Zitian Qu
LOGIGEN: Logic-Driven Generation of Verifiable Agentic Tasks
Feb 28, 2026Abstract:The evolution of Large Language Models (LLMs) from static instruction-followers to autonomous agents necessitates operating within complex, stateful environments to achieve precise state-transition objectives. However, this paradigm is bottlenecked by data scarcity, as existing tool-centric reverse-synthesis pipelines fail to capture the rigorous logic of real-world applications. We introduce \textbf{LOGIGEN}, a logic-driven framework that synthesizes verifiable training data based on three core pillars: \textbf{Hard-Compiled Policy Grounding}, \textbf{Logic-Driven Forward Synthesis}, and \textbf{Deterministic State Verification}. Specifically, a Triple-Agent Orchestration is employed: the \textbf{Architect} compiles natural-language policy into database constraints to enforce hard rules; the \textbf{Set Designer} initializes boundary-adjacent states to trigger critical policy conflicts; and the \textbf{Explorer} searches this environment to discover causal solution paths. This framework yields a dataset of 20,000 complex tasks across 8 domains, where validity is strictly guaranteed by checking exact state equivalence. Furthermore, we propose a verification-based training protocol where Supervised Fine-Tuning (SFT) on verifiable trajectories establishes compliance with hard-compiled policy, while Reinforcement Learning (RL) guided by dense state-rewards refines long-horizon goal achievement. On $τ^2$-Bench, LOGIGEN-32B(RL) achieves a \textbf{79.5\% success rate}, substantially outperforming the base model (40.7\%). These results demonstrate that logic-driven synthesis combined with verification-based training effectively constructs the causally valid trajectories needed for next-generation agents.
GENIE: Generative Note Information Extraction model for structuring EHR data
Jan 30, 2025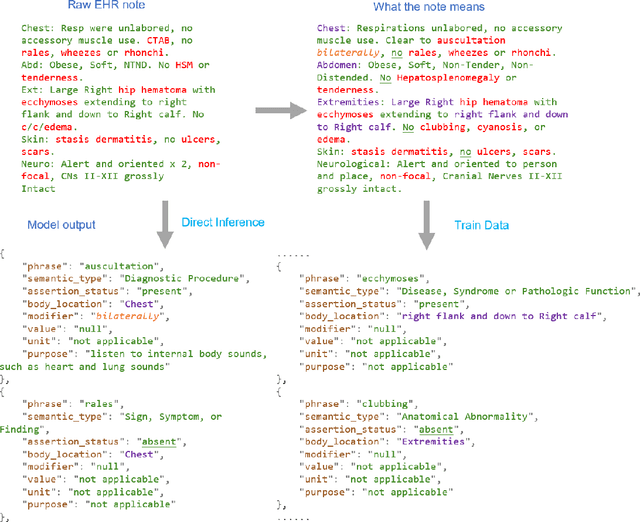
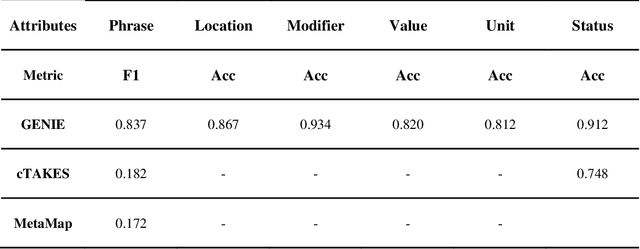
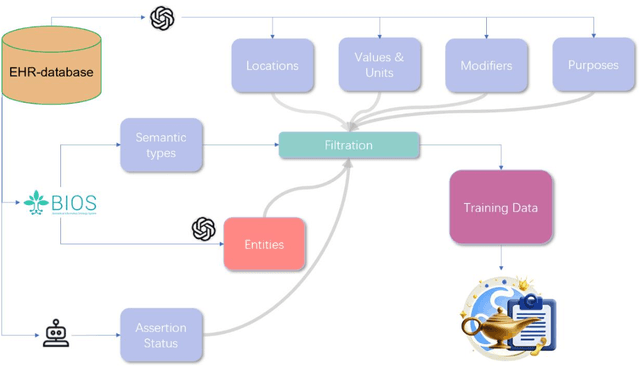
Abstract:Electronic Health Records (EHRs) hold immense potential for advancing healthcare, offering rich, longitudinal data that combines structured information with valuable insights from unstructured clinical notes. However, the unstructured nature of clinical text poses significant challenges for secondary applications. Traditional methods for structuring EHR free-text data, such as rule-based systems and multi-stage pipelines, are often limited by their time-consuming configurations and inability to adapt across clinical notes from diverse healthcare settings. Few systems provide a comprehensive attribute extraction for terminologies. While giant large language models (LLMs) like GPT-4 and LLaMA 405B excel at structuring tasks, they are slow, costly, and impractical for large-scale use. To overcome these limitations, we introduce GENIE, a Generative Note Information Extraction system that leverages LLMs to streamline the structuring of unstructured clinical text into usable data with standardized format. GENIE processes entire paragraphs in a single pass, extracting entities, assertion statuses, locations, modifiers, values, and purposes with high accuracy. Its unified, end-to-end approach simplifies workflows, reduces errors, and eliminates the need for extensive manual intervention. Using a robust data preparation pipeline and fine-tuned small scale LLMs, GENIE achieves competitive performance across multiple information extraction tasks, outperforming traditional tools like cTAKES and MetaMap and can handle extra attributes to be extracted. GENIE strongly enhances real-world applicability and scalability in healthcare systems. By open-sourcing the model and test data, we aim to encourage collaboration and drive further advancements in EHR structurization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge