Yefeng Zheng
Deep Convolutional Neural Networks for Molecular Subtyping of Gliomas Using Magnetic Resonance Imaging
Mar 10, 2022Abstract:Knowledge of molecular subtypes of gliomas can provide valuable information for tailored therapies. This study aimed to investigate the use of deep convolutional neural networks (DCNNs) for noninvasive glioma subtyping with radiological imaging data according to the new taxonomy announced by the World Health Organization in 2016. Methods: A DCNN model was developed for the prediction of the five glioma subtypes based on a hierarchical classification paradigm. This model used three parallel, weight-sharing, deep residual learning networks to process 2.5-dimensional input of trimodal MRI data, including T1-weighted, T1-weighted with contrast enhancement, and T2-weighted images. A data set comprising 1,016 real patients was collected for evaluation of the developed DCNN model. The predictive performance was evaluated via the area under the curve (AUC) from the receiver operating characteristic analysis. For comparison, the performance of a radiomics-based approach was also evaluated. Results: The AUCs of the DCNN model for the four classification tasks in the hierarchical classification paradigm were 0.89, 0.89, 0.85, and 0.66, respectively, as compared to 0.85, 0.75, 0.67, and 0.59 of the radiomics approach. Conclusion: The results showed that the developed DCNN model can predict glioma subtypes with promising performance, given sufficient, non-ill-balanced training data.
Conquering Data Variations in Resolution: A Slice-Aware Multi-Branch Decoder Network
Mar 07, 2022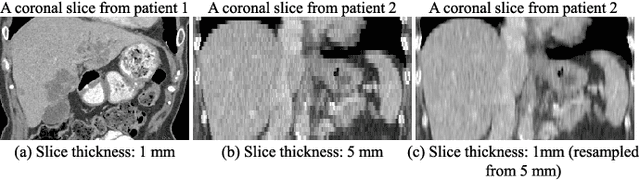
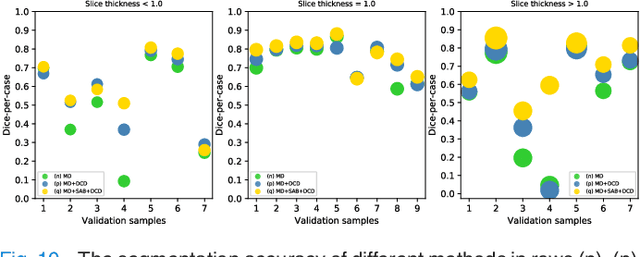
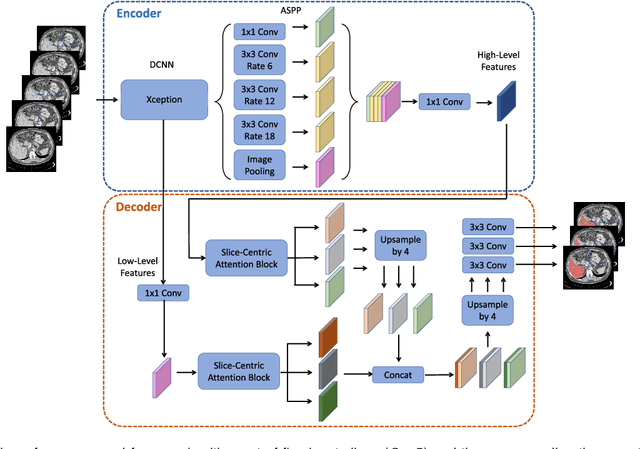
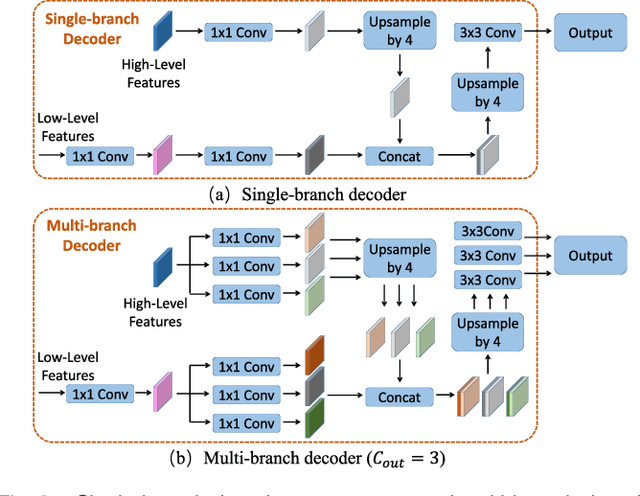
Abstract:Fully convolutional neural networks have made promising progress in joint liver and liver tumor segmentation. Instead of following the debates over 2D versus 3D networks (for example, pursuing the balance between large-scale 2D pretraining and 3D context), in this paper, we novelly identify the wide variation in the ratio between intra- and inter-slice resolutions as a crucial obstacle to the performance. To tackle the mismatch between the intra- and inter-slice information, we propose a slice-aware 2.5D network that emphasizes extracting discriminative features utilizing not only in-plane semantics but also out-of-plane coherence for each separate slice. Specifically, we present a slice-wise multi-input multi-output architecture to instantiate such a design paradigm, which contains a Multi-Branch Decoder (MD) with a Slice-centric Attention Block (SAB) for learning slice-specific features and a Densely Connected Dice (DCD) loss to regularize the inter-slice predictions to be coherent and continuous. Based on the aforementioned innovations, we achieve state-of-the-art results on the MICCAI 2017 Liver Tumor Segmentation (LiTS) dataset. Besides, we also test our model on the ISBI 2019 Segmentation of THoracic Organs at Risk (SegTHOR) dataset, and the result proves the robustness and generalizability of the proposed method in other segmentation tasks.
Simultaneous Alignment and Surface Regression Using Hybrid 2D-3D Networks for 3D Coherent Layer Segmentation of Retina OCT Images
Mar 04, 2022

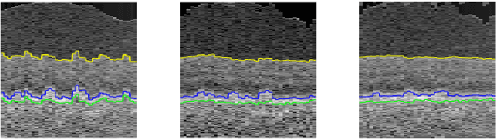
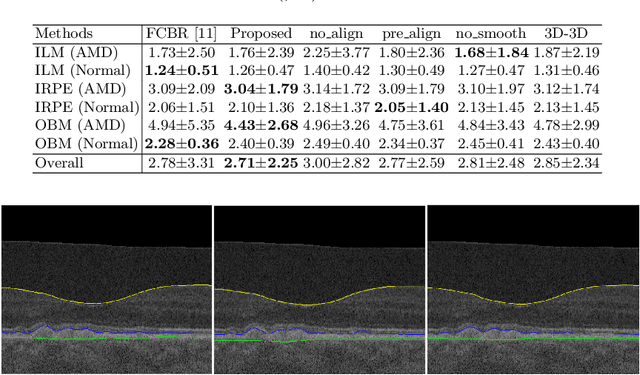
Abstract:Automated surface segmentation of retinal layer is important and challenging in analyzing optical coherence tomography (OCT). Recently, many deep learning based methods have been developed for this task and yield remarkable performance. However, due to large spatial gap and potential mismatch between the B-scans of OCT data, all of them are based on 2D segmentation of individual B-scans, which may loss the continuity information across the B-scans. In addition, 3D surface of the retina layers can provide more diagnostic information, which is crucial in quantitative image analysis. In this study, a novel framework based on hybrid 2D-3D convolutional neural networks (CNNs) is proposed to obtain continuous 3D retinal layer surfaces from OCT. The 2D features of individual B-scans are extracted by an encoder consisting of 2D convolutions. These 2D features are then used to produce the alignment displacement field and layer segmentation by two 3D decoders, which are coupled via a spatial transformer module. The entire framework is trained end-to-end. To the best of our knowledge, this is the first study that attempts 3D retinal layer segmentation in volumetric OCT images based on CNNs. Experiments on a publicly available dataset show that our framework achieves superior results to state-of-the-art 2D methods in terms of both layer segmentation accuracy and cross-B-scan 3D continuity, thus offering more clinical values than previous works.
Label Propagation for Annotation-Efficient Nuclei Segmentation from Pathology Images
Feb 16, 2022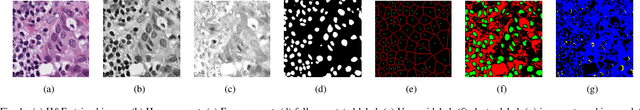
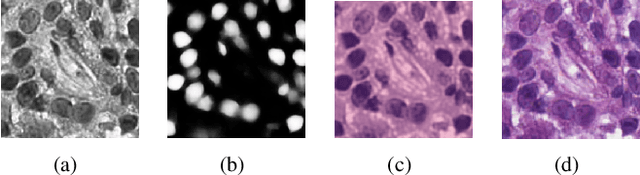
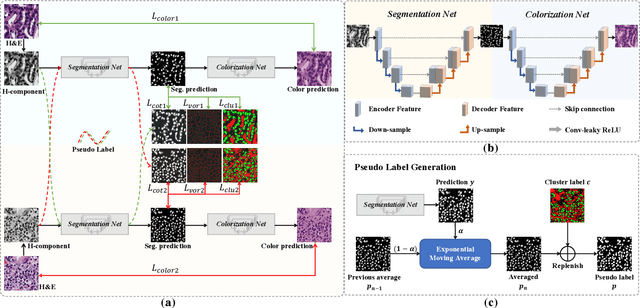
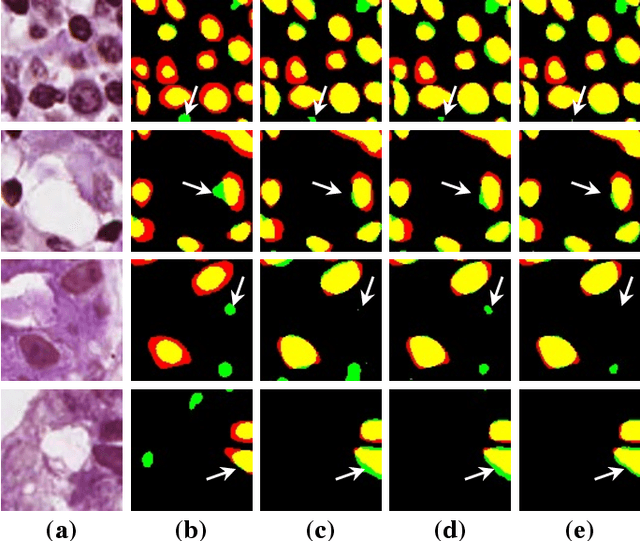

Abstract:Nuclei segmentation is a crucial task for whole slide image analysis in digital pathology. Generally, the segmentation performance of fully-supervised learning heavily depends on the amount and quality of the annotated data. However, it is time-consuming and expensive for professional pathologists to provide accurate pixel-level ground truth, while it is much easier to get coarse labels such as point annotations. In this paper, we propose a weakly-supervised learning method for nuclei segmentation that only requires point annotations for training. The proposed method achieves label propagation in a coarse-to-fine manner as follows. First, coarse pixel-level labels are derived from the point annotations based on the Voronoi diagram and the k-means clustering method to avoid overfitting. Second, a co-training strategy with an exponential moving average method is designed to refine the incomplete supervision of the coarse labels. Third, a self-supervised visual representation learning method is tailored for nuclei segmentation of pathology images that transforms the hematoxylin component images into the H\&E stained images to gain better understanding of the relationship between the nuclei and cytoplasm. We comprehensively evaluate the proposed method using two public datasets. Both visual and quantitative results demonstrate the superiority of our method to the state-of-the-art methods, and its competitive performance compared to the fully-supervised methods. The source codes for implementing the experiments will be released after acceptance.
FedMed-ATL: Misaligned Unpaired Brain Image Synthesis via Affine Transform Loss
Feb 16, 2022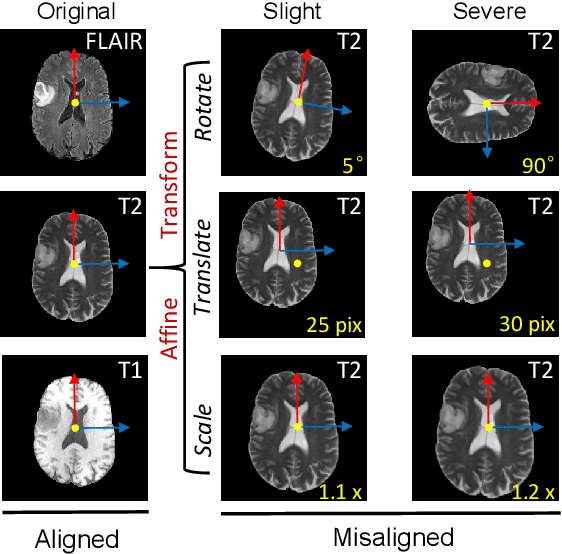
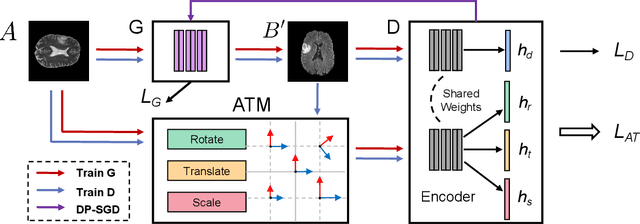
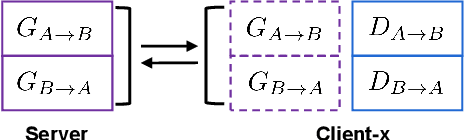
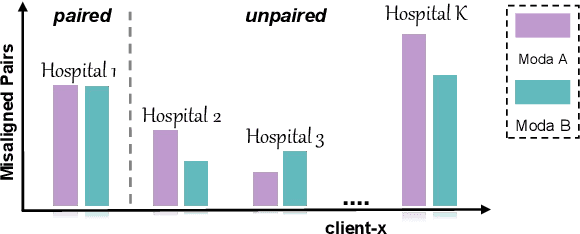
Abstract:The existence of completely aligned and paired multi-modal neuroimaging data has proved its effectiveness in the diagnosis of brain diseases. However, collecting the full set of well-aligned and paired data is impractical or even luxurious, since the practical difficulties may include high cost, long time acquisition, image corruption, and privacy issues. Previously, the misaligned unpaired neuroimaging data (termed as MUD) are generally treated as noisy label. However, such a noisy label-based method could not work very well when misaligned data occurs distortions severely, for example, different angles of rotation. In this paper, we propose a novel federated self-supervised learning (FedMed) for brain image synthesis. An affine transform loss (ATL) was formulated to make use of severely distorted images without violating privacy legislation for the hospital. We then introduce a new data augmentation procedure for self-supervised training and fed it into three auxiliary heads, namely auxiliary rotation, auxiliary translation, and auxiliary scaling heads. The proposed method demonstrates advanced performance in both the quality of synthesized results under a severely misaligned and unpaired data setting, and better stability than other GAN-based algorithms. The proposed method also reduces the demand for deformable registration while encouraging to realize the usage of those misaligned and unpaired data. Experimental results verify the outstanding ability of our learning paradigm compared to other state-of-the-art approaches. Our code is available on the website: https://github.com/FedMed-Meta/FedMed-ATL
A Survey of Cross-Modality Brain Image Synthesis
Feb 16, 2022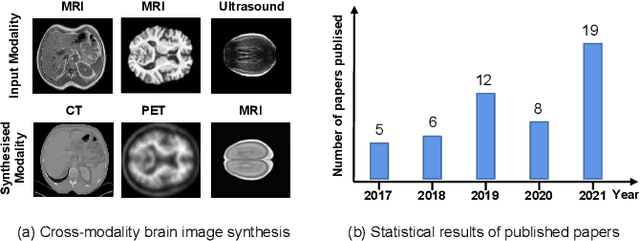
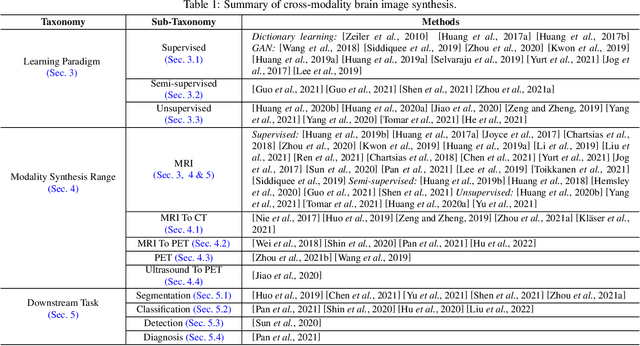
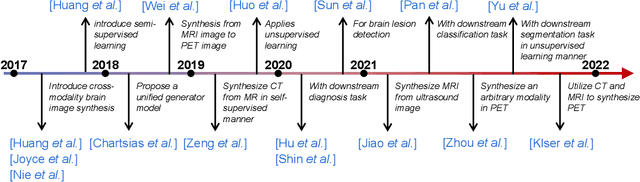
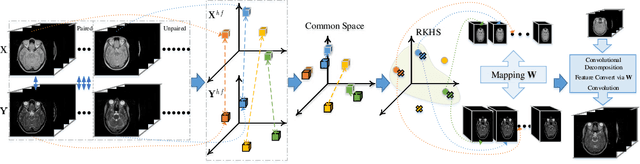
Abstract:The existence of completely aligned and paired multi-modal neuroimaging data has proved its effectiveness in diagnosis of brain diseases. However, collecting the full set of well-aligned and paired data is impractical or even luxurious, since the practical difficulties may include high cost, long time acquisition, image corruption, and privacy issues. A realistic solution is to explore either an unsupervised learning or a semi-supervised learning to synthesize the absent neuroimaging data. In this paper, we tend to approach multi-modality brain image synthesis task from different perspectives, which include the level of supervision, the range of modality synthesis, and the synthesis-based downstream tasks. Particularly, we provide in-depth analysis on how cross-modality brain image synthesis can improve the performance of different downstream tasks. Finally, we evaluate the challenges and provide several open directions for this community. All resources are available at https://github.com/M-3LAB/awesome-multimodal-brain-image-systhesis
FedMed-GAN: Federated Multi-Modal Unsupervised Brain Image Synthesis
Jan 22, 2022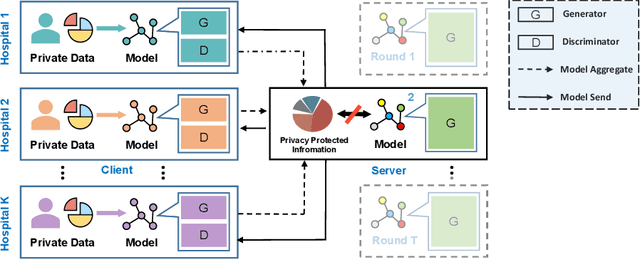

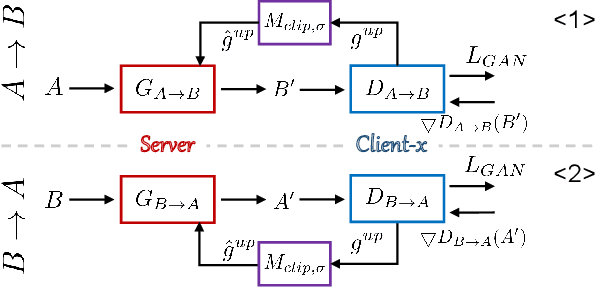
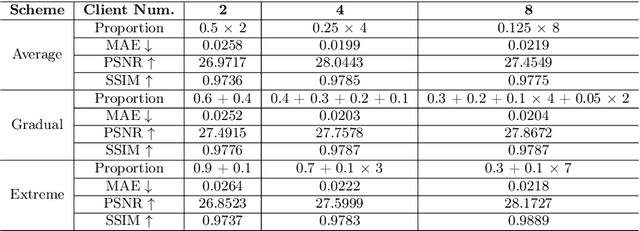
Abstract:Utilizing the paired multi-modal neuroimaging data has been proved to be effective to investigate human cognitive activities and certain pathologies. However, it is not practical to obtain the full set of paired neuroimaging data centrally since the collection faces several constraints, e.g., high examination costs, long acquisition time, and even image corruption. In addition, most of the paired neuroimaging data are dispersed into different medical institutions and cannot group together for centralized training considering the privacy issues. Under the circumstance, there is a clear need to launch federated learning and facilitate the integration of other unpaired data from different hospitals or data owners. In this paper, we build up a new benchmark for federated multi-modal unsupervised brain image synthesis (termed as FedMed-GAN) to bridge the gap between federated learning and medical GAN. Moreover, based on the similarity of edge information across multi-modal neuroimaging data, we propose a novel edge loss to solve the generative mode collapse issue of FedMed-GAN and mitigate the performance drop resulting from differential privacy. Compared with the state-of-the-art method shown in our built benchmark, our novel edge loss could significantly speed up the generator convergence rate without sacrificing performance under different unpaired data distribution settings.
Revisiting Experience Replay: Continual Learning by Adaptively Tuning Task-wise Relationship
Jan 06, 2022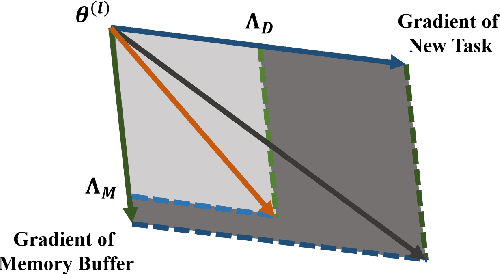
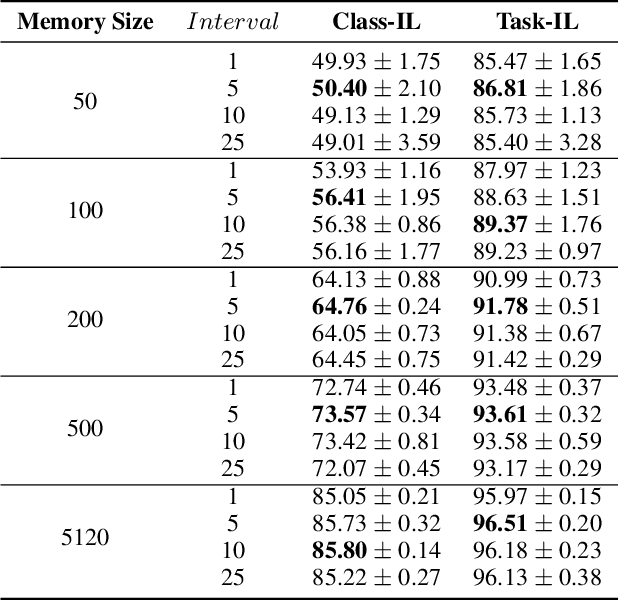
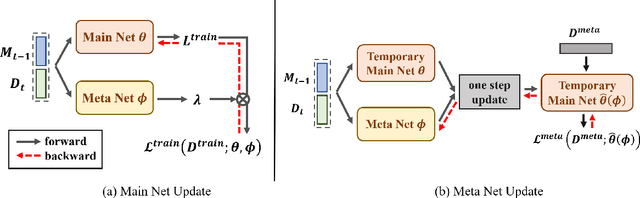
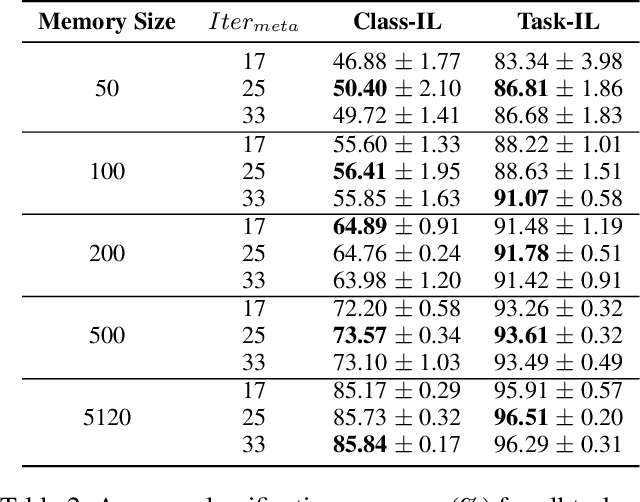
Abstract:Continual learning requires models to learn new tasks while maintaining previously learned knowledge. Various algorithms have been proposed to address this real challenge. Till now, rehearsal-based methods, such as experience replay, have achieved state-of-the-art performance. These approaches save a small part of the data of the past tasks as a memory buffer to prevent models from forgetting previously learned knowledge. However, most of them treat every new task equally, i.e., fixed the hyperparameters of the framework while learning different new tasks. Such a setting lacks the consideration of the relationship/similarity between past and new tasks. For example, the previous knowledge/features learned from dogs are more beneficial for the identification of cats (new task), compared to those learned from buses. In this regard, we propose a meta learning algorithm based on bi-level optimization to adaptively tune the relationship between the knowledge extracted from the past and new tasks. Therefore, the model can find an appropriate direction of gradient during continual learning and avoid the serious overfitting problem on memory buffer. Extensive experiments are conducted on three publicly available datasets (i.e., CIFAR-10, CIFAR-100, and Tiny ImageNet). The experimental results demonstrate that the proposed method can consistently improve the performance of all baselines.
InDuDoNet+: A Model-Driven Interpretable Dual Domain Network for Metal Artifact Reduction in CT Images
Dec 23, 2021



Abstract:During the computed tomography (CT) imaging process, metallic implants within patients always cause harmful artifacts, which adversely degrade the visual quality of reconstructed CT images and negatively affect the subsequent clinical diagnosis. For the metal artifact reduction (MAR) task, current deep learning based methods have achieved promising performance. However, most of them share two main common limitations: 1) the CT physical imaging geometry constraint is not comprehensively incorporated into deep network structures; 2) the entire framework has weak interpretability for the specific MAR task; hence, the role of every network module is difficult to be evaluated. To alleviate these issues, in the paper, we construct a novel interpretable dual domain network, termed InDuDoNet+, into which CT imaging process is finely embedded. Concretely, we derive a joint spatial and Radon domain reconstruction model and propose an optimization algorithm with only simple operators for solving it. By unfolding the iterative steps involved in the proposed algorithm into the corresponding network modules, we easily build the InDuDoNet+ with clear interpretability. Furthermore, we analyze the CT values among different tissues, and merge the prior observations into a prior network for our InDuDoNet+, which significantly improve its generalization performance. Comprehensive experiments on synthesized data and clinical data substantiate the superiority of the proposed methods as well as the superior generalization performance beyond the current state-of-the-art (SOTA) MAR methods. Code is available at \url{https://github.com/hongwang01/InDuDoNet_plus}.
Alleviating Noisy-label Effects in Image Classification via Probability Transition Matrix
Oct 19, 2021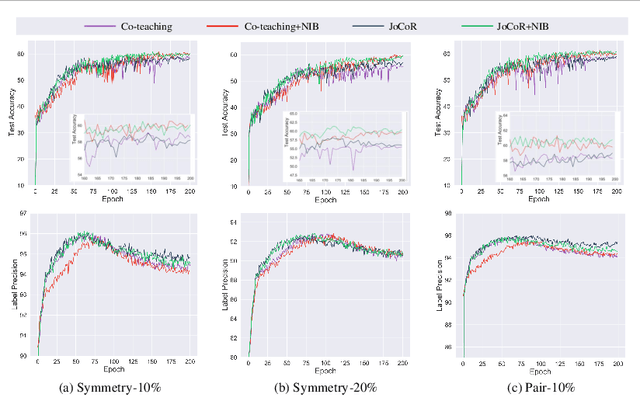
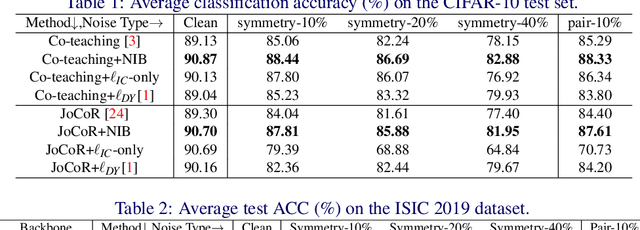
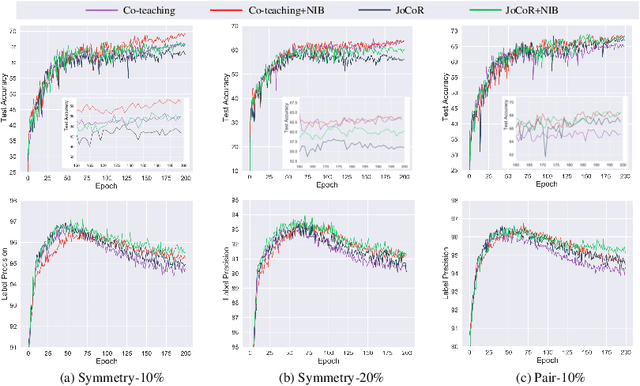
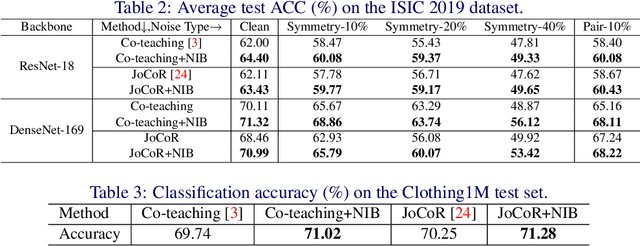
Abstract:Deep-learning-based image classification frameworks often suffer from the noisy label problem caused by the inter-observer variation. Recent studies employed learning-to-learn paradigms (e.g., Co-teaching and JoCoR) to filter the samples with noisy labels from the training set. However, most of them use a simple cross-entropy loss as the criterion for noisy label identification. The hard samples, which are beneficial for classifier learning, are often mistakenly treated as noises in such a setting since both the hard samples and ones with noisy labels lead to a relatively larger loss value than the easy cases. In this paper, we propose a plugin module, namely noise ignoring block (NIB), consisting of a probability transition matrix and an inter-class correlation (IC) loss, to separate the hard samples from the mislabeled ones, and further boost the accuracy of image classification network trained with noisy labels. Concretely, our IC loss is calculated as Kullback-Leibler divergence between the network prediction and the accumulative soft label generated by the probability transition matrix. Such that, with the lower value of IC loss, the hard cases can be easily distinguished from mislabeled cases. Extensive experiments are conducted on natural and medical image datasets (CIFAR-10 and ISIC 2019). The experimental results show that our NIB module consistently improves the performances of the state-of-the-art robust training methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge