Vince Lyzinski
The Importance of Being Correlated: Implications of Dependence in Joint Spectral Inference across Multiple Networks
Aug 01, 2020
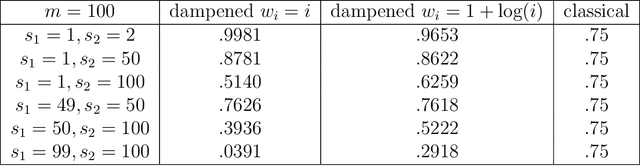

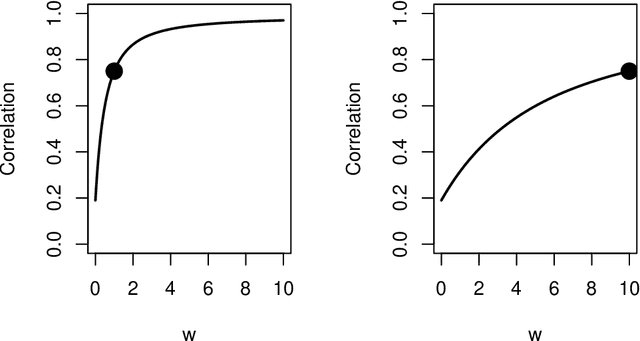
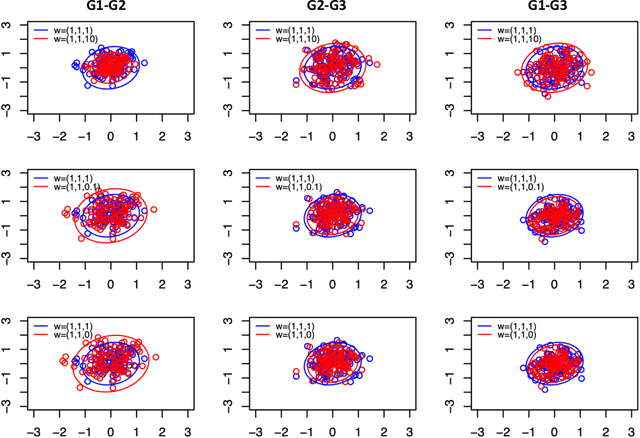
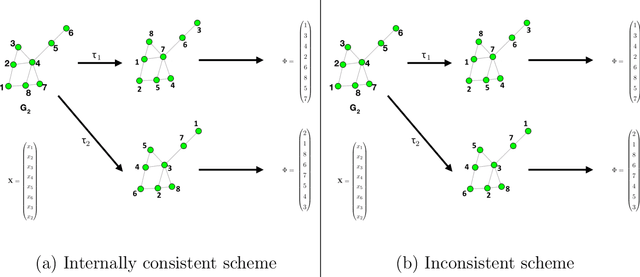
Abstract:Spectral inference on multiple networks is a rapidly-developing subfield of graph statistics. Recent work has demonstrated that joint, or simultaneous, spectral embedding of multiple independent network realizations can deliver more accurate estimation than individual spectral decompositions of those same networks. Little attention has been paid, however, to the network correlation that such joint embedding procedures necessarily induce. In this paper, we present a detailed analysis of induced correlation in a {\em generalized omnibus} embedding for multiple networks. We show that our embedding procedure is flexible and robust, and, moreover, we prove a central limit theorem for this embedding and explicitly compute the limiting covariance. We examine how this covariance can impact inference in a network time series, and we construct an appropriately calibrated omnibus embedding that can detect changes in real biological networks that previous embedding procedures could not discern. Our analysis confirms that the effect of induced correlation can be both subtle and transformative, with import in theory and practice.
On the role of features in vertex nomination: Content and context together are better
May 06, 2020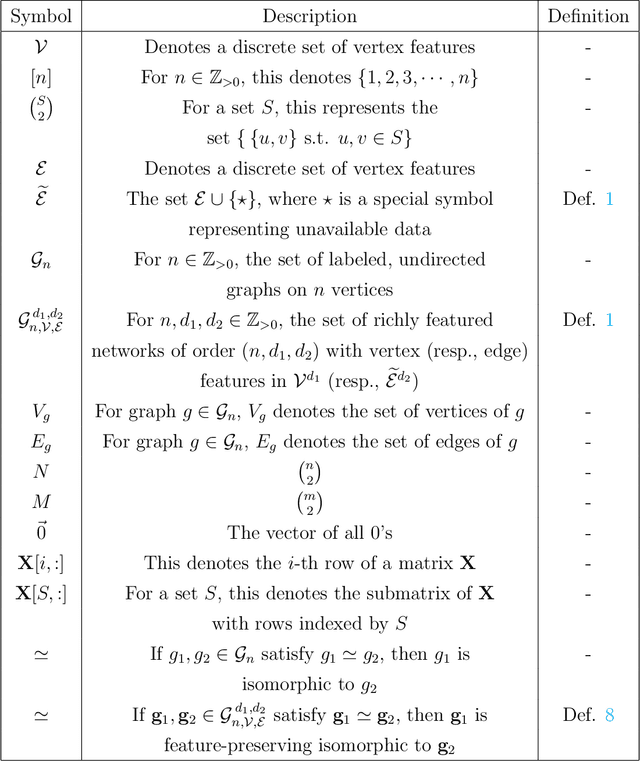
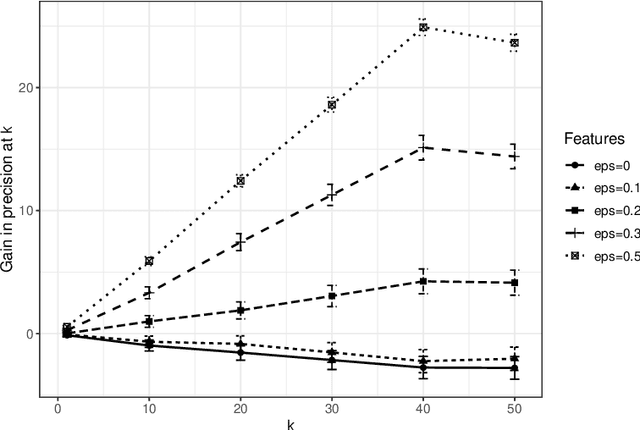

Abstract:Vertex nomination is a lightly-supervised network information retrieval (IR) task in which vertices of interest in one graph are used to query a second graph to discover vertices of interest in the second graph. Similar to other IR tasks, the output of a vertex nomination scheme is a ranked list of the vertices in the second graph, with the heretofore unknown vertices of interest ideally concentrating at the top of the list. Vertex nomination schemes provide a useful suite of tools for efficiently mining complex networks for pertinent information. In this paper, we explore, both theoretically and practically, the dual roles of content (i.e., edge and vertex attributes) and context (i.e., network topology) in vertex nomination. We provide necessary and sufficient conditions under which vertex nomination schemes that leverage both content and context outperform schemes that leverage only content or context separately. While the joint utility of both content and context has been demonstrated empirically in the literature, the framework presented in this paper provides a novel theoretical basis for understanding the potential complementary roles of network features and topology.
Graph matching between bipartite and unipartite networks: to collapse, or not to collapse, that is the question
Feb 05, 2020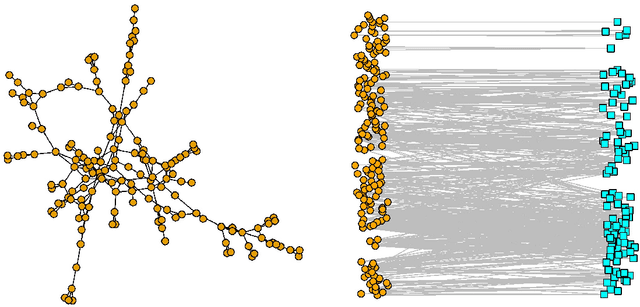
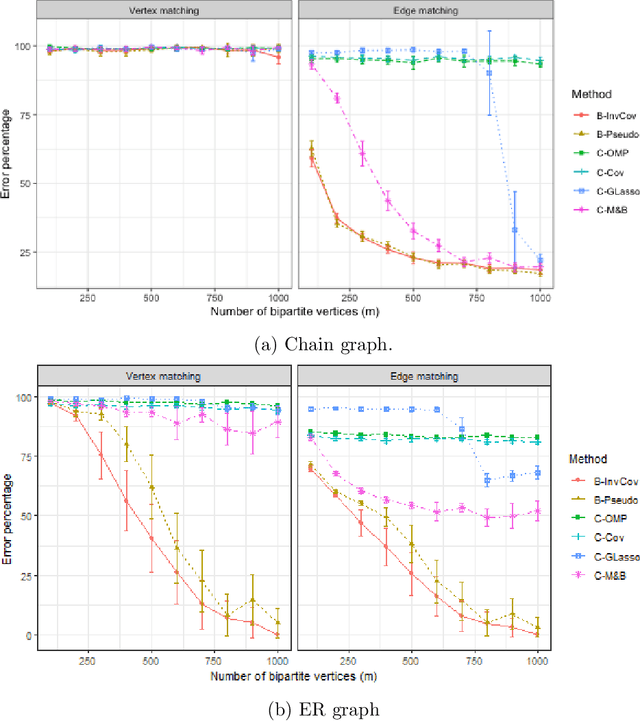
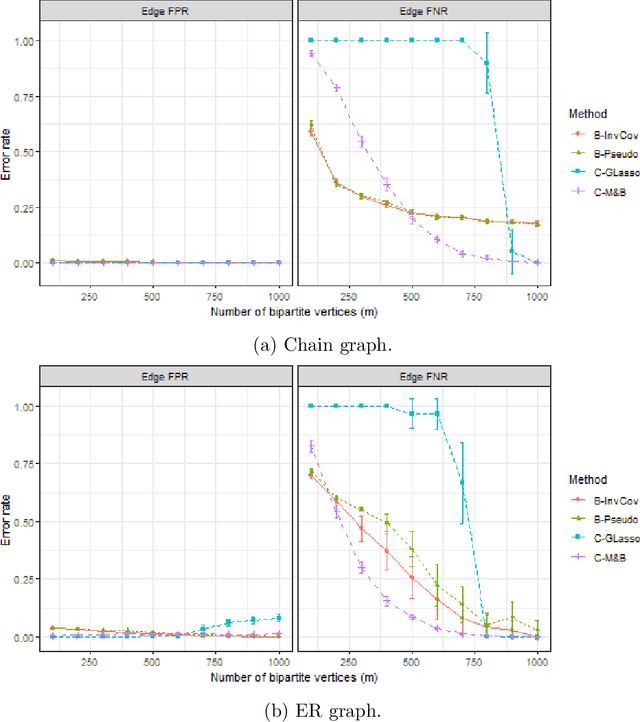
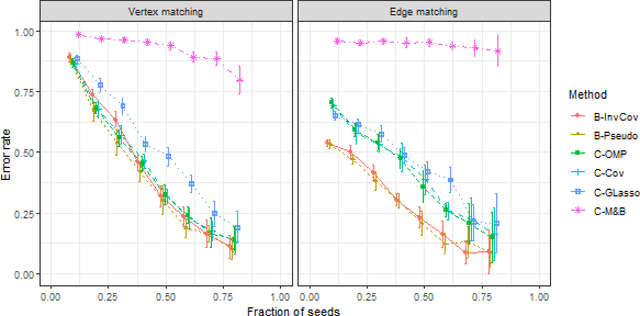
Abstract:Graph matching consists of aligning the vertices of two unlabeled graphs in order to maximize the shared structure across networks; when the graphs are unipartite, this is commonly formulated as minimizing their edge disagreements. In this paper, we address the common setting in which one of the graphs to match is a bipartite network and one is unipartite. Commonly, the bipartite networks are collapsed or projected into a unipartite graph, and graph matching proceeds as in the classical setting. This potentially leads to noisy edge estimates and loss of information. We formulate the graph matching problem between a bipartite and a unipartite graph using an undirected graphical model, and introduce methods to find the alignment with this model without collapsing. In simulations and real data examples, we show how our methods can result in a more accurate matching than the naive approach of transforming the bipartite networks into unipartite, and we demonstrate the performance gains achieved by our method in simulated and real data networks, including a co-authorship-citation network pair and brain structural and functional data.
Vertex Nomination, Consistent Estimation, and Adversarial Modification
May 15, 2019
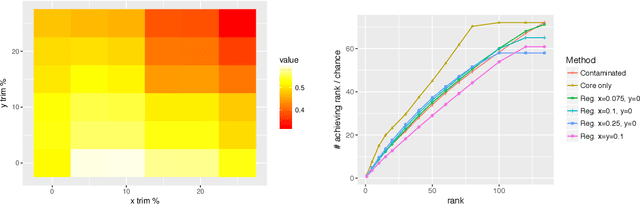
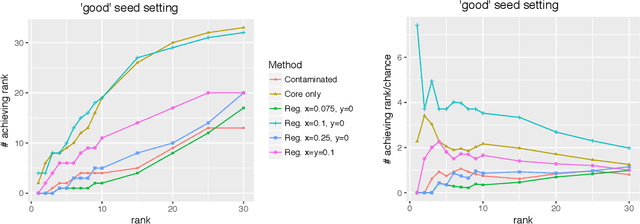
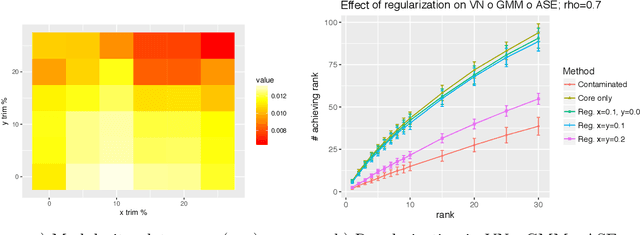
Abstract:Given a pair of graphs $G_1$ and $G_2$ and a vertex set of interest in $G_1$, the vertex nomination problem seeks to find the corresponding vertices of interest in $G_2$ (if they exist) and produce a rank list of the vertices in $G_2$, with the corresponding vertices of interest in $G_2$ concentrating, ideally, at the top of the rank list. In this paper we study the effect of an adversarial contamination model on the performance of a spectral graph embedding-based vertex nomination scheme. In both real and simulated examples, we demonstrate that this vertex nomination scheme performs effectively in the uncontaminated setting; adversarial network contamination adversely impacts the performance of our VN scheme; and network regularization successfully mitigates the impact of the contamination. In addition to furthering the theoretic basis of consistency in vertex nomination, the adversarial noise model posited herein is grounded in theoretical developments that allow us to frame the role of an adversary in terms of maximal vertex nomination consistency classes.
Maximum Likelihood Estimation and Graph Matching in Errorfully Observed Networks
Dec 26, 2018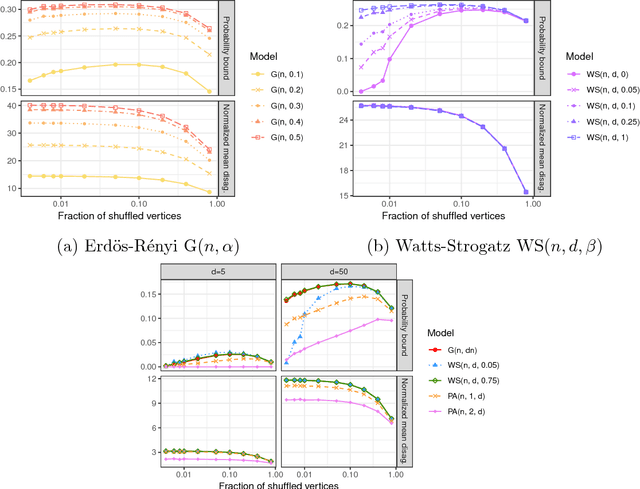
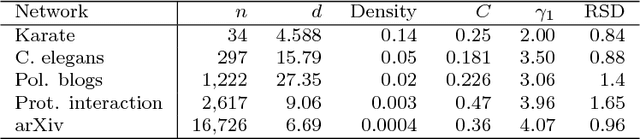
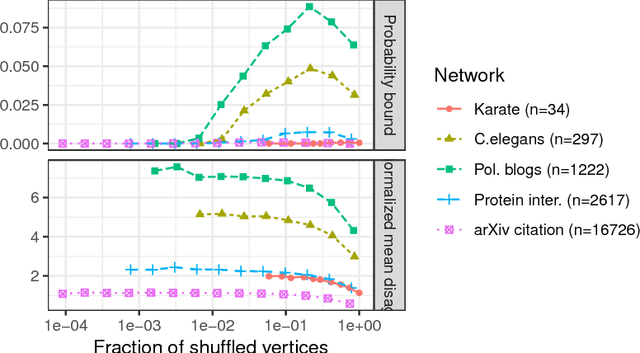

Abstract:Given a pair of graphs with the same number of vertices, the inexact graph matching problem consists in finding a correspondence between the vertices of these graphs that minimizes the total number of induced edge disagreements. We study this problem from a statistical framework in which one of the graphs is an errorfully observed copy of the other. We introduce a corrupting channel model, and show that in this model framework, the solution to the graph matching problem is a maximum likelihood estimator. Necessary and sufficient conditions for consistency of this MLE are presented, as well as a relaxed notion of consistency in which a negligible fraction of the vertices need not be matched correctly. The results are used to study matchability in several families of random graphs, including edge independent models, random regular graphs and small-world networks. We also use these results to introduce measures of matching feasibility, and experimentally validate the results on simulated and real-world networks.
On a 'Two Truths' Phenomenon in Spectral Graph Clustering
Sep 07, 2018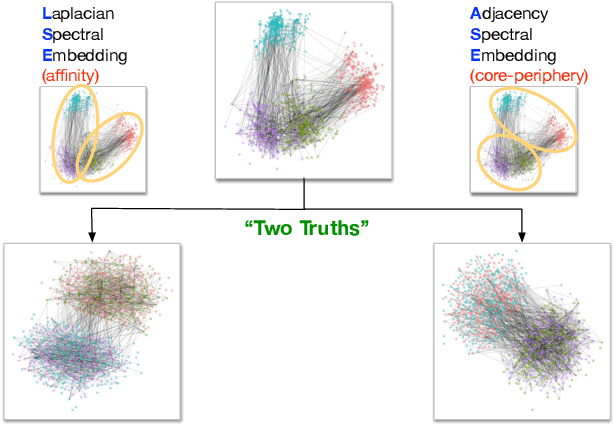
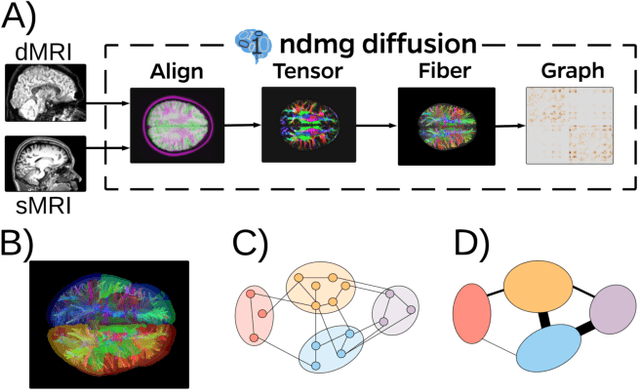
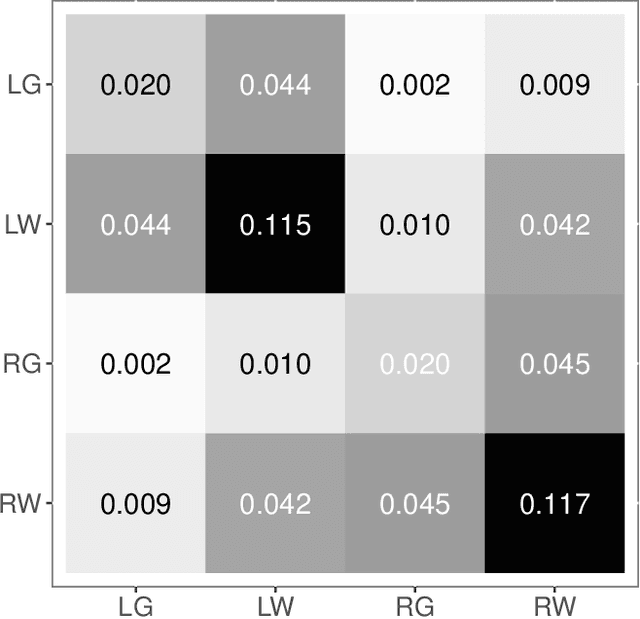
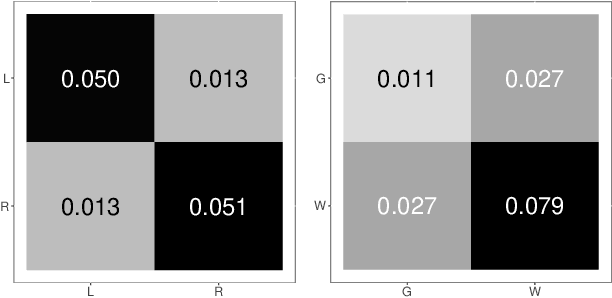
Abstract:Clustering is concerned with coherently grouping observations without any explicit concept of true groupings. Spectral graph clustering - clustering the vertices of a graph based on their spectral embedding - is commonly approached via K-means (or, more generally, Gaussian mixture model) clustering composed with either Laplacian or Adjacency spectral embedding (LSE or ASE). Recent theoretical results provide new understanding of the problem and solutions, and lead us to a 'Two Truths' LSE vs. ASE spectral graph clustering phenomenon convincingly illustrated here via a diffusion MRI connectome data set: the different embedding methods yield different clustering results, with LSE capturing left hemisphere/right hemisphere affinity structure and ASE capturing gray matter/white matter core-periphery structure.
On consistent vertex nomination schemes
May 29, 2018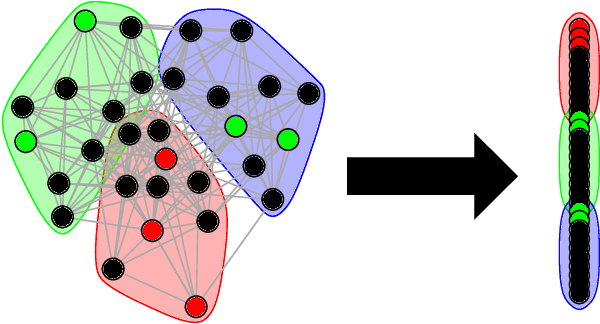
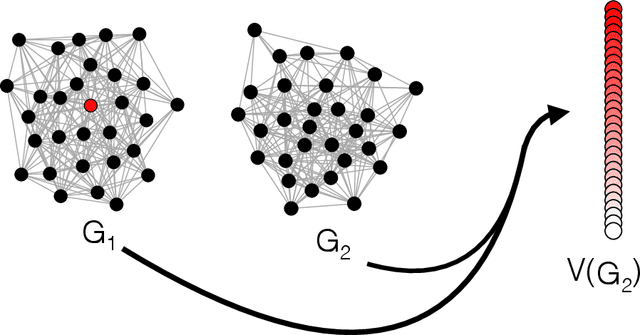
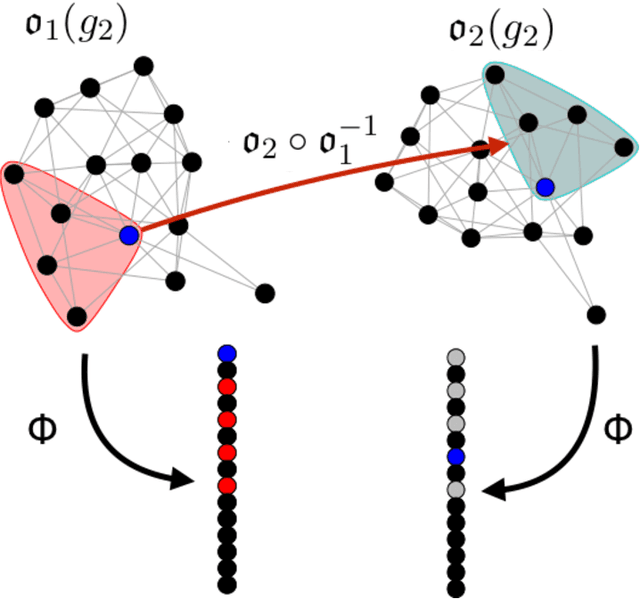
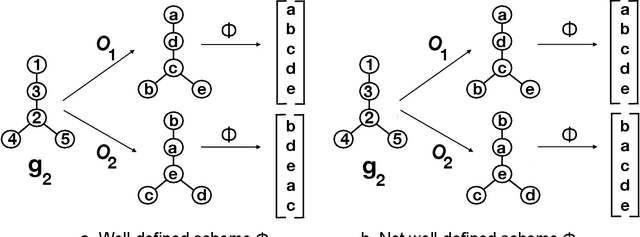
Abstract:Given a vertex of interest in a network $G_1$, the vertex nomination problem seeks to find the corresponding vertex of interest (if it exists) in a second network $G_2$. A vertex nomination scheme produces a rank list of the vertices in $G_2$, where the vertices are ranked by how likely they are judged to be the corresponding vertex of interest in $G_2$. The vertex nomination problem and related information retrieval tasks have attracted much attention in the machine learning literature, with numerous applications in social and biological networks. However, the current framework has often been confined to a comparatively small class of network models, and the concept of statistically consistent vertex nomination schemes has been only shallowly explored. In this paper, we extend the vertex nomination problem to a very general statistical model of graphs. Further, drawing inspiration from the long-established classification framework in the pattern recognition literature, we provide definitions for the key notions of Bayes optimality and consistency in our extended vertex nomination framework, including a derivation of the Bayes optimal vertex nomination scheme. In addition, we prove that no universally consistent vertex nomination schemes exist. Illustrative examples are provided throughout.
Seeded Graph Matching
Apr 10, 2018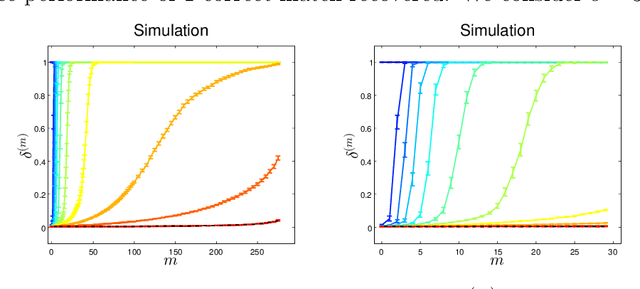
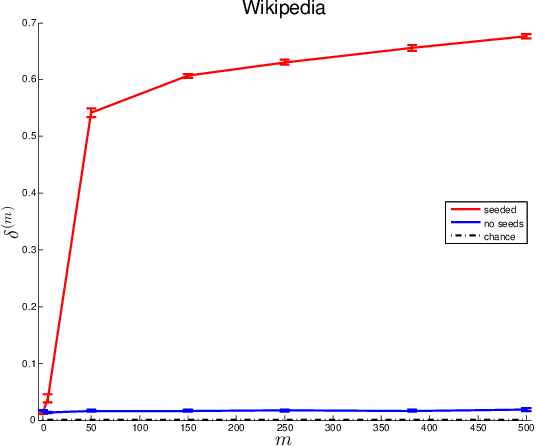
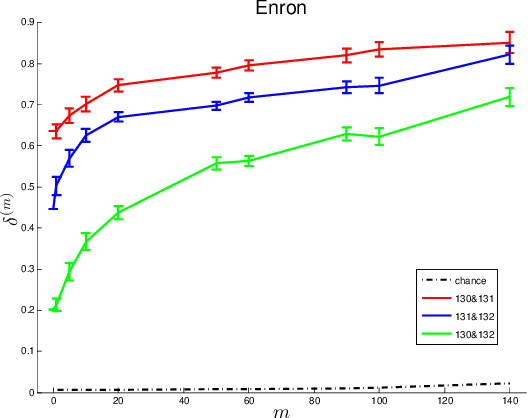
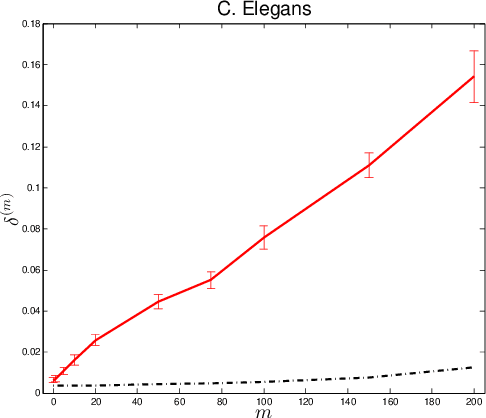
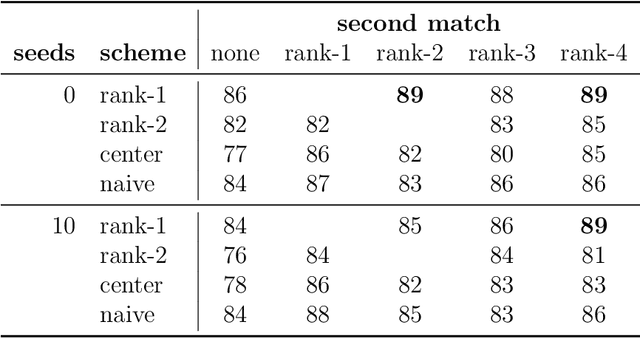
Abstract:Given two graphs, the graph matching problem is to align the two vertex sets so as to minimize the number of adjacency disagreements between the two graphs. The seeded graph matching problem is the graph matching problem when we are first given a partial alignment that we are tasked with completing. In this paper, we modify the state-of-the-art approximate graph matching algorithm "FAQ" of Vogelstein et al. (2015) to make it a fast approximate seeded graph matching algorithm, adapt its applicability to include graphs with differently sized vertex sets, and extend the algorithm so as to provide, for each individual vertex, a nomination list of likely matches. We demonstrate the effectiveness of our algorithm via simulation and real data experiments; indeed, knowledge of even a few seeds can be extremely effective when our seeded graph matching algorithm is used to recover a naturally existing alignment that is only partially observed.
Matched Filters for Noisy Induced Subgraph Detection
Mar 06, 2018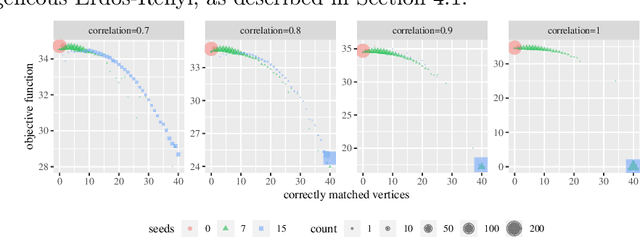
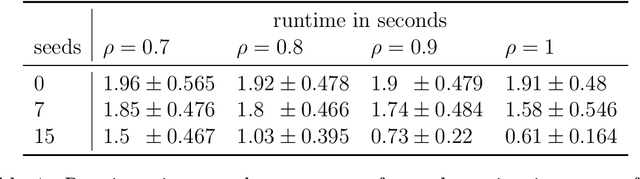

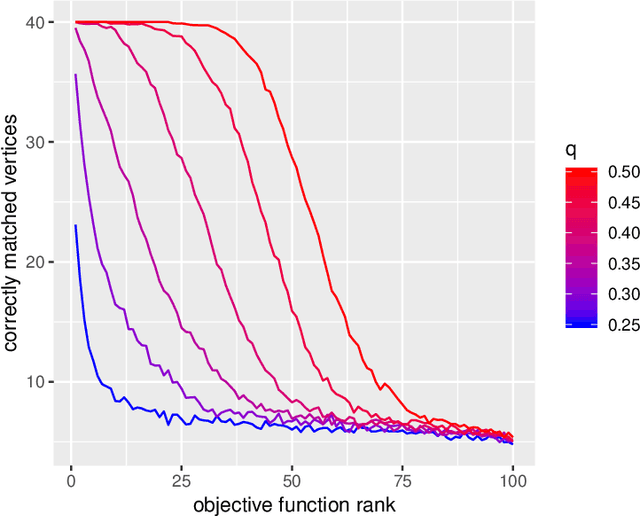
Abstract:We consider the problem of finding the vertex correspondence between two graphs with different number of vertices where the smaller graph is still potentially large. We propose a solution to this problem via a graph matching matched filter: padding the smaller graph in different ways and then using graph matching methods to align it to the larger network. Under a statistical model for correlated pairs of graphs, which yields a noisy copy of the small graph within the larger graph, the resulting optimization problem can be guaranteed to recover the true vertex correspondence between the networks, though there are currently no efficient algorithms for solving this problem. We consider an approach that exploits a partially known correspondence and show via varied simulations and applications to the Drosophila connectome that in practice this approach can achieve good performance.
Vertex nomination: The canonical sampling and the extended spectral nomination schemes
Feb 14, 2018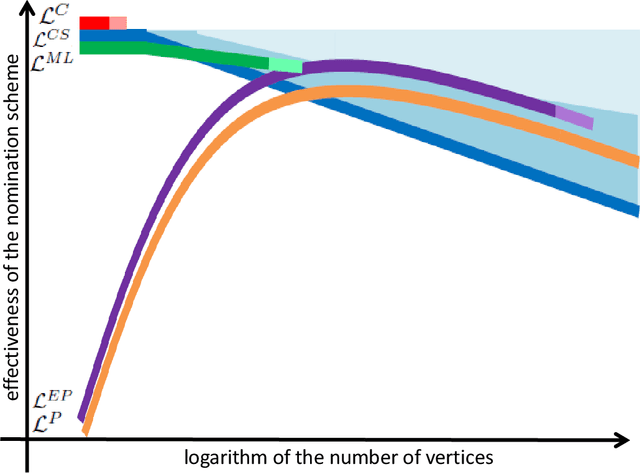
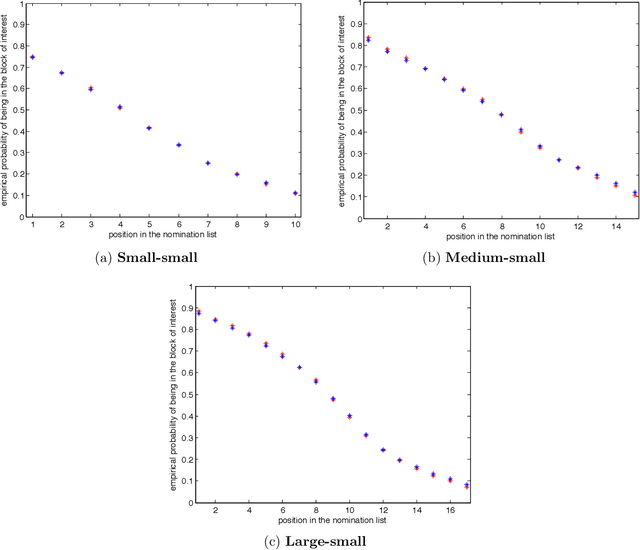
Abstract:Suppose that one particular block in a stochastic block model is deemed "interesting," but block labels are only observed for a few of the vertices. Utilizing a graph realized from the model, the vertex nomination task is to order the vertices with unobserved block labels into a "nomination list" with the goal of having an abundance of interesting vertices near the top of the list. In this paper we extend and enhance two basic vertex nomination schemes; the canonical nomination scheme ${\mathcal L}^C$ and the spectral partitioning nomination scheme ${\mathcal L}^P$. The canonical nomination scheme ${\mathcal L}^C$ is provably optimal, but is computationally intractable, being impractical to implement even on modestly sized graphs. With this in mind, we introduce a scalable, Markov chain Monte Carlo-based nomination scheme, called the {\it canonical sampling nomination scheme} ${\mathcal L}^{CS}$, that converges to the canonical nomination scheme ${\mathcal L}^{C}$ as the amount of sampling goes to infinity. We also introduce a novel spectral partitioning nomination scheme called the {\it extended spectral partitioning nomination scheme} ${\mathcal L}^{EP}$. Real-data and simulation experiments are employed to illustrate the effectiveness of these vertex nomination schemes, as well as their empirical computational complexity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge