Yizhe Zhang
Data-Driven Deep Supervision for Skin Lesion Classification
Sep 04, 2022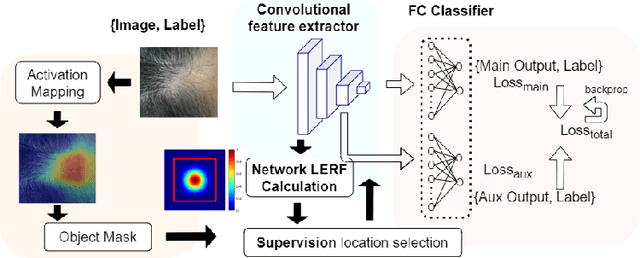
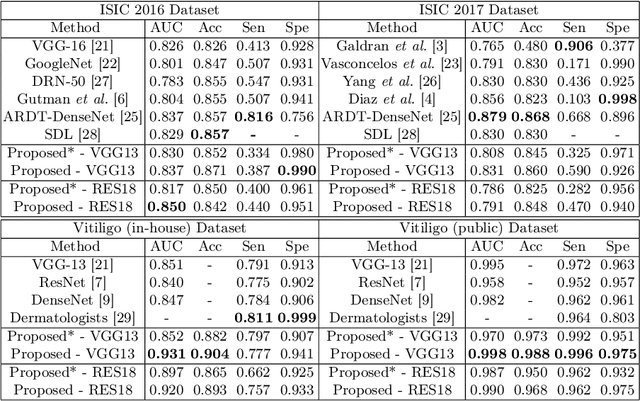
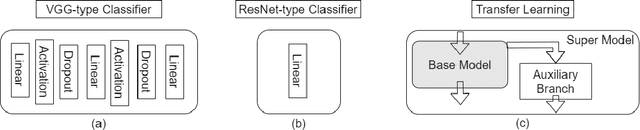
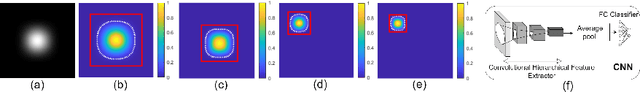
Abstract:Automatic classification of pigmented, non-pigmented, and depigmented non-melanocytic skin lesions have garnered lots of attention in recent years. However, imaging variations in skin texture, lesion shape, depigmentation contrast, lighting condition, etc. hinder robust feature extraction, affecting classification accuracy. In this paper, we propose a new deep neural network that exploits input data for robust feature extraction. Specifically, we analyze the convolutional network's behavior (field-of-view) to find the location of deep supervision for improved feature extraction. To achieve this, first, we perform activation mapping to generate an object mask, highlighting the input regions most critical for classification output generation. Then the network layer whose layer-wise effective receptive field matches the approximated object shape in the object mask is selected as our focus for deep supervision. Utilizing different types of convolutional feature extractors and classifiers on three melanoma detection datasets and two vitiligo detection datasets, we verify the effectiveness of our new method.
Usable Region Estimate for Assessing Practical Usability of Medical Image Segmentation Models
Jul 01, 2022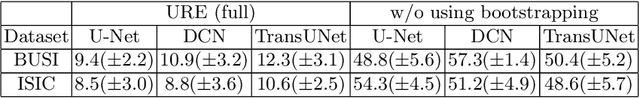
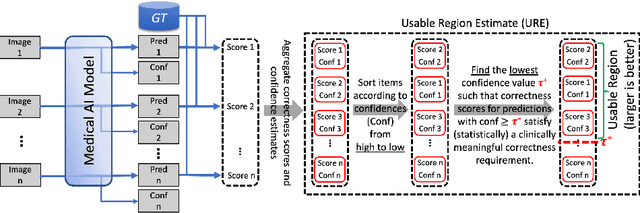
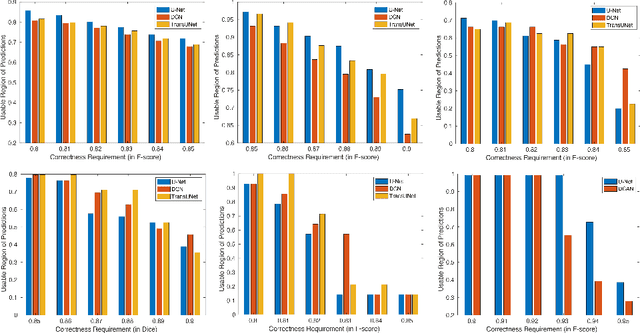
Abstract:We aim to quantitatively measure the practical usability of medical image segmentation models: to what extent, how often, and on which samples a model's predictions can be used/trusted. We first propose a measure, Correctness-Confidence Rank Correlation (CCRC), to capture how predictions' confidence estimates correlate with their correctness scores in rank. A model with a high value of CCRC means its prediction confidences reliably suggest which samples' predictions are more likely to be correct. Since CCRC does not capture the actual prediction correctness, it alone is insufficient to indicate whether a prediction model is both accurate and reliable to use in practice. Therefore, we further propose another method, Usable Region Estimate (URE), which simultaneously quantifies predictions' correctness and reliability of confidence assessments in one estimate. URE provides concrete information on to what extent a model's predictions are usable. In addition, the sizes of usable regions (UR) can be utilized to compare models: A model with a larger UR can be taken as a more usable and hence better model. Experiments on six datasets validate that the proposed evaluation methods perform well, providing a concrete and concise measure for the practical usability of medical image segmentation models. Code is made available at https://github.com/yizhezhang2000/ure.
H-EMD: A Hierarchical Earth Mover's Distance Method for Instance Segmentation
Jun 02, 2022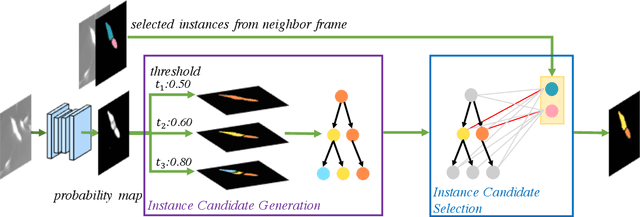
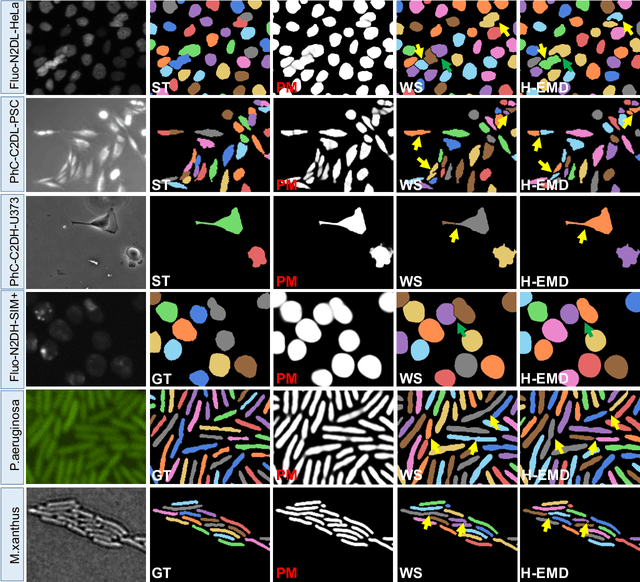
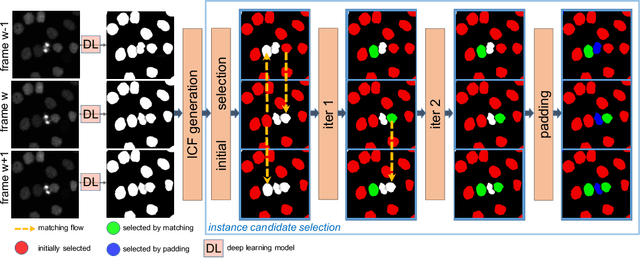
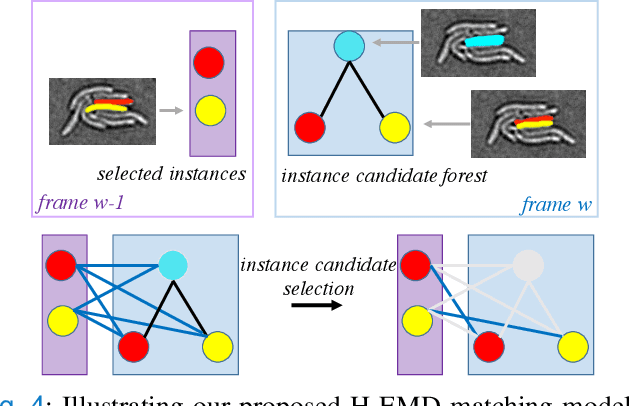

Abstract:Deep learning (DL) based semantic segmentation methods have achieved excellent performance in biomedical image segmentation, producing high quality probability maps to allow extraction of rich instance information to facilitate good instance segmentation. While numerous efforts were put into developing new DL semantic segmentation models, less attention was paid to a key issue of how to effectively explore their probability maps to attain the best possible instance segmentation. We observe that probability maps by DL semantic segmentation models can be used to generate many possible instance candidates, and accurate instance segmentation can be achieved by selecting from them a set of "optimized" candidates as output instances. Further, the generated instance candidates form a well-behaved hierarchical structure (a forest), which allows selecting instances in an optimized manner. Hence, we propose a novel framework, called hierarchical earth mover's distance (H-EMD), for instance segmentation in biomedical 2D+time videos and 3D images, which judiciously incorporates consistent instance selection with semantic-segmentation-generated probability maps. H-EMD contains two main stages. (1) Instance candidate generation: capturing instance-structured information in probability maps by generating many instance candidates in a forest structure. (2) Instance candidate selection: selecting instances from the candidate set for final instance segmentation. We formulate a key instance selection problem on the instance candidate forest as an optimization problem based on the earth mover's distance (EMD), and solve it by integer linear programming. Extensive experiments on eight biomedical video or 3D datasets demonstrate that H-EMD consistently boosts DL semantic segmentation models and is highly competitive with state-of-the-art methods.
Linearizing Transformer with Key-Value Memory Bank
Mar 26, 2022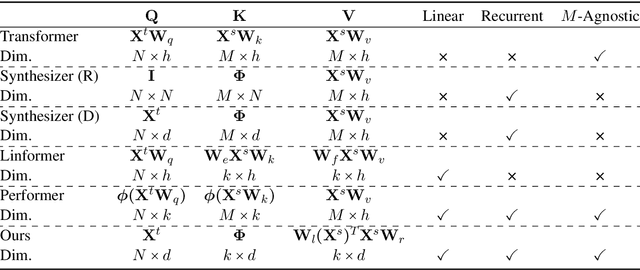
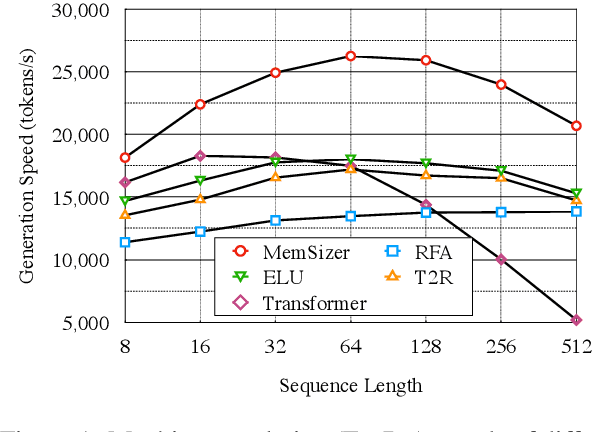
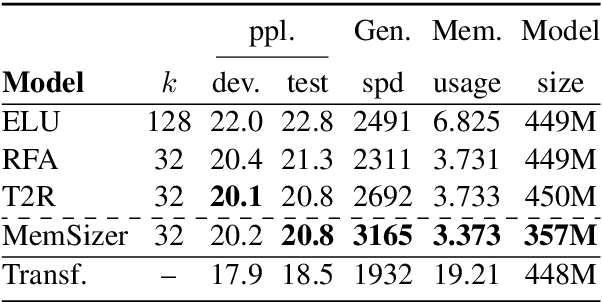
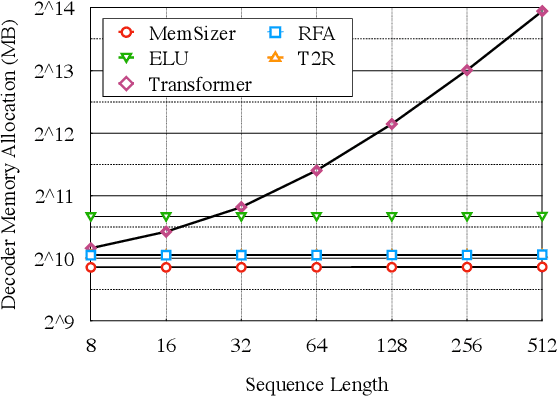
Abstract:Transformer has brought great success to a wide range of natural language processing tasks. Nevertheless, the computational overhead of the vanilla transformer scales quadratically with sequence length. Many efforts have been made to develop more efficient transformer variants. A line of work (e.g., Linformer) projects the input sequence into a low-rank space, achieving linear time complexity. However, Linformer does not suit well for text generation tasks as the sequence length must be pre-specified. We propose MemSizer, an approach also projects the source sequence into lower dimension representation but can take input with dynamic length, with a different perspective of the attention mechanism. MemSizer not only achieves the same linear time complexity but also enjoys efficient recurrent-style autoregressive generation, which yields constant memory complexity and reduced computation at inference. We demonstrate that MemSizer provides an improved tradeoff between efficiency and accuracy over the vanilla transformer and other linear variants in language modeling and machine translation tasks, revealing a viable direction towards further inference efficiency improvement.
Report from the NSF Future Directions Workshop on Automatic Evaluation of Dialog: Research Directions and Challenges
Mar 18, 2022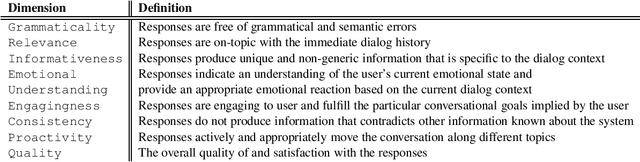

Abstract:This is a report on the NSF Future Directions Workshop on Automatic Evaluation of Dialog. The workshop explored the current state of the art along with its limitations and suggested promising directions for future work in this important and very rapidly changing area of research.
Towards More Efficient Insertion Transformer with Fractional Positional Encoding
Dec 12, 2021
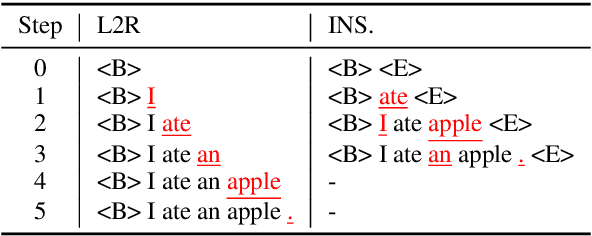
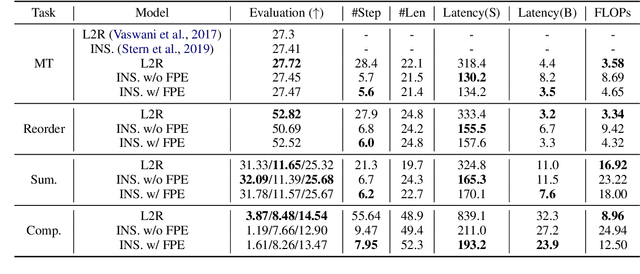
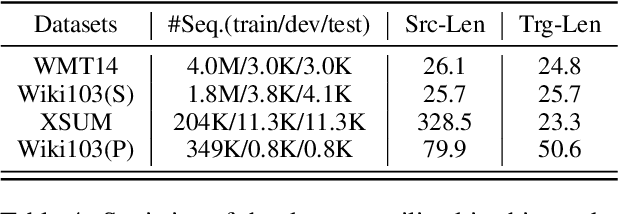
Abstract:Auto-regressive neural sequence models have been shown to be effective across text generation tasks. However, their left-to-right decoding order prevents generation from being parallelized. Insertion Transformer (Stern et al., 2019) is an attractive alternative that allows outputting multiple tokens in a single generation step. Nevertheless, due to the incompatibility of absolute positional encoding and insertion-based generation schemes, it needs to refresh the encoding of every token in the generated partial hypotheses at each step, which could be costly. We design a novel incremental positional encoding scheme for insertion transformers called Fractional Positional Encoding (FPE), which allows reusing representations calculated in previous steps. Empirical studies on various language generation tasks demonstrate the effectiveness of FPE, which leads to reduction of floating point operations and latency improvements on batched decoding.
HS3: Learning with Proper Task Complexity in Hierarchically Supervised Semantic Segmentation
Nov 03, 2021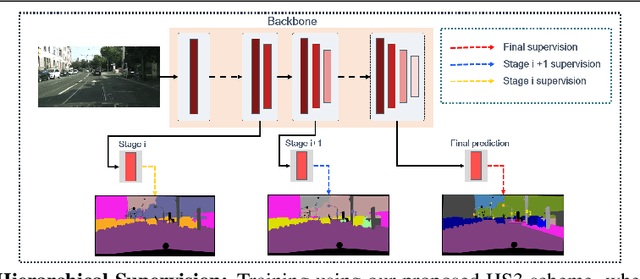
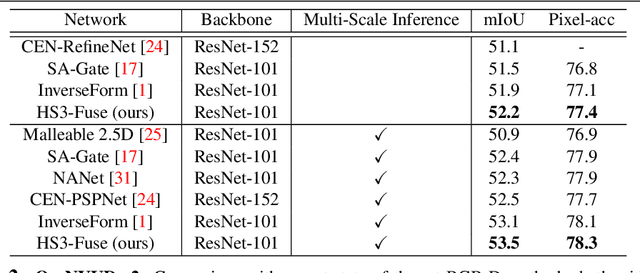
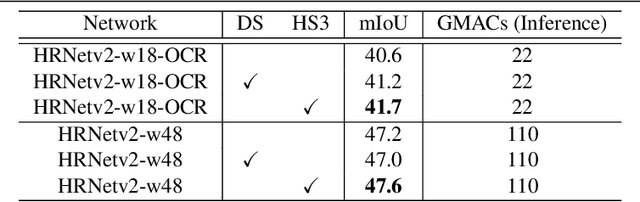
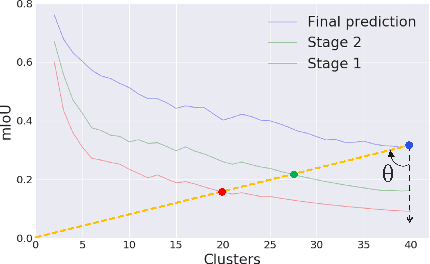

Abstract:While deeply supervised networks are common in recent literature, they typically impose the same learning objective on all transitional layers despite their varying representation powers. In this paper, we propose Hierarchically Supervised Semantic Segmentation (HS3), a training scheme that supervises intermediate layers in a segmentation network to learn meaningful representations by varying task complexity. To enforce a consistent performance vs. complexity trade-off throughout the network, we derive various sets of class clusters to supervise each transitional layer of the network. Furthermore, we devise a fusion framework, HS3-Fuse, to aggregate the hierarchical features generated by these layers, which can provide rich semantic contexts and further enhance the final segmentation. Extensive experiments show that our proposed HS3 scheme considerably outperforms vanilla deep supervision with no added inference cost. Our proposed HS3-Fuse framework further improves segmentation predictions and achieves state-of-the-art results on two large segmentation benchmarks: NYUD-v2 and Cityscapes.
X-Distill: Improving Self-Supervised Monocular Depth via Cross-Task Distillation
Oct 24, 2021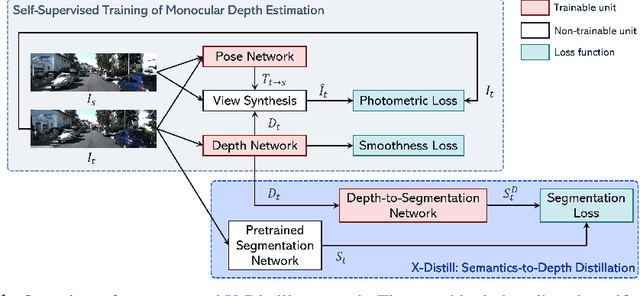
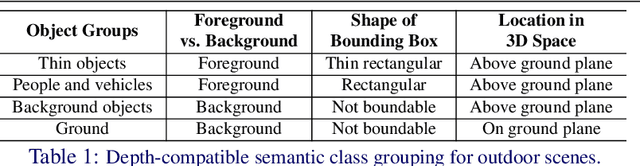

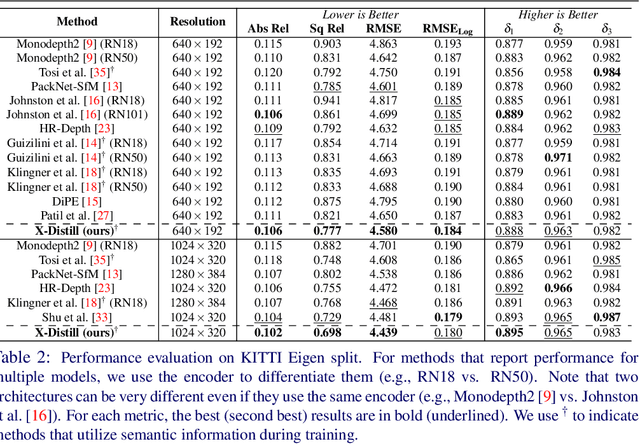
Abstract:In this paper, we propose a novel method, X-Distill, to improve the self-supervised training of monocular depth via cross-task knowledge distillation from semantic segmentation to depth estimation. More specifically, during training, we utilize a pretrained semantic segmentation teacher network and transfer its semantic knowledge to the depth network. In order to enable such knowledge distillation across two different visual tasks, we introduce a small, trainable network that translates the predicted depth map to a semantic segmentation map, which can then be supervised by the teacher network. In this way, this small network enables the backpropagation from the semantic segmentation teacher's supervision to the depth network during training. In addition, since the commonly used object classes in semantic segmentation are not directly transferable to depth, we study the visual and geometric characteristics of the objects and design a new way of grouping them that can be shared by both tasks. It is noteworthy that our approach only modifies the training process and does not incur additional computation during inference. We extensively evaluate the efficacy of our proposed approach on the standard KITTI benchmark and compare it with the latest state of the art. We further test the generalizability of our approach on Make3D. Overall, the results show that our approach significantly improves the depth estimation accuracy and outperforms the state of the art.
Perceptual Consistency in Video Segmentation
Oct 24, 2021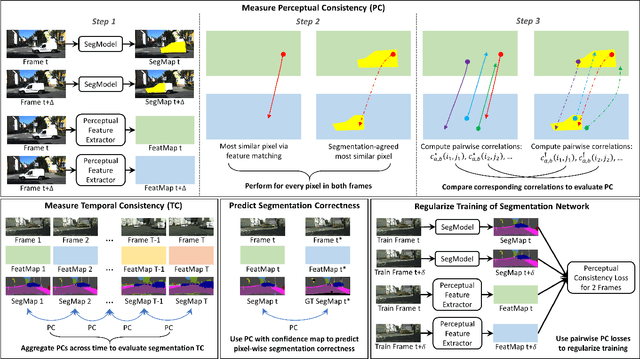
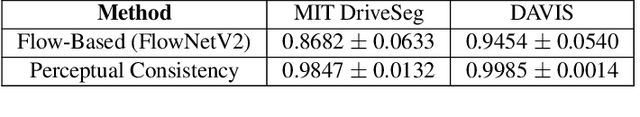
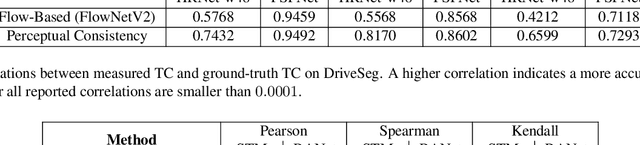
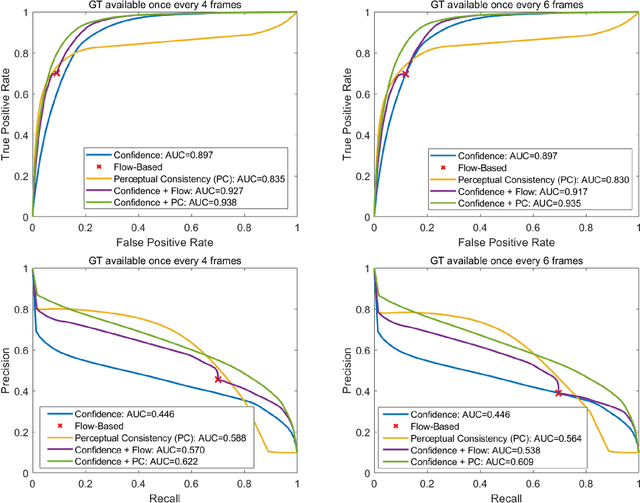
Abstract:In this paper, we present a novel perceptual consistency perspective on video semantic segmentation, which can capture both temporal consistency and pixel-wise correctness. Given two nearby video frames, perceptual consistency measures how much the segmentation decisions agree with the pixel correspondences obtained via matching general perceptual features. More specifically, for each pixel in one frame, we find the most perceptually correlated pixel in the other frame. Our intuition is that such a pair of pixels are highly likely to belong to the same class. Next, we assess how much the segmentation agrees with such perceptual correspondences, based on which we derive the perceptual consistency of the segmentation maps across these two frames. Utilizing perceptual consistency, we can evaluate the temporal consistency of video segmentation by measuring the perceptual consistency over consecutive pairs of segmentation maps in a video. Furthermore, given a sparsely labeled test video, perceptual consistency can be utilized to aid with predicting the pixel-wise correctness of the segmentation on an unlabeled frame. More specifically, by measuring the perceptual consistency between the predicted segmentation and the available ground truth on a nearby frame and combining it with the segmentation confidence, we can accurately assess the classification correctness on each pixel. Our experiments show that the proposed perceptual consistency can more accurately evaluate the temporal consistency of video segmentation as compared to flow-based measures. Furthermore, it can help more confidently predict segmentation accuracy on unlabeled test frames, as compared to using classification confidence alone. Finally, our proposed measure can be used as a regularizer during the training of segmentation models, which leads to more temporally consistent video segmentation while maintaining accuracy.
AuxAdapt: Stable and Efficient Test-Time Adaptation for Temporally Consistent Video Semantic Segmentation
Oct 24, 2021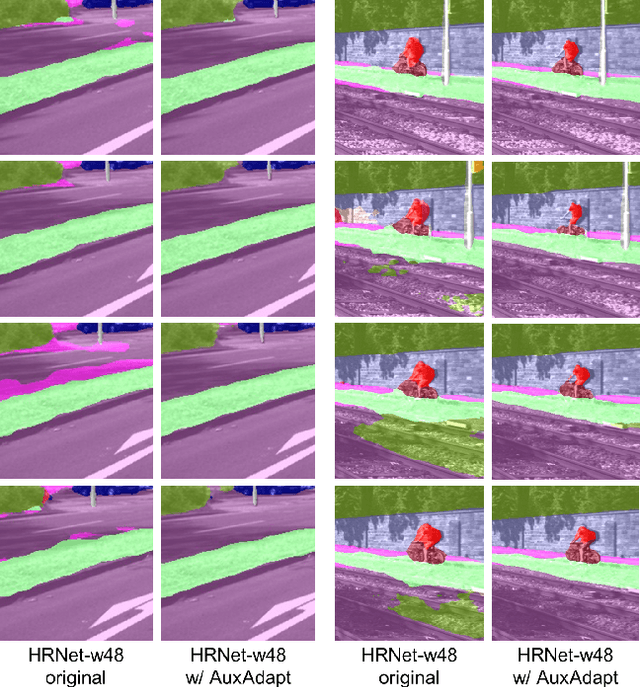
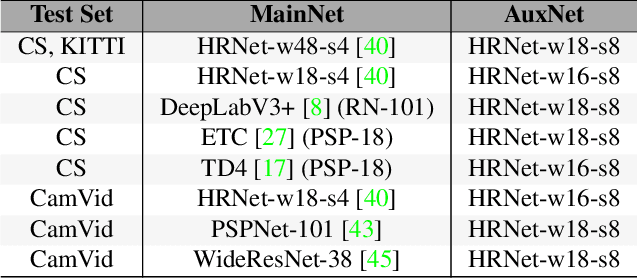
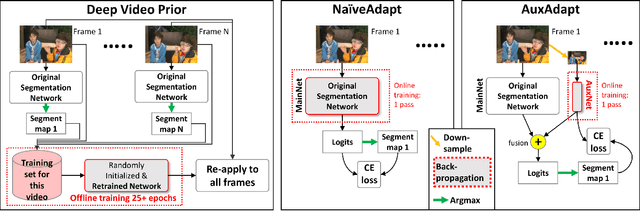
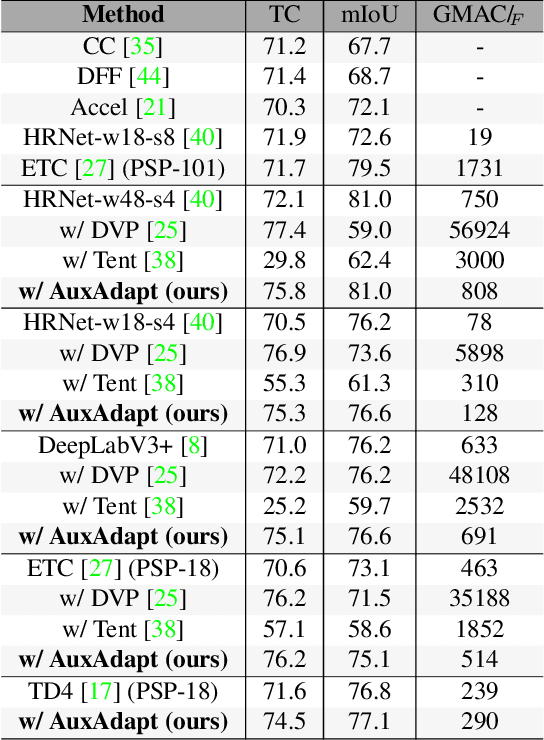
Abstract:In video segmentation, generating temporally consistent results across frames is as important as achieving frame-wise accuracy. Existing methods rely either on optical flow regularization or fine-tuning with test data to attain temporal consistency. However, optical flow is not always avail-able and reliable. Besides, it is expensive to compute. Fine-tuning the original model in test time is cost sensitive. This paper presents an efficient, intuitive, and unsupervised online adaptation method, AuxAdapt, for improving the temporal consistency of most neural network models. It does not require optical flow and only takes one pass of the video. Since inconsistency mainly arises from the model's uncertainty in its output, we propose an adaptation scheme where the model learns from its own segmentation decisions as it streams a video, which allows producing more confident and temporally consistent labeling for similarly-looking pixels across frames. For stability and efficiency, we leverage a small auxiliary segmentation network (AuxNet) to assist with this adaptation. More specifically, AuxNet readjusts the decision of the original segmentation network (Main-Net) by adding its own estimations to that of MainNet. At every frame, only AuxNet is updated via back-propagation while keeping MainNet fixed. We extensively evaluate our test-time adaptation approach on standard video benchmarks, including Cityscapes, CamVid, and KITTI. The results demonstrate that our approach provides label-wise accurate, temporally consistent, and computationally efficient adaptation (5+ folds overhead reduction comparing to state-of-the-art test-time adaptation methods).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge