Yong Fan
from the iSTAGING consortium, for the ADNI
Disentangling Alzheimer's disease neurodegeneration from typical brain aging using machine learning
Sep 08, 2021
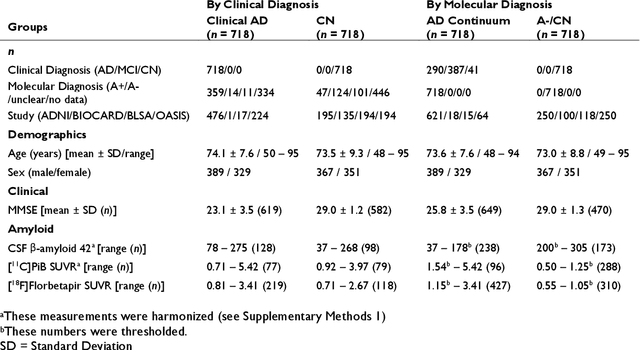
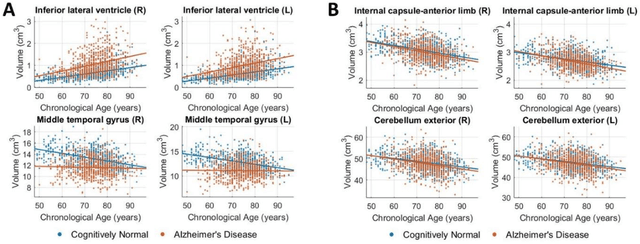
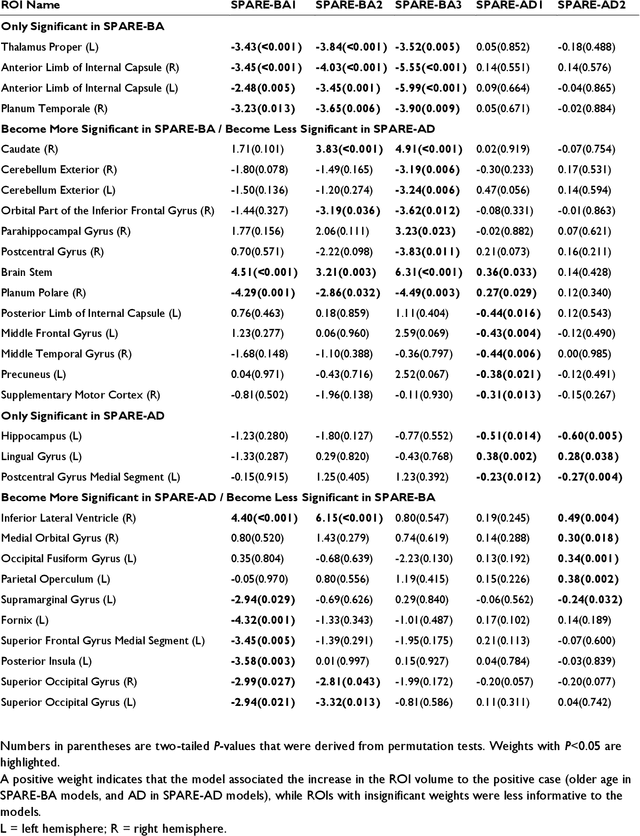
Abstract:Neuroimaging biomarkers that distinguish between typical brain aging and Alzheimer's disease (AD) are valuable for determining how much each contributes to cognitive decline. Machine learning models can derive multi-variate brain change patterns related to the two processes, including the SPARE-AD (Spatial Patterns of Atrophy for Recognition of Alzheimer's Disease) and SPARE-BA (of Brain Aging) investigated herein. However, substantial overlap between brain regions affected in the two processes confounds measuring them independently. We present a methodology toward disentangling the two. T1-weighted MRI images of 4,054 participants (48-95 years) with AD, mild cognitive impairment (MCI), or cognitively normal (CN) diagnoses from the iSTAGING (Imaging-based coordinate SysTem for AGIng and NeurodeGenerative diseases) consortium were analyzed. First, a subset of AD patients and CN adults were selected based purely on clinical diagnoses to train SPARE-BA1 (regression of age using CN individuals) and SPARE-AD1 (classification of CN versus AD). Second, analogous groups were selected based on clinical and molecular markers to train SPARE-BA2 and SPARE-AD2: amyloid-positive (A+) AD continuum group (consisting of A+AD, A+MCI, and A+ and tau-positive CN individuals) and amyloid-negative (A-) CN group. Finally, the combined group of the AD continuum and A-/CN individuals was used to train SPARE-BA3, with the intention to estimate brain age regardless of AD-related brain changes. Disentangled SPARE models derived brain patterns that were more specific to the two types of the brain changes. Correlation between the SPARE-BA and SPARE-AD was significantly reduced. Correlation of disentangled SPARE-AD was non-inferior to the molecular measurements and to the number of APOE4 alleles, but was less to AD-related psychometric test scores, suggesting contribution of advanced brain aging to these scores.
Unsupervised deep learning for individualized brain functional network identification
Dec 11, 2020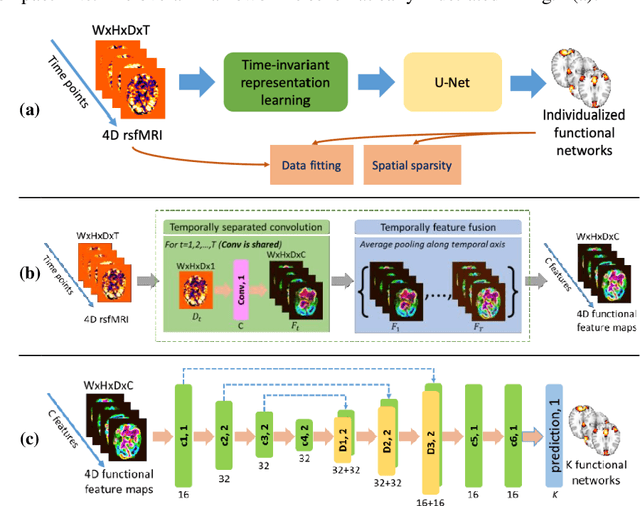
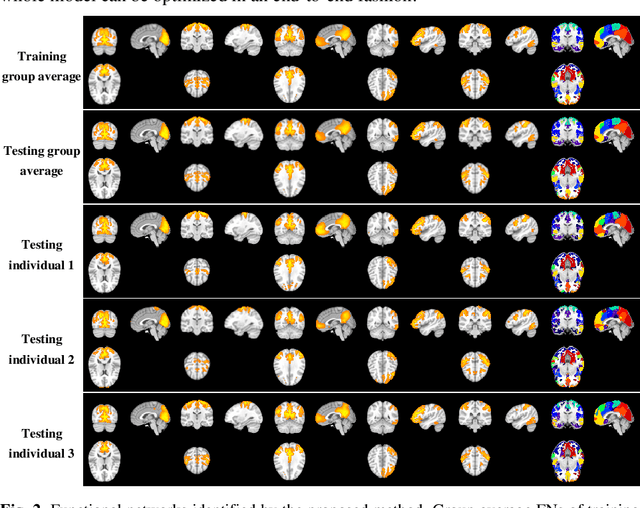
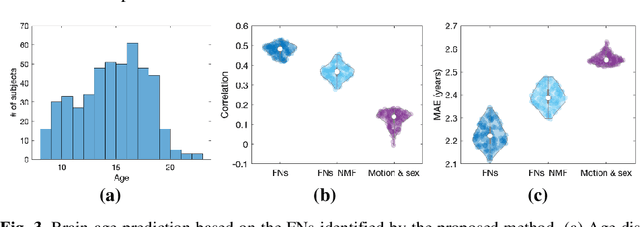
Abstract:A novel unsupervised deep learning method is developed to identify individual-specific large scale brain functional networks (FNs) from resting-state fMRI (rsfMRI) in an end-to-end learning fashion. Our method leverages deep Encoder-Decoder networks and conventional brain decomposition models to identify individual-specific FNs in an unsupervised learning framework and facilitate fast inference for new individuals with one forward pass of the deep network. Particularly, convolutional neural networks (CNNs) with an Encoder-Decoder architecture are adopted to identify individual-specific FNs from rsfMRI data by optimizing their data fitting and sparsity regularization terms that are commonly used in brain decomposition models. Moreover, a time-invariant representation learning module is designed to learn features invariant to temporal orders of time points of rsfMRI data. The proposed method has been validated based on a large rsfMRI dataset and experimental results have demonstrated that our method could obtain individual-specific FNs which are consistent with well-established FNs and are informative for predicting brain age, indicating that the individual-specific FNs identified truly captured the underlying variability of individualized functional neuroanatomy.
Meta-Learning for Natural Language Understanding under Continual Learning Framework
Nov 03, 2020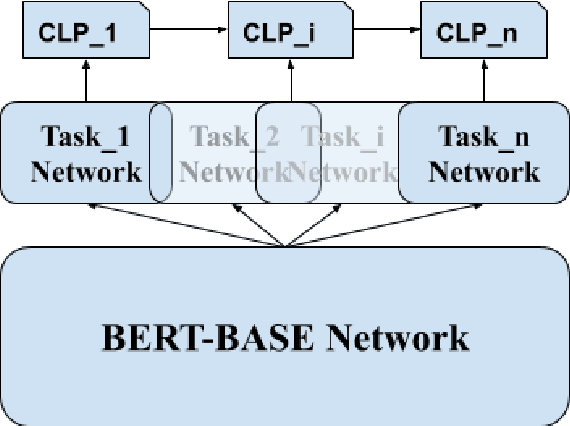
Abstract:Neural network has been recognized with its accomplishments on tackling various natural language understanding (NLU) tasks. Methods have been developed to train a robust model to handle multiple tasks to gain a general representation of text. In this paper, we implement the model-agnostic meta-learning (MAML) and Online aware Meta-learning (OML) meta-objective under the continual framework for NLU tasks. We validate our methods on selected SuperGLUE and GLUE benchmark.
Medical Image Harmonization Using Deep Learning Based Canonical Mapping: Toward Robust and Generalizable Learning in Imaging
Oct 11, 2020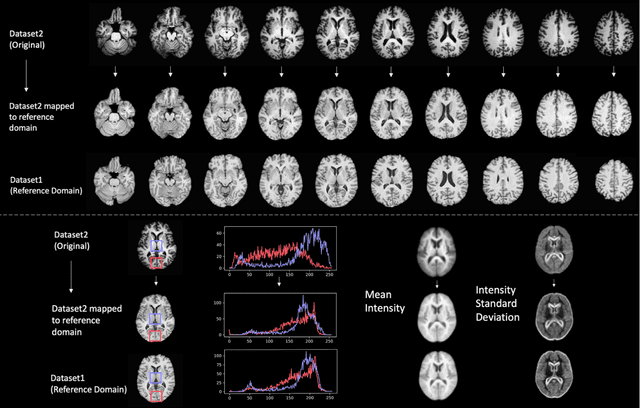
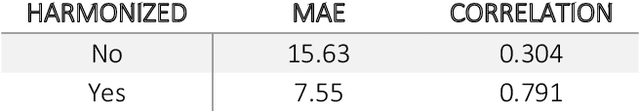
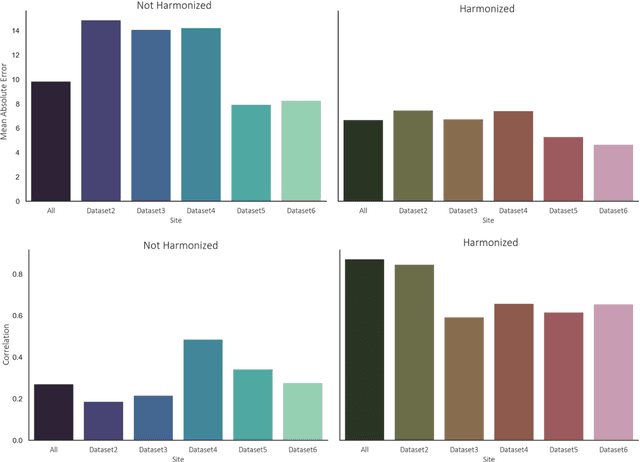
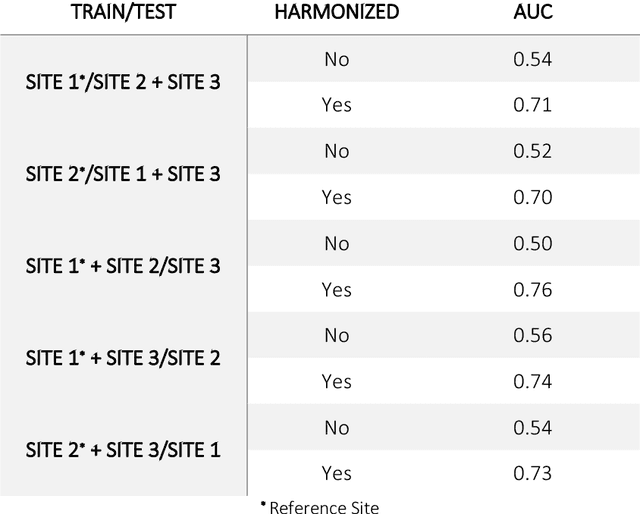
Abstract:Conventional and deep learning-based methods have shown great potential in the medical imaging domain, as means for deriving diagnostic, prognostic, and predictive biomarkers, and by contributing to precision medicine. However, these methods have yet to see widespread clinical adoption, in part due to limited generalization performance across various imaging devices, acquisition protocols, and patient populations. In this work, we propose a new paradigm in which data from a diverse range of acquisition conditions are "harmonized" to a common reference domain, where accurate model learning and prediction can take place. By learning an unsupervised image to image canonical mapping from diverse datasets to a reference domain using generative deep learning models, we aim to reduce confounding data variation while preserving semantic information, thereby rendering the learning task easier in the reference domain. We test this approach on two example problems, namely MRI-based brain age prediction and classification of schizophrenia, leveraging pooled cohorts of neuroimaging MRI data spanning 9 sites and 9701 subjects. Our results indicate a substantial improvement in these tasks in out-of-sample data, even when training is restricted to a single site.
MDReg-Net: Multi-resolution diffeomorphic image registration using fully convolutional networks with deep self-supervision
Oct 04, 2020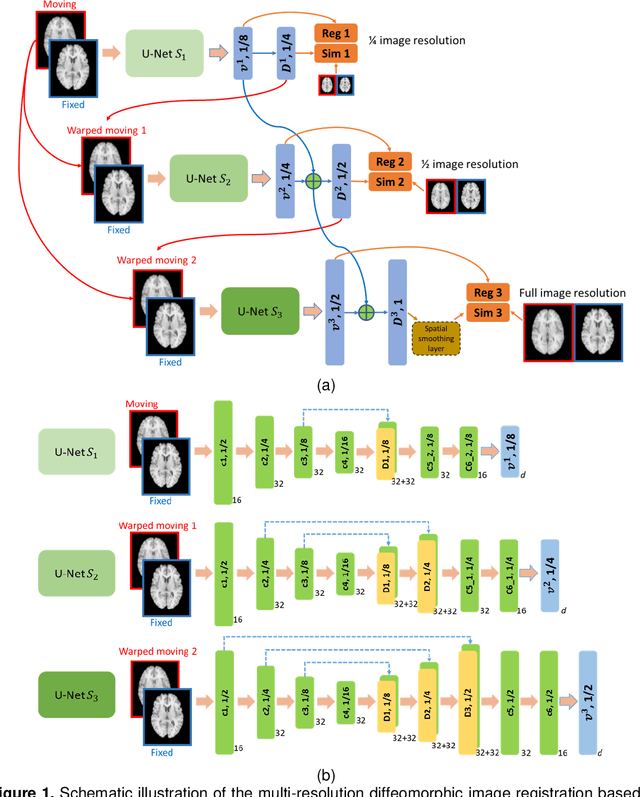
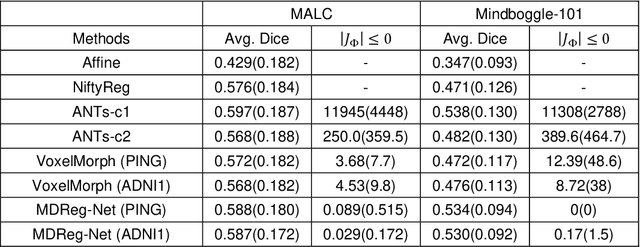
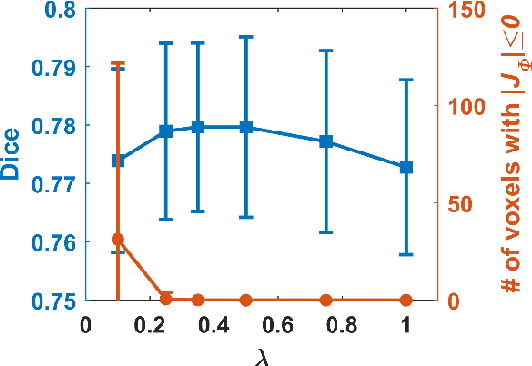
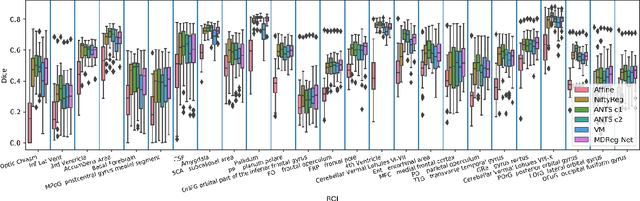
Abstract:We present a diffeomorphic image registration algorithm to learn spatial transformations between pairs of images to be registered using fully convolutional networks (FCNs) under a self-supervised learning setting. The network is trained to estimate diffeomorphic spatial transformations between pairs of images by maximizing an image-wise similarity metric between fixed and warped moving images, similar to conventional image registration algorithms. It is implemented in a multi-resolution image registration framework to optimize and learn spatial transformations at different image resolutions jointly and incrementally with deep self-supervision in order to better handle large deformation between images. A spatial Gaussian smoothing kernel is integrated with the FCNs to yield sufficiently smooth deformation fields to achieve diffeomorphic image registration. Particularly, spatial transformations learned at coarser resolutions are utilized to warp the moving image, which is subsequently used for learning incremental transformations at finer resolutions. This procedure proceeds recursively to the full image resolution and the accumulated transformations serve as the final transformation to warp the moving image at the finest resolution. Experimental results for registering high resolution 3D structural brain magnetic resonance (MR) images have demonstrated that image registration networks trained by our method obtain robust, diffeomorphic image registration results within seconds with improved accuracy compared with state-of-the-art image registration algorithms.
Adaptive convolutional neural networks for k-space data interpolation in fast magnetic resonance imaging
Jun 09, 2020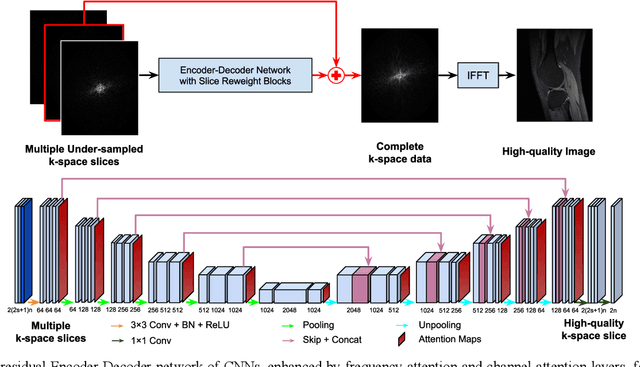
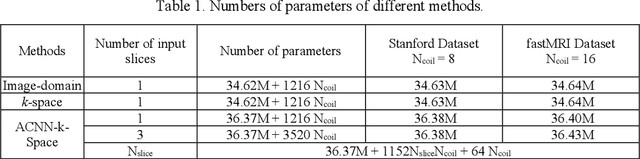
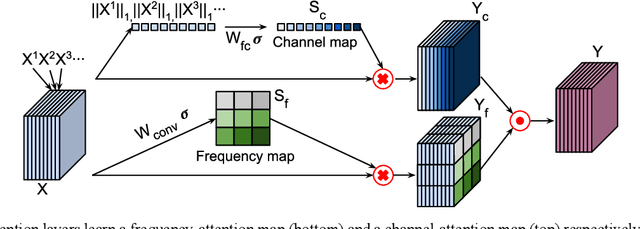
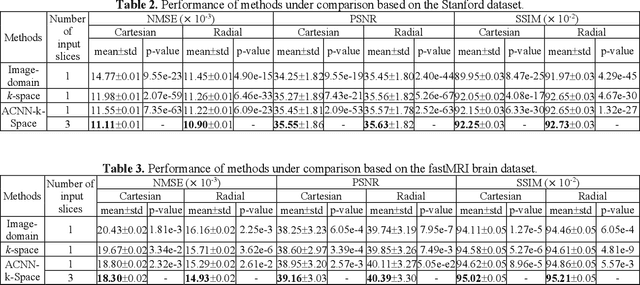
Abstract:Deep learning in k-space has demonstrated great potential for image reconstruction from undersampled k-space data in fast magnetic resonance imaging (MRI). However, existing deep learning-based image reconstruction methods typically apply weight-sharing convolutional neural networks (CNNs) to k-space data without taking into consideration the k-space data's spatial frequency properties, leading to ineffective learning of the image reconstruction models. Moreover, complementary information of spatially adjacent slices is often ignored in existing deep learning methods. To overcome such limitations, we develop a deep learning algorithm, referred to as adaptive convolutional neural networks for k-space data interpolation (ACNN-k-Space), which adopts a residual Encoder-Decoder network architecture to interpolate the undersampled k-space data by integrating spatially contiguous slices as multi-channel input, along with k-space data from multiple coils if available. The network is enhanced by self-attention layers to adaptively focus on k-space data at different spatial frequencies and channels. We have evaluated our method on two public datasets and compared it with state-of-the-art existing methods. Ablation studies and experimental results demonstrate that our method effectively reconstructs images from undersampled k-space data and achieves significantly better image reconstruction performance than current state-of-the-art techniques.
ACEnet: Anatomical Context-Encoding Network for Neuroanatomy Segmentation
Feb 13, 2020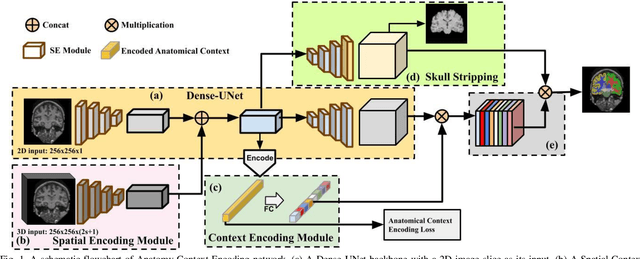
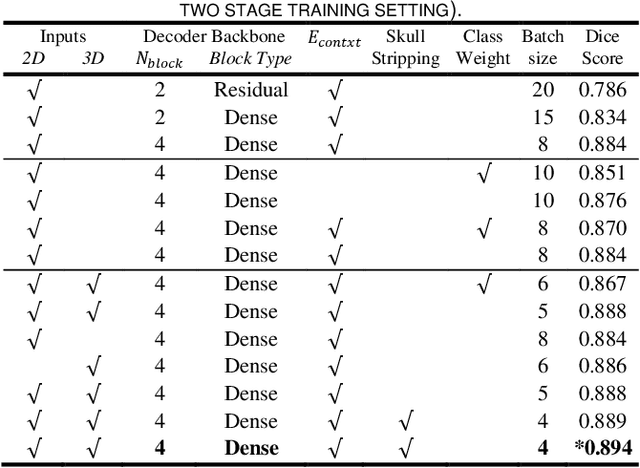
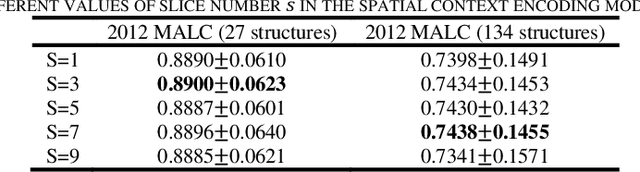
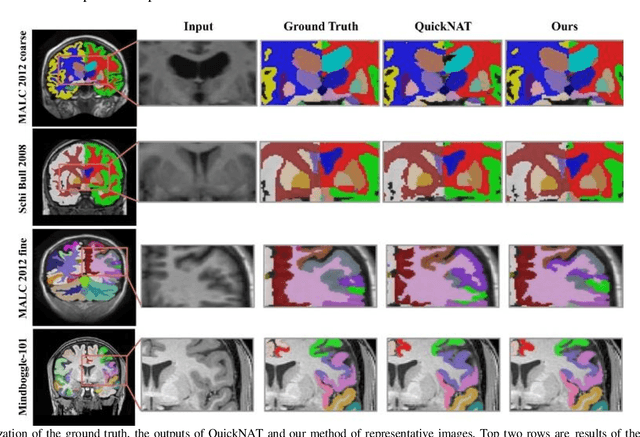
Abstract:Segmentation of brain structures from magnetic resonance (MR) scans plays an important role in the quantification of brain morphology. Since 3D deep learning models suffer from high computational cost, 2D deep learning methods are favored for their computational efficiency. However, existing 2D deep learning methods are not equipped to effectively capture 3D spatial contextual information that is needed to achieve accurate brain structure segmentation. In order to overcome this limitation, we develop an Anatomical Context-Encoding Network (ACEnet) to incorporate 3D spatial and anatomical contexts in 2D convolutional neural networks (CNNs) for efficient and accurate segmentation of brain structures from MR scans, consisting of 1) an anatomical context encoding module to incorporate anatomical information in 2D CNNs, 2) a spatial context encoding module to integrate 3D image information in 2D CNNs, and 3) a skull stripping module to guide 2D CNNs to attend to the brain. Extensive experiments on three benchmark datasets have demonstrated that our method outperforms state-of-the-art alternative methods for brain structure segmentation in terms of both computational efficiency and segmentation accuracy.
Context-endcoding for neural network based skull stripping in magnetic resonance imaging
Oct 23, 2019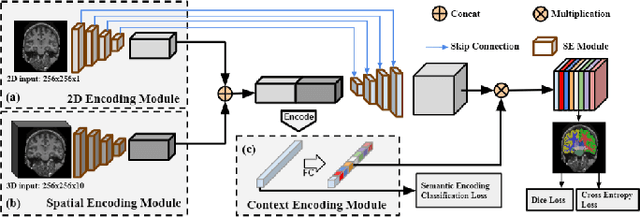
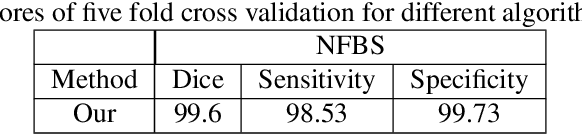
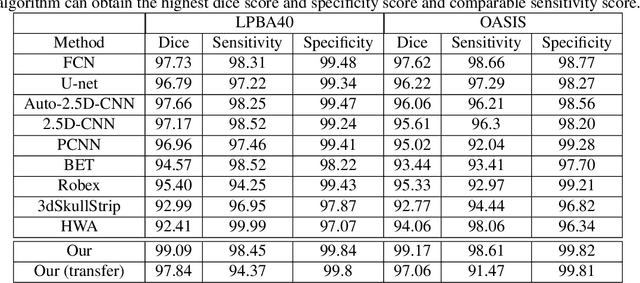
Abstract:Skull stripping is usually the first step for most brain analysisprocess in magnetic resonance images. A lot of deep learn-ing neural network based methods have been developed toachieve higher accuracy. Since the 3D deep learning modelssuffer from high computational cost and are subject to GPUmemory limit challenge, a variety of 2D deep learning meth-ods have been developed. However, existing 2D deep learn-ing methods are not equipped to effectively capture 3D se-mantic information that is needed to achieve higher accuracy.In this paper, we propose a context-encoding method to em-power the 2D network to capture the 3D context information.For the context-encoding method, firstly we encode the 2Dfeatures of original 2D network, secondly we encode the sub-volume of 3D MRI images, finally we fuse the encoded 2Dfeatures and 3D features with semantic encoding classifica-tion loss. To get computational efficiency, although we en-code the sub-volume of 3D MRI images instead of buildinga 3D neural network, extensive experiments on three bench-mark Datasets demonstrate our method can achieve superioraccuracy to state-of-the-art alternative methods with the dicescore 99.6% on NFBS and 99.09 % on LPBA40 and 99.17 %on OASIS.
Feature-Fused Context-Encoding Network for Neuroanatomy Segmentation
May 07, 2019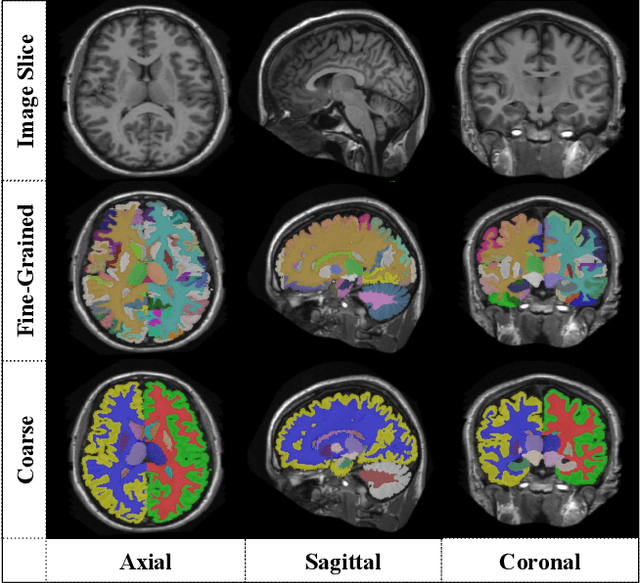
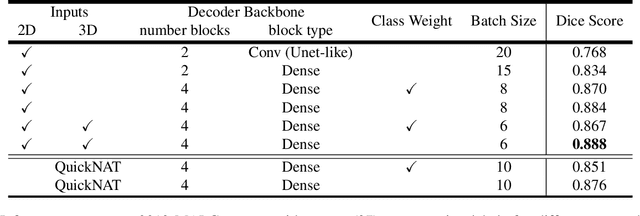
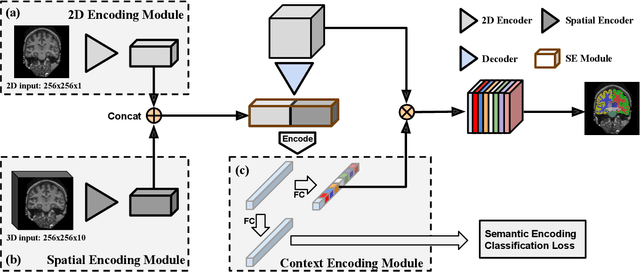
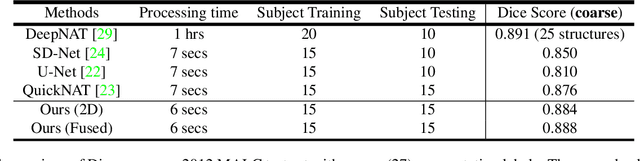
Abstract:Automatic segmentation of fine-grained brain structures remains a challenging task. Current segmentation methods mainly utilize 2D and 3D deep neural networks. The 2D networks take image slices as input to produce coarse segmentation in less processing time, whereas the 3D networks take the whole image volumes to generated fine-detailed segmentation with more computational burden. In order to obtain accurate fine-grained segmentation efficiently, in this paper, we propose an end-to-end Feature-Fused Context-Encoding Network for brain structure segmentation from MR (magnetic resonance) images. Our model is implemented based on a 2D convolutional backbone, which integrates a 2D encoding module to acquire planar image features and a spatial encoding module to extract spatial context information. A global context encoding module is further introduced to capture global context semantics from the fused 2D encoding and spatial features. The proposed network aims to fully leverage the global anatomical prior knowledge learned from context semantics, which is represented by a structure-aware attention factor to recalibrate the outputs of the network. In this way, the network is guaranteed to be aware of the class-dependent feature maps to facilitate the segmentation. We evaluate our model on 2012 Brain Multi-Atlas Labelling Challenge dataset for 134 fine-grained structure segmentation. Besides, we validate our network on 27 coarse structure segmentation tasks. Experimental results have demonstrated that our model can achieve improved performance compared with the state-of-the-art approaches.
A deep learning model for early prediction of Alzheimer's disease dementia based on hippocampal MRI
Apr 15, 2019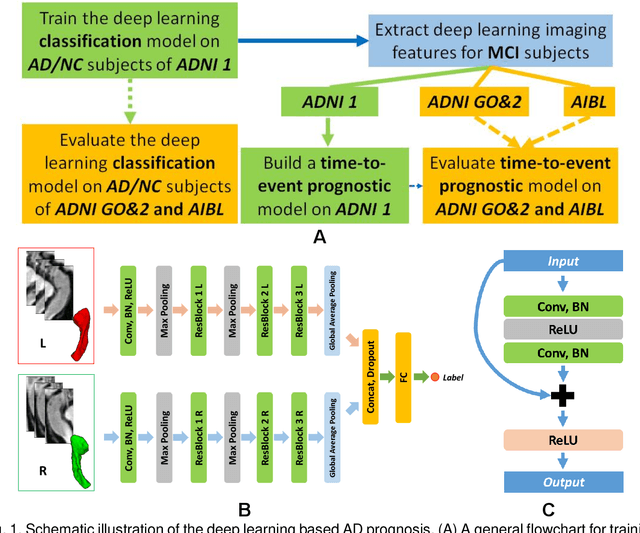
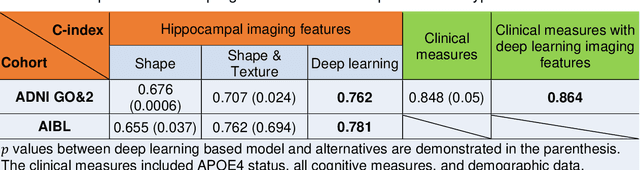
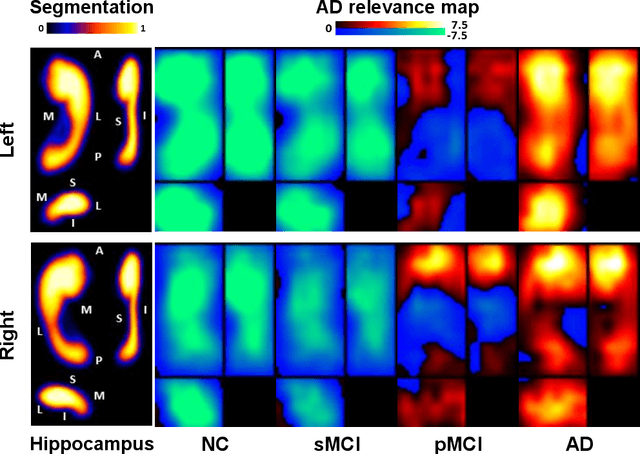
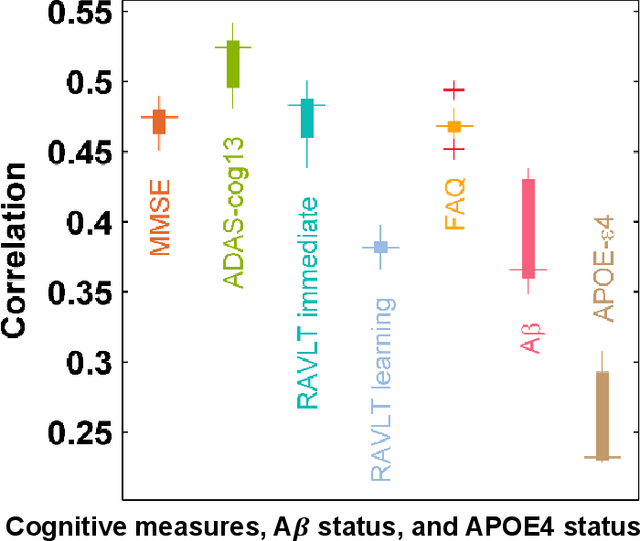
Abstract:Introduction: It is challenging at baseline to predict when and which individuals who meet criteria for mild cognitive impairment (MCI) will ultimately progress to Alzheimer's disease (AD) dementia. Methods: A deep learning method is developed and validated based on MRI scans of 2146 subjects (803 for training and 1343 for validation) to predict MCI subjects' progression to AD dementia in a time-to-event analysis setting. Results: The deep learning time-to-event model predicted individual subjects' progression to AD dementia with a concordance index (C-index) of 0.762 on 439 ADNI testing MCI subjects with follow-up duration from 6 to 78 months (quartiles: [24, 42, 54]) and a C-index of 0.781 on 40 AIBL testing MCI subjects with follow-up duration from 18-54 months (quartiles: [18, 36,54]). The predicted progression risk also clustered individual subjects into subgroups with significant differences in their progression time to AD dementia (p<0.0002). Improved performance for predicting progression to AD dementia (C-index=0.864) was obtained when the deep learning based progression risk was combined with baseline clinical measures. Conclusion: Our method provides a cost effective and accurate means for prognosis and potentially to facilitate enrollment in clinical trials with individuals likely to progress within a specific temporal period.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge