Zhuoyan Xu
Efficient Table Retrieval and Understanding with Multimodal Large Language Models
Feb 07, 2026Abstract:Tabular data is frequently captured in image form across a wide range of real-world scenarios such as financial reports, handwritten records, and document scans. These visual representations pose unique challenges for machine understanding, as they combine both structural and visual complexities. While recent advances in Multimodal Large Language Models (MLLMs) show promising results in table understanding, they typically assume the relevant table is readily available. However, a more practical scenario involves identifying and reasoning over relevant tables from large-scale collections to answer user queries. To address this gap, we propose TabRAG, a framework that enables MLLMs to answer queries over large collections of table images. Our approach first retrieves candidate tables using jointly trained visual-text foundation models, then leverages MLLMs to perform fine-grained reranking of these candidates, and finally employs MLLMs to reason over the selected tables for answer generation. Through extensive experiments on a newly constructed dataset comprising 88,161 training and 9,819 testing samples across 8 benchmarks with 48,504 unique tables, we demonstrate that our framework significantly outperforms existing methods by 7.0% in retrieval recall and 6.1% in answer accuracy, offering a practical solution for real-world table understanding tasks.
Can Language Models Compose Skills In-Context?
Oct 27, 2025Abstract:Composing basic skills from simple tasks to accomplish composite tasks is crucial for modern intelligent systems. We investigate the in-context composition ability of language models to perform composite tasks that combine basic skills demonstrated in in-context examples. This is more challenging than the standard setting, where skills and their composition can be learned in training. We conduct systematic experiments on various representative open-source language models, utilizing linguistic and logical tasks designed to probe composition abilities. The results reveal that simple task examples can have a surprising negative impact on the performance, because the models generally struggle to recognize and assemble the skills correctly, even with Chain-of-Thought examples. Theoretical analysis further shows that it is crucial to align examples with the corresponding steps in the composition. This inspires a method for the probing tasks, whose improved performance provides positive support for our insights.
Neural at ArchEHR-QA 2025: Agentic Prompt Optimization for Evidence-Grounded Clinical Question Answering
Jun 12, 2025Abstract:Automated question answering (QA) over electronic health records (EHRs) can bridge critical information gaps for clinicians and patients, yet it demands both precise evidence retrieval and faithful answer generation under limited supervision. In this work, we present Neural, the runner-up in the BioNLP 2025 ArchEHR-QA shared task on evidence-grounded clinical QA. Our proposed method decouples the task into (1) sentence-level evidence identification and (2) answer synthesis with explicit citations. For each stage, we automatically explore the prompt space with DSPy's MIPROv2 optimizer, jointly tuning instructions and few-shot demonstrations on the development set. A self-consistency voting scheme further improves evidence recall without sacrificing precision. On the hidden test set, our method attains an overall score of 51.5, placing second stage while outperforming standard zero-shot and few-shot prompting by over 20 and 10 points, respectively. These results indicate that data-driven prompt optimization is a cost-effective alternative to model fine-tuning for high-stakes clinical QA, advancing the reliability of AI assistants in healthcare.
Learning to Inference Adaptively for Multimodal Large Language Models
Mar 13, 2025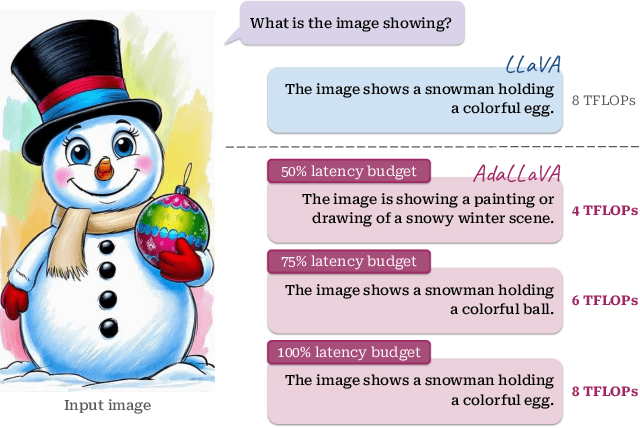
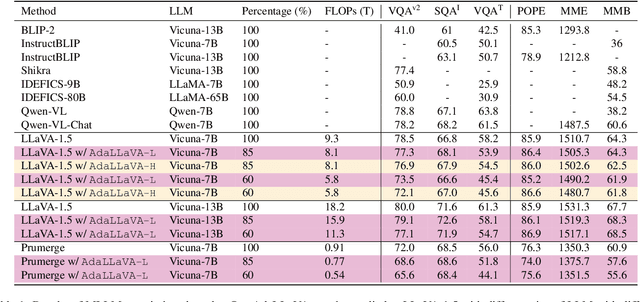
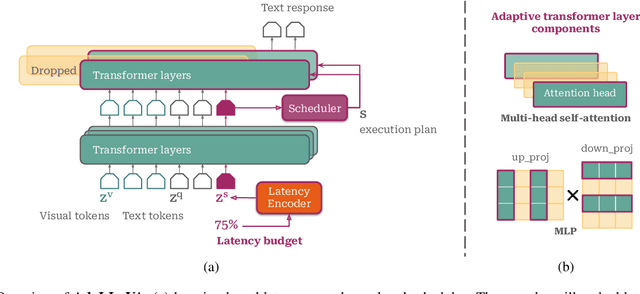
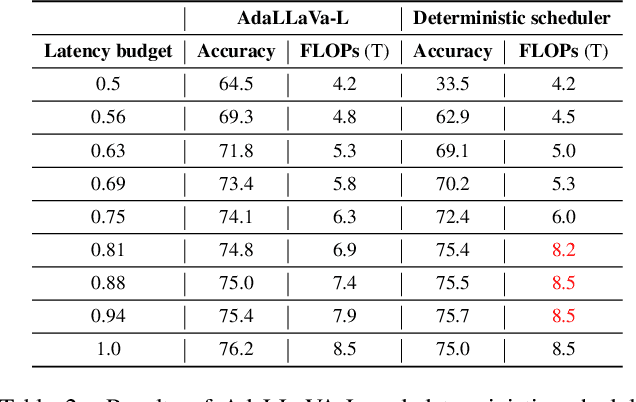
Abstract:Multimodal Large Language Models (MLLMs) have shown impressive capabilities in reasoning, yet come with substantial computational cost, limiting their deployment in resource-constrained settings. Despite recent efforts on improving the efficiency of MLLMs, prior solutions fall short in responding to varying runtime conditions, in particular changing resource availability (e.g., contention due to the execution of other programs on the device). To bridge this gap, we introduce AdaLLaVA, an adaptive inference framework that learns to dynamically reconfigure operations in an MLLM during inference, accounting for the input data and a latency budget. We conduct extensive experiments across benchmarks involving question-answering, reasoning, and hallucination. Our results show that AdaLLaVA effectively adheres to input latency budget, achieving varying accuracy and latency tradeoffs at runtime. Further, we demonstrate that AdaLLaVA adapts to both input latency and content, can be integrated with token selection for enhanced efficiency, and generalizes across MLLMs.Our project webpage with code release is at https://zhuoyan-xu.github.io/ada-llava/.
Do Large Language Models Have Compositional Ability? An Investigation into Limitations and Scalability
Jul 22, 2024
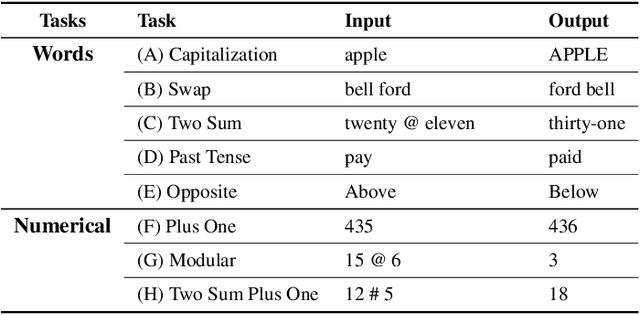
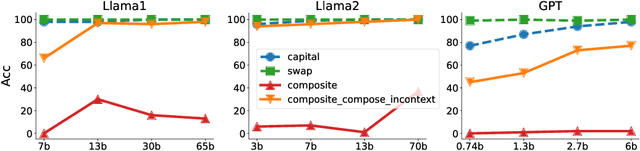
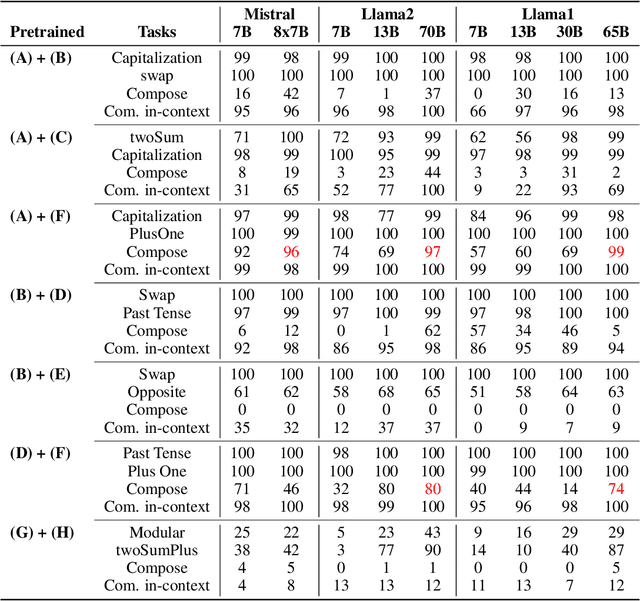
Abstract:Large language models (LLMs) have emerged as powerful tools for many AI problems and exhibit remarkable in-context learning (ICL) capabilities. Compositional ability, solving unseen complex tasks that combine two or more simple tasks, is an essential reasoning ability for Artificial General Intelligence. Despite LLM's tremendous success, how they approach composite tasks, especially those not encountered during the pretraining phase, remains an open question and largely ununderstood. In this study, we delve into the ICL capabilities of LLMs on composite tasks, with only simple tasks as in-context examples. We develop a test suite of composite tasks that include linguistic and logical challenges and perform empirical studies across different LLM families. We observe that models exhibit divergent behaviors: (1) For simpler composite tasks that apply distinct mapping mechanisms to different input segments, the models demonstrate decent compositional ability, while scaling up the model enhances this ability; (2) for more complex composite tasks that involving reasoning multiple steps, where each step represent one task, models typically underperform, and scaling up generally provide no improvements. We offer theoretical analysis in a simplified setting, explaining that models exhibit compositional capability when the task handles different input parts separately. We believe our work sheds new light on the capabilities of LLMs in solving composite tasks regarding the nature of the tasks and model scale. Our dataset and code are available at {\url{https://github.com/OliverXUZY/LLM_Compose}}.
Why Larger Language Models Do In-context Learning Differently?
May 30, 2024


Abstract:Large language models (LLM) have emerged as a powerful tool for AI, with the key ability of in-context learning (ICL), where they can perform well on unseen tasks based on a brief series of task examples without necessitating any adjustments to the model parameters. One recent interesting mysterious observation is that models of different scales may have different ICL behaviors: larger models tend to be more sensitive to noise in the test context. This work studies this observation theoretically aiming to improve the understanding of LLM and ICL. We analyze two stylized settings: (1) linear regression with one-layer single-head linear transformers and (2) parity classification with two-layer multiple attention heads transformers (non-linear data and non-linear model). In both settings, we give closed-form optimal solutions and find that smaller models emphasize important hidden features while larger ones cover more hidden features; thus, smaller models are more robust to noise while larger ones are more easily distracted, leading to different ICL behaviors. This sheds light on where transformers pay attention to and how that affects ICL. Preliminary experimental results on large base and chat models provide positive support for our analysis.
Towards Few-Shot Adaptation of Foundation Models via Multitask Finetuning
Feb 22, 2024



Abstract:Foundation models have emerged as a powerful tool for many AI problems. Despite the tremendous success of foundation models, effective adaptation to new tasks, particularly those with limited labels, remains an open question and lacks theoretical understanding. An emerging solution with recent success in vision and NLP involves finetuning a foundation model on a selection of relevant tasks, before its adaptation to a target task with limited labeled samples. In this paper, we study the theoretical justification of this multitask finetuning approach. Our theoretical analysis reveals that with a diverse set of related tasks, this multitask finetuning leads to reduced error in the target task, in comparison to directly adapting the same pretrained model. We quantify the relationship between finetuning tasks and target tasks by diversity and consistency metrics, and further propose a practical task selection algorithm. We substantiate our theoretical claims with extensive empirical evidence. Further, we present results affirming our task selection algorithm adeptly chooses related finetuning tasks, providing advantages to the model performance on target tasks. We believe our study shed new light on the effective adaptation of foundation models to new tasks that lack abundant labels. Our code is available at https://github.com/OliverXUZY/Foudation-Model_Multitask.
Spatial Transcriptomics Dimensionality Reduction using Wavelet Bases
May 19, 2022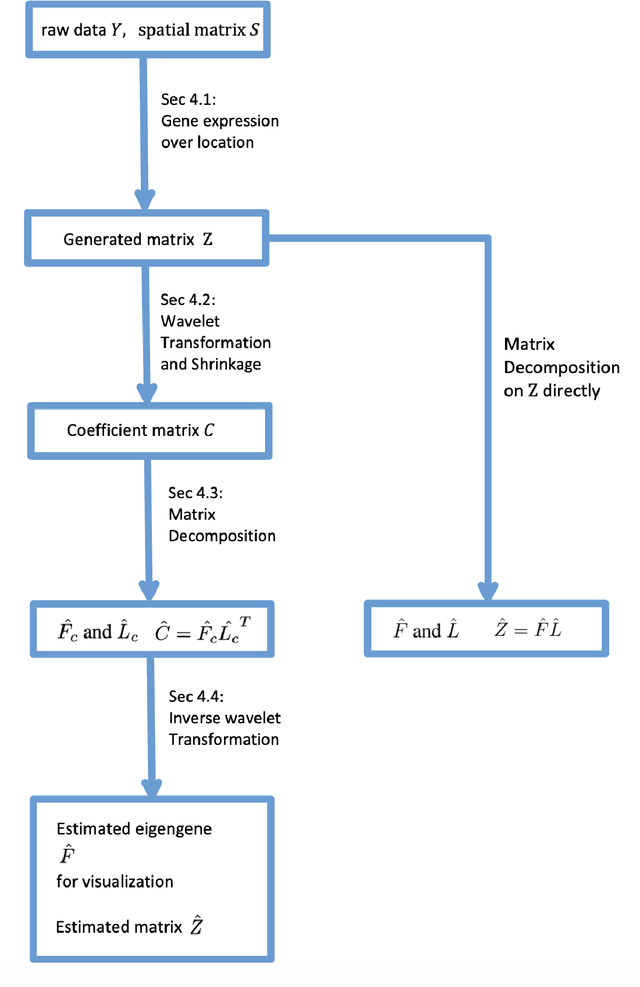
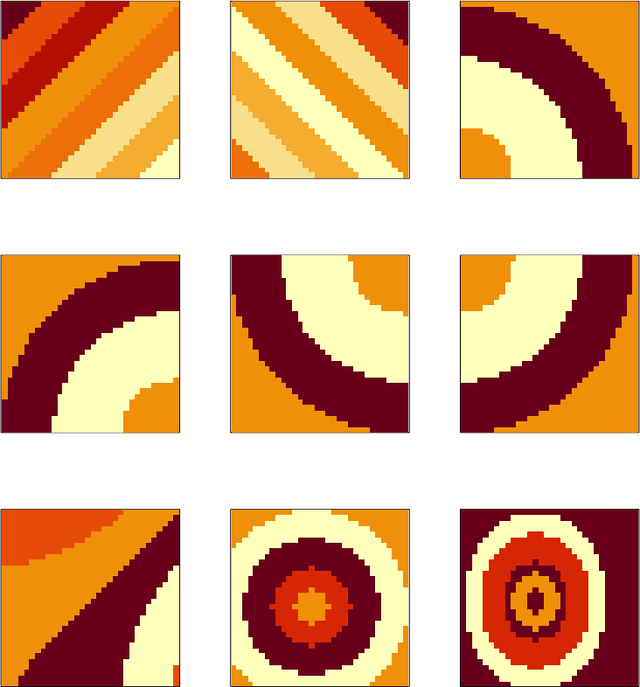
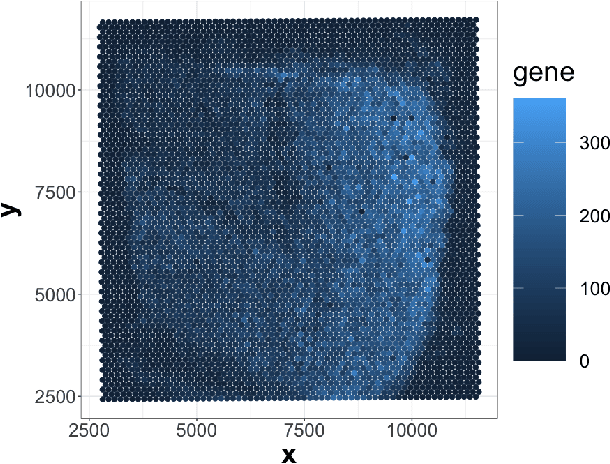
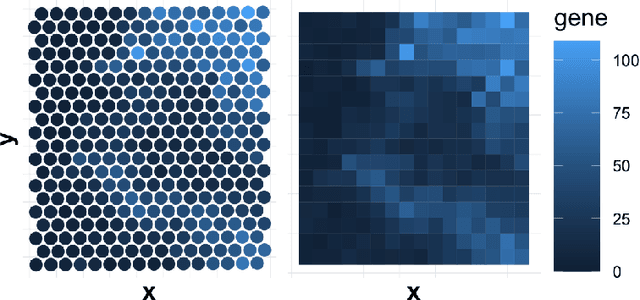

Abstract:Spatially resolved transcriptomics (ST) measures gene expression along with the spatial coordinates of the measurements. The analysis of ST data involves significant computation complexity. In this work, we propose gene expression dimensionality reduction algorithm that retains spatial structure. We combine the wavelet transformation with matrix factorization to select spatially-varying genes. We extract a low-dimensional representation of these genes. We consider Empirical Bayes setting, imposing regularization through the prior distribution of factor genes. Additionally, We provide visualization of extracted representation genes capturing the global spatial pattern. We illustrate the performance of our methods by spatial structure recovery and gene expression reconstruction in simulation. In real data experiments, our method identifies spatial structure of gene factors and outperforms regular decomposition regarding reconstruction error. We found the connection between the fluctuation of gene patterns and wavelet technique, providing smoother visualization. We develop the package and share the workflow generating reproducible quantitative results and gene visualization. The package is available at https://github.com/OliverXUZY/waveST.
Generalized tensor regression with covariates on multiple modes
Oct 21, 2019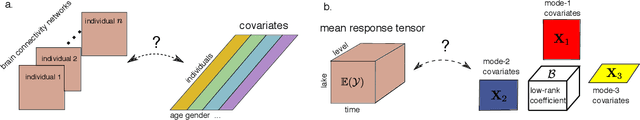
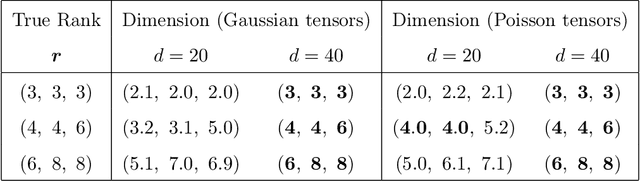
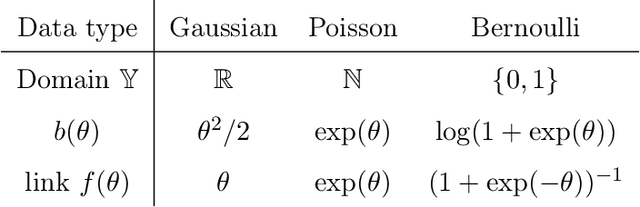

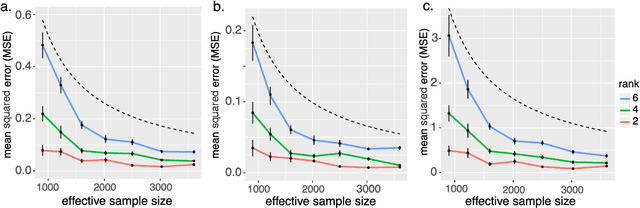
Abstract:We consider the problem of tensor-response regression given covariates on multiple modes. Such data problems arise frequently in applications such as neuroimaging, network analysis, and spatial-temporal modeling. We propose a new family of tensor response regression models that incorporate covariates, and establish the theoretical accuracy guarantees. Unlike earlier methods, our estimation allows high-dimensionality in both the tensor response and the covariate matrices on multiple modes. An efficient alternating updating algorithm is further developed. Our proposal handles a broad range of data types, including continuous, count, and binary observations. Through simulation and applications to two real datasets, we demonstrate the outperformance of our approach over the state-of-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge