Yiqin Yu
Medical visual question answering using joint self-supervised learning
Feb 25, 2023



Abstract:Visual Question Answering (VQA) becomes one of the most active research problems in the medical imaging domain. A well-known VQA challenge is the intrinsic diversity between the image and text modalities, and in the medical VQA task, there is another critical problem relying on the limited size of labelled image-question-answer data. In this study we propose an encoder-decoder framework that leverages the image-text joint representation learned from large-scaled medical image-caption data and adapted to the small-sized medical VQA task. The encoder embeds across the image-text dual modalities with self-attention mechanism and is independently pre-trained on the large-scaled medical image-caption dataset by multiple self-supervised learning tasks. Then the decoder is connected to the top of the encoder and fine-tuned using the small-sized medical VQA dataset. The experiment results present that our proposed method achieves better performance comparing with the baseline and SOTA methods.
Adversarial Sample Enhanced Domain Adaptation: A Case Study on Predictive Modeling with Electronic Health Records
Jan 13, 2021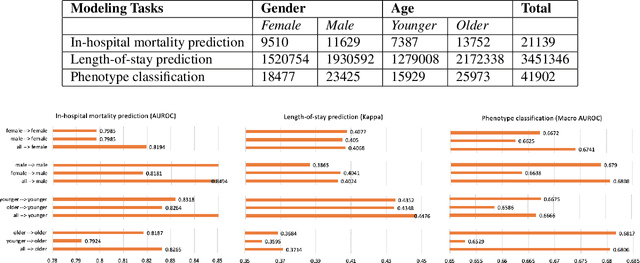
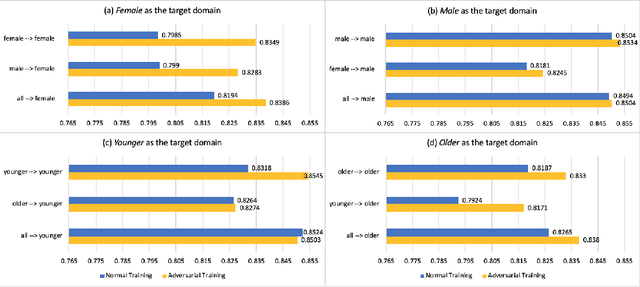
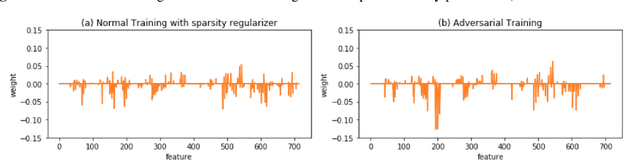
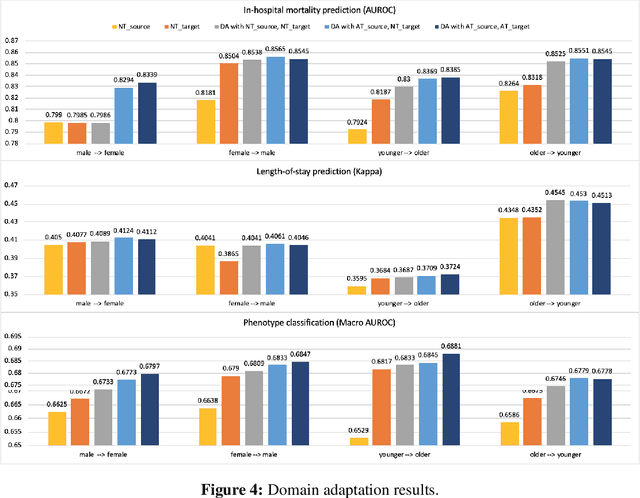
Abstract:With the successful adoption of machine learning on electronic health records (EHRs), numerous computational models have been deployed to address a variety of clinical problems. However, due to the heterogeneity of EHRs, models trained on different patient groups suffer from poor generalizability. How to mitigate domain shifts between the source patient group where the model is built upon and the target one where the model will be deployed becomes a critical issue. In this paper, we propose a data augmentation method to facilitate domain adaptation, which leverages knowledge from the source patient group when training model on the target one. Specifically, adversarially generated samples are used during domain adaptation to fill the generalization gap between the two patient groups. The proposed method is evaluated by a case study on different predictive modeling tasks on MIMIC-III EHR dataset. Results confirm the effectiveness of our method and the generality on different tasks.
Dynamic Knowledge Distillation for Black-box Hypothesis Transfer Learning
Aug 07, 2020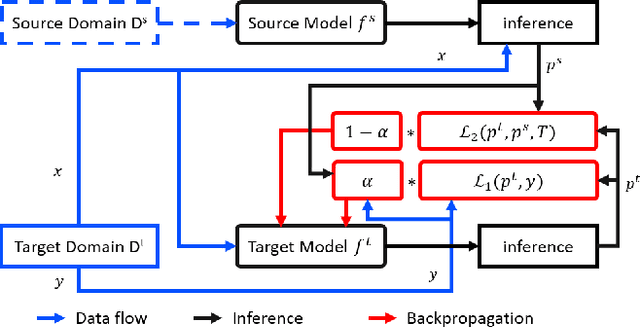
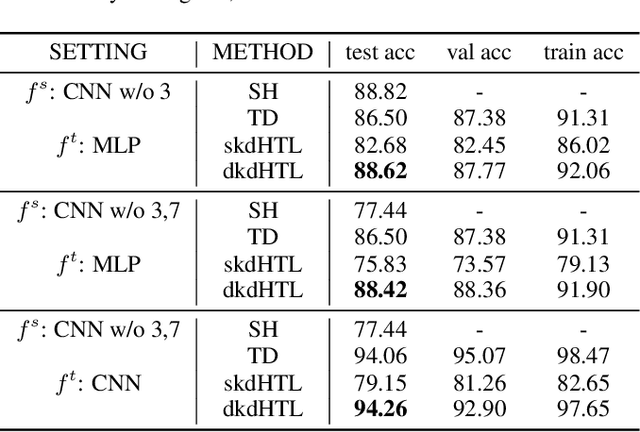
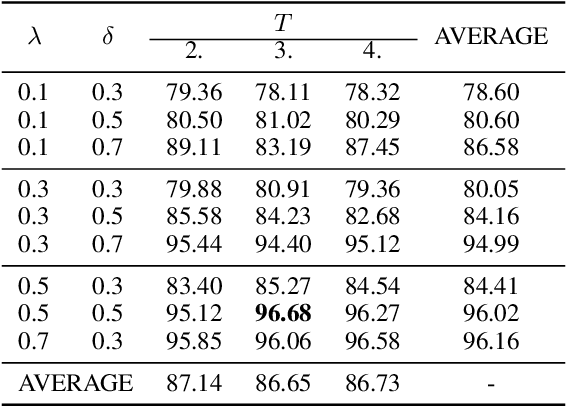

Abstract:In real world applications like healthcare, it is usually difficult to build a machine learning prediction model that works universally well across different institutions. At the same time, the available model is often proprietary, i.e., neither the model parameter nor the data set used for model training is accessible. In consequence, leveraging the knowledge hidden in the available model (aka. the hypothesis) and adapting it to a local data set becomes extremely challenging. Motivated by this situation, in this paper we aim to address such a specific case within the hypothesis transfer learning framework, in which 1) the source hypothesis is a black-box model and 2) the source domain data is unavailable. In particular, we introduce a novel algorithm called dynamic knowledge distillation for hypothesis transfer learning (dkdHTL). In this method, we use knowledge distillation with instance-wise weighting mechanism to adaptively transfer the "dark" knowledge from the source hypothesis to the target domain.The weighting coefficients of the distillation loss and the standard loss are determined by the consistency between the predicted probability of the source hypothesis and the target ground-truth label.Empirical results on both transfer learning benchmark datasets and a healthcare dataset demonstrate the effectiveness of our method.
Unlocking the Power of Deep PICO Extraction: Step-wise Medical NER Identification
Apr 30, 2020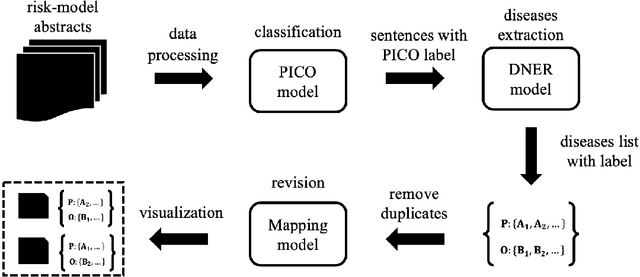
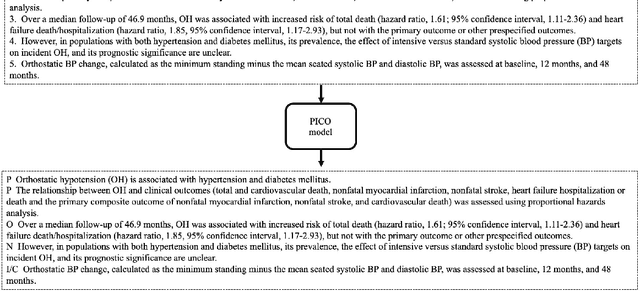
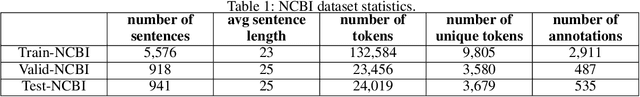

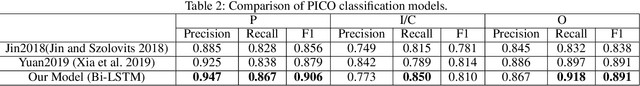
Abstract:The PICO framework (Population, Intervention, Comparison, and Outcome) is usually used to formulate evidence in the medical domain. The major task of PICO extraction is to extract sentences from medical literature and classify them into each class. However, in most circumstances, there will be more than one evidences in an extracted sentence even it has been categorized to a certain class. In order to address this problem, we propose a step-wise disease Named Entity Recognition (DNER) extraction and PICO identification method. With our method, sentences in paper title and abstract are first classified into different classes of PICO, and medical entities are then identified and classified into P and O. Different kinds of deep learning frameworks are used and experimental results show that our method will achieve high performance and fine-grained extraction results comparing with conventional PICO extraction works.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge