"Image": models, code, and papers
CLANet: A Comprehensive Framework for Cross-Batch Cell Line Identification Using Brightfield Images
Jun 28, 2023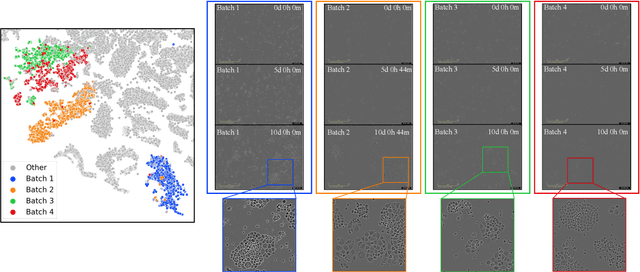

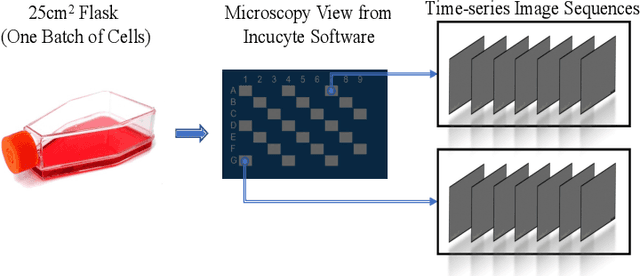

Cell line authentication plays a crucial role in the biomedical field, ensuring researchers work with accurately identified cells. Supervised deep learning has made remarkable strides in cell line identification by studying cell morphological features through cell imaging. However, batch effects, a significant issue stemming from the different times at which data is generated, lead to substantial shifts in the underlying data distribution, thus complicating reliable differentiation between cell lines from distinct batch cultures. To address this challenge, we introduce CLANet, a pioneering framework for cross-batch cell line identification using brightfield images, specifically designed to tackle three distinct batch effects. We propose a cell cluster-level selection method to efficiently capture cell density variations, and a self-supervised learning strategy to manage image quality variations, thus producing reliable patch representations. Additionally, we adopt multiple instance learning(MIL) for effective aggregation of instance-level features for cell line identification. Our innovative time-series segment sampling module further enhances MIL's feature-learning capabilities, mitigating biases from varying incubation times across batches. We validate CLANet using data from 32 cell lines across 93 experimental batches from the AstraZeneca Global Cell Bank. Our results show that CLANet outperforms related approaches (e.g. domain adaptation, MIL), demonstrating its effectiveness in addressing batch effects in cell line identification.
Continuous Cost Aggregation for Dual-Pixel Disparity Extraction
Jun 13, 2023
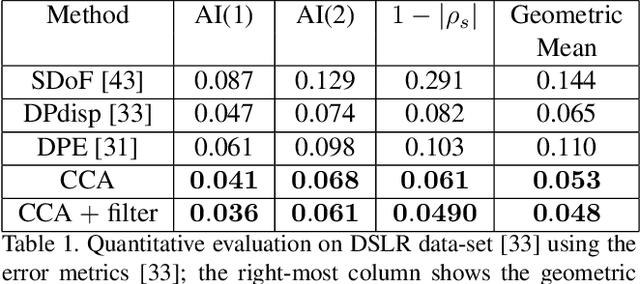
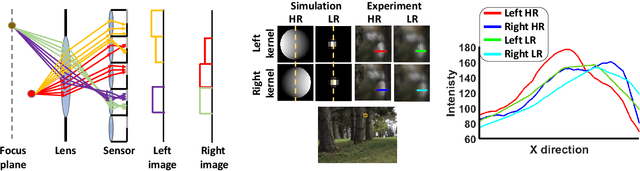
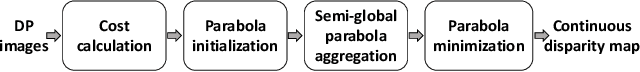
Recent works have shown that depth information can be obtained from Dual-Pixel (DP) sensors. A DP arrangement provides two views in a single shot, thus resembling a stereo image pair with a tiny baseline. However, the different point spread function (PSF) per view, as well as the small disparity range, makes the use of typical stereo matching algorithms problematic. To address the above shortcomings, we propose a Continuous Cost Aggregation (CCA) scheme within a semi-global matching framework that is able to provide accurate continuous disparities from DP images. The proposed algorithm fits parabolas to matching costs and aggregates parabola coefficients along image paths. The aggregation step is performed subject to a quadratic constraint that not only enforces the disparity smoothness but also maintains the quadratic form of the total costs. This gives rise to an inherently efficient disparity propagation scheme with a pixel-wise minimization in closed-form. Furthermore, the continuous form allows for a robust multi-scale aggregation that better compensates for the varying PSF. Experiments on DP data from both DSLR and phone cameras show that the proposed scheme attains state-of-the-art performance in DP disparity estimation.
TinySiamese Network for Biometric Analysis
Jul 02, 2023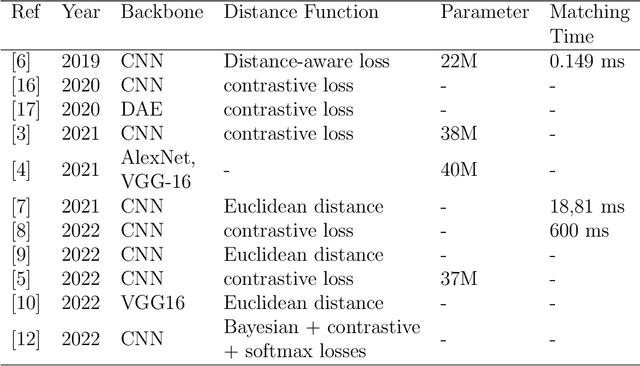
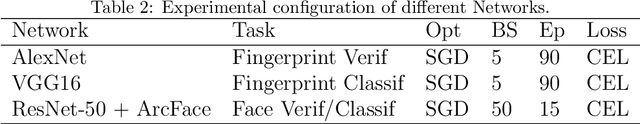
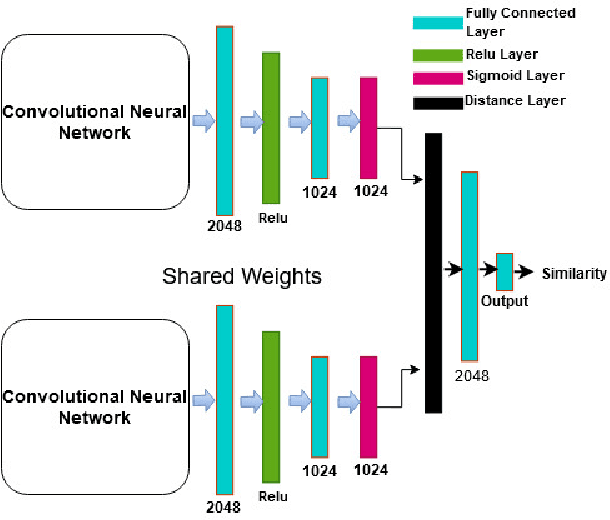
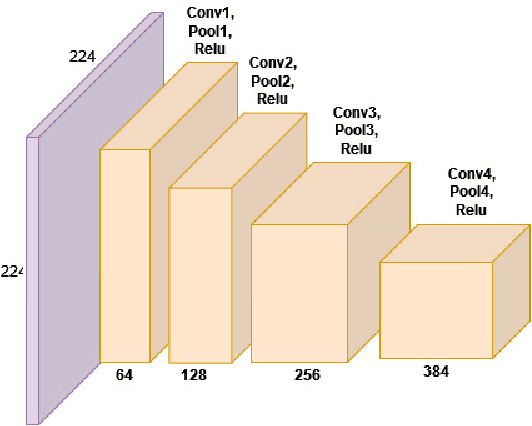

Biometric recognition is the process of verifying or classifying human characteristics in images or videos. It is a complex task that requires machine learning algorithms, including convolutional neural networks (CNNs) and Siamese networks. Besides, there are several limitations to consider when using these algorithms for image verification and classification tasks. In fact, training may be computationally intensive, requiring specialized hardware and significant computational resources to train and deploy. Moreover, it necessitates a large amount of labeled data, which can be time-consuming and costly to obtain. The main advantage of the proposed TinySiamese compared to the standard Siamese is that it does not require the whole CNN for training. In fact, using a pre-trained CNN as a feature extractor and the TinySiamese to learn the extracted features gave almost the same performance and efficiency as the standard Siamese for biometric verification. In this way, the TinySiamese solves the problems of memory and computational time with a small number of layers which did not exceed 7. It can be run under low-power machines which possess a normal GPU and cannot allocate a large RAM space. Using TinySiamese with only 8 GO of memory, the matching time decreased by 76.78% on the B2F (Biometric images of Fingerprints and Faces), FVC2000, FVC2002 and FVC2004 while the training time for 10 epochs went down by approximately 93.14% on the B2F, FVC2002, THDD-part1 and CASIA-B datasets. The accuracy of the fingerprint, gait (NM-angle 180 degree) and face verification tasks was better than the accuracy of a standard Siamese by 0.87%, 20.24% and 3.85% respectively. TinySiamese achieved comparable accuracy with related works for the fingerprint and gait classification tasks.
L1BSR: Exploiting Detector Overlap for Self-Supervised Single-Image Super-Resolution of Sentinel-2 L1B Imagery
Apr 17, 2023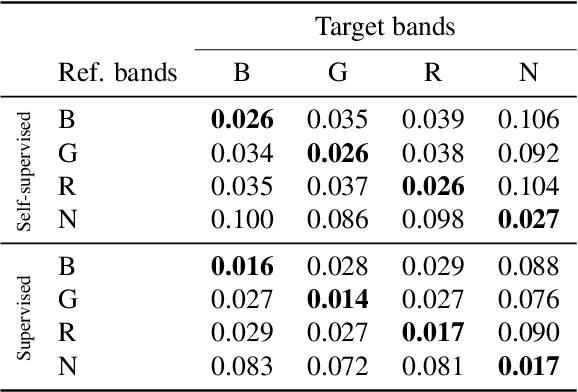
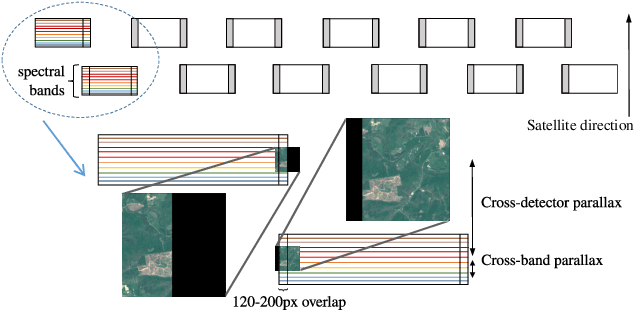
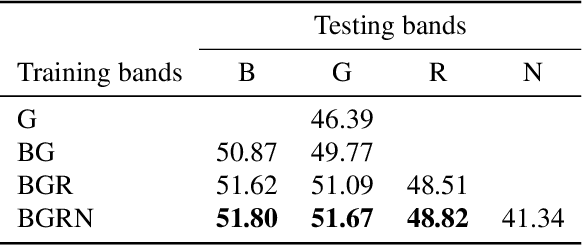
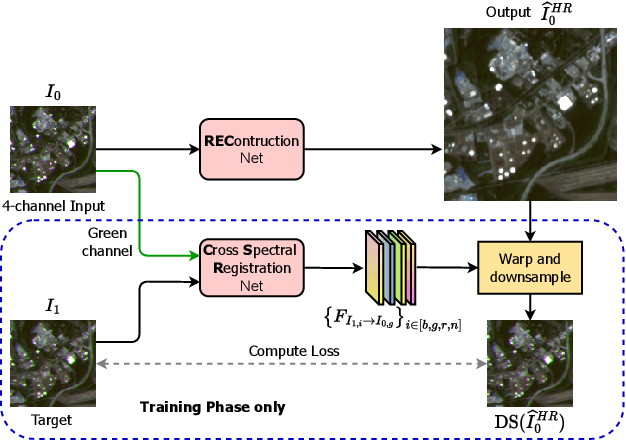
High-resolution satellite imagery is a key element for many Earth monitoring applications. Satellites such as Sentinel-2 feature characteristics that are favorable for super-resolution algorithms such as aliasing and band-misalignment. Unfortunately the lack of reliable high-resolution (HR) ground truth limits the application of deep learning methods to this task. In this work we propose L1BSR, a deep learning-based method for single-image super-resolution and band alignment of Sentinel-2 L1B 10m bands. The method is trained with self-supervision directly on real L1B data by leveraging overlapping areas in L1B images produced by adjacent CMOS detectors, thus not requiring HR ground truth. Our self-supervised loss is designed to enforce the super-resolved output image to have all the bands correctly aligned. This is achieved via a novel cross-spectral registration network (CSR) which computes an optical flow between images of different spectral bands. The CSR network is also trained with self-supervision using an Anchor-Consistency loss, which we also introduce in this work. We demonstrate the performance of the proposed approach on synthetic and real L1B data, where we show that it obtains comparable results to supervised methods.
Live image-based neurosurgical guidance and roadmap generation using unsupervised embedding
Mar 31, 2023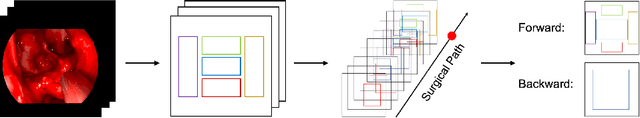
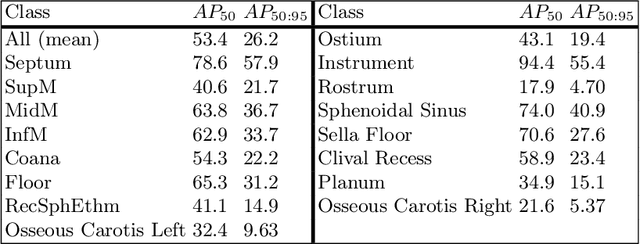
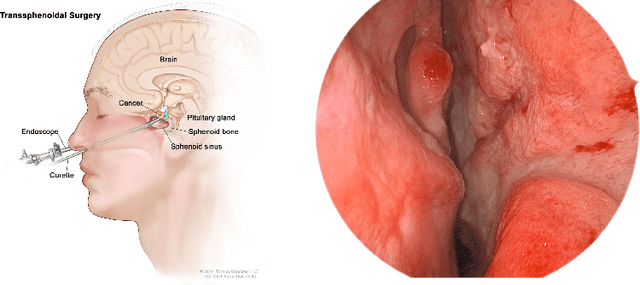
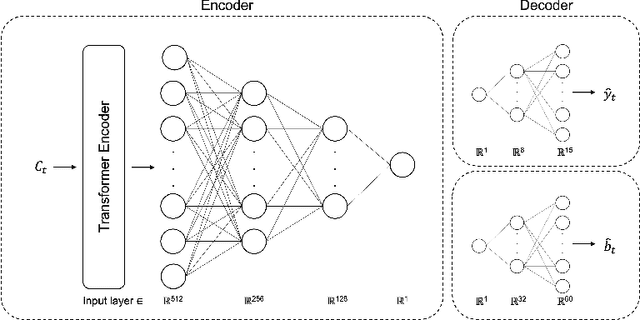
Advanced minimally invasive neurosurgery navigation relies mainly on Magnetic Resonance Imaging (MRI) guidance. MRI guidance, however, only provides pre-operative information in the majority of the cases. Once the surgery begins, the value of this guidance diminishes to some extent because of the anatomical changes due to surgery. Guidance with live image feedback coming directly from the surgical device, e.g., endoscope, can complement MRI-based navigation or be an alternative if MRI guidance is not feasible. With this motivation, we present a method for live image-only guidance leveraging a large data set of annotated neurosurgical videos.First, we report the performance of a deep learning-based object detection method, YOLO, on detecting anatomical structures in neurosurgical images. Second, we present a method for generating neurosurgical roadmaps using unsupervised embedding without assuming exact anatomical matches between patients, presence of an extensive anatomical atlas, or the need for simultaneous localization and mapping. A generated roadmap encodes the common anatomical paths taken in surgeries in the training set. At inference, the roadmap can be used to map a surgeon's current location using live image feedback on the path to provide guidance by being able to predict which structures should appear going forward or backward, much like a mapping application. Even though the embedding is not supervised by position information, we show that it is correlated to the location inside the brain and on the surgical path. We trained and evaluated the proposed method with a data set of 166 transsphenoidal adenomectomy procedures.
Ducho: A Unified Framework for the Extraction of Multimodal Features in Recommendation
Jun 29, 2023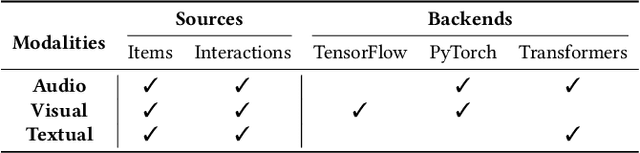
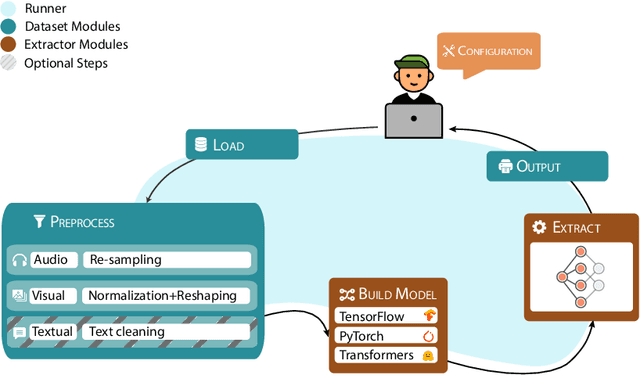
In multimodal-aware recommendation, the extraction of meaningful multimodal features is at the basis of high-quality recommendations. Generally, each recommendation framework implements its multimodal extraction procedures with specific strategies and tools. This is limiting for two reasons: (i) different extraction strategies do not ease the interdependence among multimodal recommendation frameworks; thus, they cannot be efficiently and fairly compared; (ii) given the large plethora of pre-trained deep learning models made available by different open source tools, model designers do not have access to shared interfaces to extract features. Motivated by the outlined aspects, we propose Ducho, a unified framework for the extraction of multimodal features in recommendation. By integrating three widely-adopted deep learning libraries as backends, namely, TensorFlow, PyTorch, and Transformers, we provide a shared interface to extract and process features where each backend's specific methods are abstracted to the end user. Noteworthy, the extraction pipeline is easily configurable with a YAML-based file where the user can specify, for each modality, the list of models (and their specific backends/parameters) to perform the extraction. Finally, to make Ducho accessible to the community, we build a public Docker image equipped with a ready-to-use CUDA environment and propose three demos to test its functionalities for different scenarios and tasks. The GitHub repository and the documentation is accessible at this link: https://github.com/sisinflab/Ducho.
MedNeXt: Transformer-driven Scaling of ConvNets for Medical Image Segmentation
Mar 22, 2023
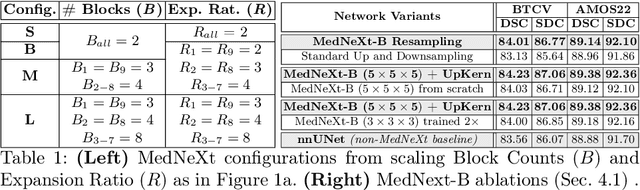

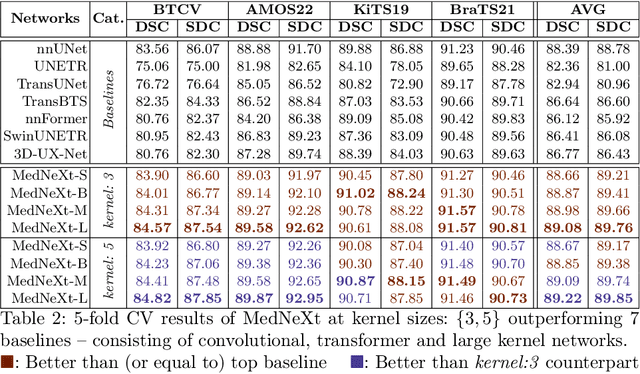
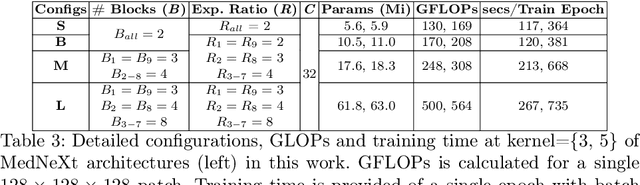
There has been exploding interest in embracing Transformer-based architectures for medical image segmentation. However, the lack of large-scale annotated medical datasets make achieving performances equivalent to those in natural images challenging. Convolutional networks, in contrast, have higher inductive biases and consequently, are easily trainable to high performance. Recently, the ConvNeXt architecture attempted to modernize the standard ConvNet by mirroring Transformer blocks. In this work, we improve upon this to design a modernized and scalable convolutional architecture customized to challenges of data-scarce medical settings. We introduce MedNeXt, a Transformer-inspired large kernel segmentation network which introduces - 1) A fully ConvNeXt 3D Encoder-Decoder Network for medical image segmentation, 2) Residual ConvNeXt up and downsampling blocks to preserve semantic richness across scales, 3) A novel technique to iteratively increase kernel sizes by upsampling small kernel networks, to prevent performance saturation on limited medical data, 4) Compound scaling at multiple levels (depth, width, kernel size) of MedNeXt. This leads to state-of-the-art performance on 4 tasks on CT and MRI modalities and varying dataset sizes, representing a modernized deep architecture for medical image segmentation.
Alteration-free and Model-agnostic Origin Attribution of Generated Images
May 29, 2023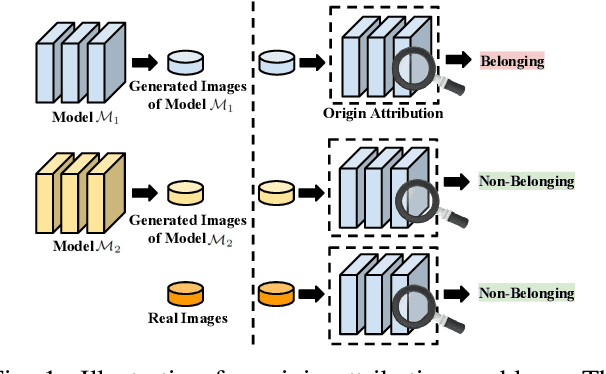
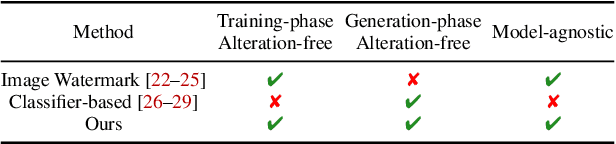

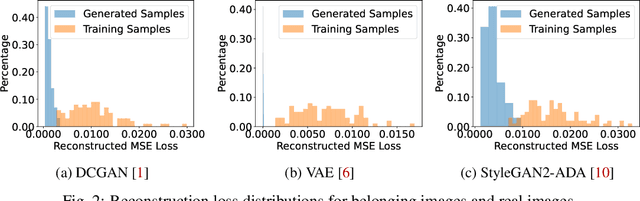
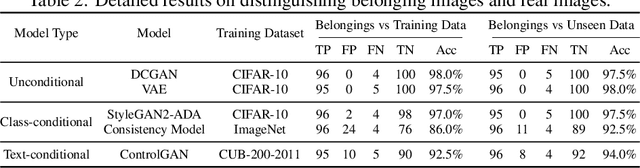
Recently, there has been a growing attention in image generation models. However, concerns have emerged regarding potential misuse and intellectual property (IP) infringement associated with these models. Therefore, it is necessary to analyze the origin of images by inferring if a specific image was generated by a particular model, i.e., origin attribution. Existing methods are limited in their applicability to specific types of generative models and require additional steps during training or generation. This restricts their use with pre-trained models that lack these specific operations and may compromise the quality of image generation. To overcome this problem, we first develop an alteration-free and model-agnostic origin attribution method via input reverse-engineering on image generation models, i.e., inverting the input of a particular model for a specific image. Given a particular model, we first analyze the differences in the hardness of reverse-engineering tasks for the generated images of the given model and other images. Based on our analysis, we propose a method that utilizes the reconstruction loss of reverse-engineering to infer the origin. Our proposed method effectively distinguishes between generated images from a specific generative model and other images, including those generated by different models and real images.
OTRE: Where Optimal Transport Guided Unpaired Image-to-Image Translation Meets Regularization by Enhancing
Feb 09, 2023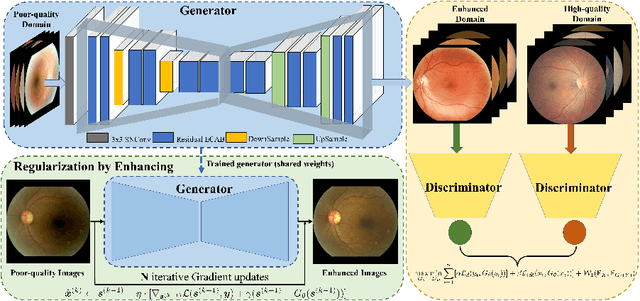
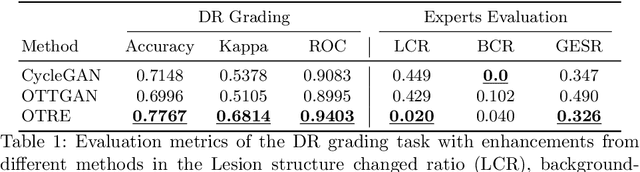
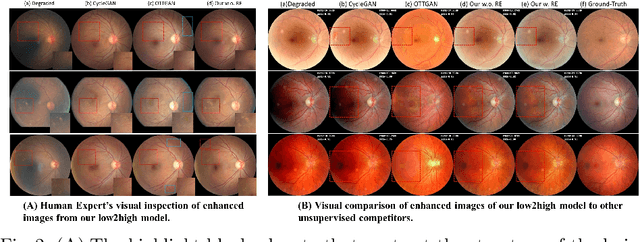
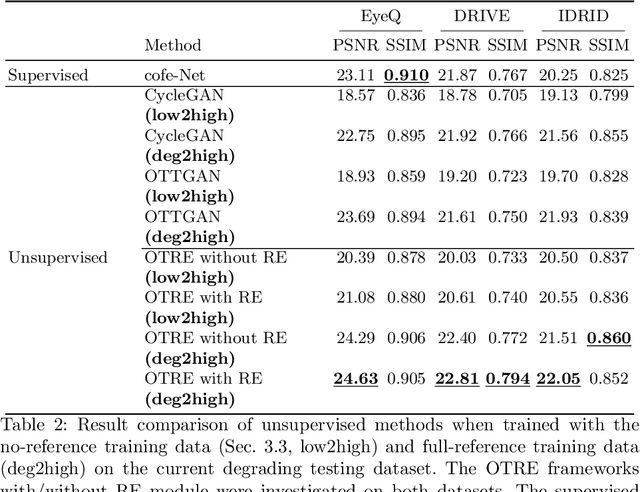
Non-mydriatic retinal color fundus photography (CFP) is widely available due to the advantage of not requiring pupillary dilation, however, is prone to poor quality due to operators, systemic imperfections, or patient-related causes. Optimal retinal image quality is mandated for accurate medical diagnoses and automated analyses. Herein, we leveraged the Optimal Transport (OT) theory to propose an unpaired image-to-image translation scheme for mapping low-quality retinal CFPs to high-quality counterparts. Furthermore, to improve the flexibility, robustness, and applicability of our image enhancement pipeline in the clinical practice, we generalized a state-of-the-art model-based image reconstruction method, regularization by denoising, by plugging in priors learned by our OT-guided image-to-image translation network. We named it as regularization by enhancing (RE). We validated the integrated framework, OTRE, on three publicly available retinal image datasets by assessing the quality after enhancement and their performance on various downstream tasks, including diabetic retinopathy grading, vessel segmentation, and diabetic lesion segmentation. The experimental results demonstrated the superiority of our proposed framework over some state-of-the-art unsupervised competitors and a state-of-the-art supervised method.
Capturing functional connectomics using Riemannian partial least squares
Jun 30, 2023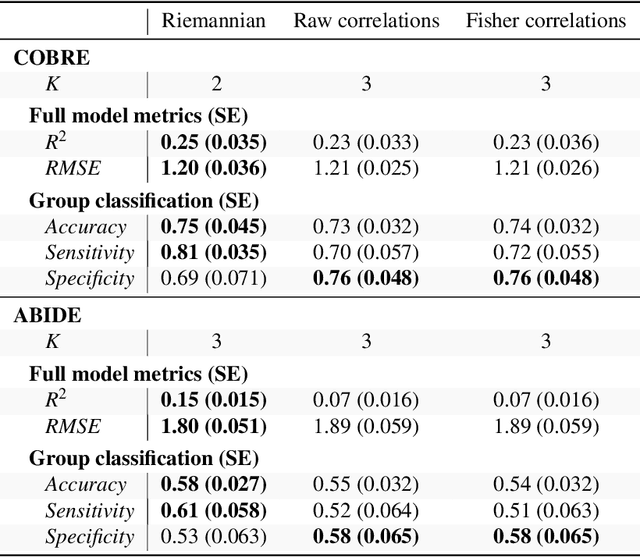
For neurological disorders and diseases, functional and anatomical connectomes of the human brain can be used to better inform targeted interventions and treatment strategies. Functional magnetic resonance imaging (fMRI) is a non-invasive neuroimaging technique that captures spatio-temporal brain function through blood flow over time. FMRI can be used to study the functional connectome through the functional connectivity matrix; that is, Pearson's correlation matrix between time series from the regions of interest of an fMRI image. One approach to analysing functional connectivity is using partial least squares (PLS), a multivariate regression technique designed for high-dimensional predictor data. However, analysing functional connectivity with PLS ignores a key property of the functional connectivity matrix; namely, these matrices are positive definite. To account for this, we introduce a generalisation of PLS to Riemannian manifolds, called R-PLS, and apply it to symmetric positive definite matrices with the affine invariant geometry. We apply R-PLS to two functional imaging datasets: COBRE, which investigates functional differences between schizophrenic patients and healthy controls, and; ABIDE, which compares people with autism spectrum disorder and neurotypical controls. Using the variable importance in the projection statistic on the results of R-PLS, we identify key functional connections in each dataset that are well represented in the literature. Given the generality of R-PLS, this method has potential to open up new avenues for multi-model imaging analysis linking structural and functional connectomics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge