-side training jobs that are slow or fragile. Here, we introduce \texttt{flwr-serverless}, a wrapper around the Flower package that extends its functionality to allow for both synchronous and asynchronous federated learning with minimal modification to Flower's design paradigm. Furthermore, our approach to federated learning allows the process to run without a central server, which increases the domains of application and accessibility of its use. This paper presents the design details and usage of this approach through a series of experiments that were conducted using public datasets. Overall, we believe that our approach decreases the time and cost to run federated training and provides an easier way to implement and experiment with federated learning systems.
Modeling Path Importance for Effective Alzheimer's Disease Drug Repurposing
Oct 23, 2023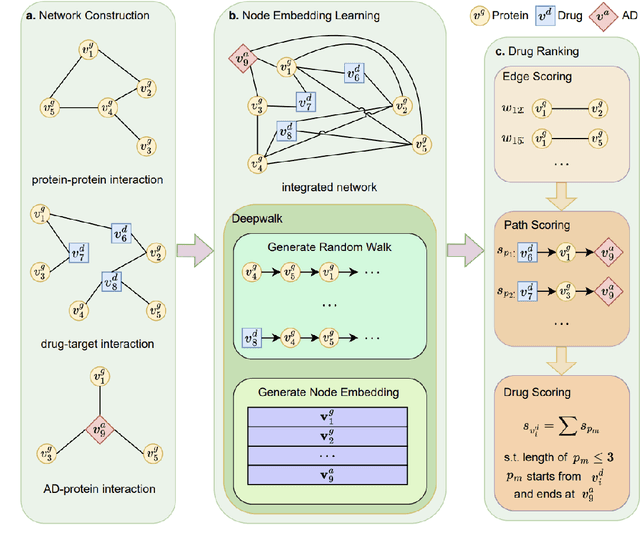
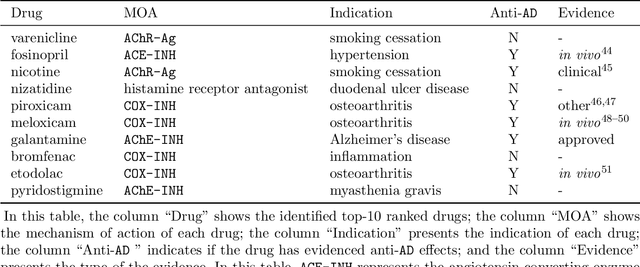
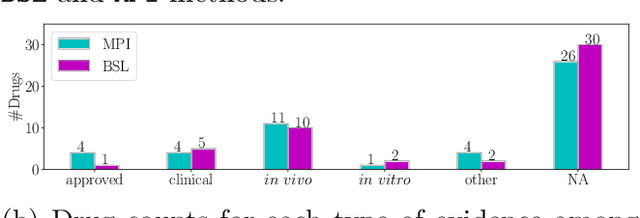

Recently, drug repurposing has emerged as an effective and resource-efficient paradigm for AD drug discovery. Among various methods for drug repurposing, network-based methods have shown promising results as they are capable of leveraging complex networks that integrate multiple interaction types, such as protein-protein interactions, to more effectively identify candidate drugs. However, existing approaches typically assume paths of the same length in the network have equal importance in identifying the therapeutic effect of drugs. Other domains have found that same length paths do not necessarily have the same importance. Thus, relying on this assumption may be deleterious to drug repurposing attempts. In this work, we propose MPI (Modeling Path Importance), a novel network-based method for AD drug repurposing. MPI is unique in that it prioritizes important paths via learned node embeddings, which can effectively capture a network's rich structural information. Thus, leveraging learned embeddings allows MPI to effectively differentiate the importance among paths. We evaluate MPI against a commonly used baseline method that identifies anti-AD drug candidates primarily based on the shortest paths between drugs and AD in the network. We observe that among the top-50 ranked drugs, MPI prioritizes 20.0% more drugs with anti-AD evidence compared to the baseline. Finally, Cox proportional-hazard models produced from insurance claims data aid us in identifying the use of etodolac, nicotine, and BBB-crossing ACE-INHs as having a reduced risk of AD, suggesting such drugs may be viable candidates for repurposing and should be explored further in future studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge