Ursula Schmidt-Erfurth
on behalf of the PINNACLE consortium
Self-supervised learning via inter-modal reconstruction and feature projection networks for label-efficient 3D-to-2D segmentation
Jul 13, 2023Abstract:Deep learning has become a valuable tool for the automation of certain medical image segmentation tasks, significantly relieving the workload of medical specialists. Some of these tasks require segmentation to be performed on a subset of the input dimensions, the most common case being 3D-to-2D. However, the performance of existing methods is strongly conditioned by the amount of labeled data available, as there is currently no data efficient method, e.g. transfer learning, that has been validated on these tasks. In this work, we propose a novel convolutional neural network (CNN) and self-supervised learning (SSL) method for label-efficient 3D-to-2D segmentation. The CNN is composed of a 3D encoder and a 2D decoder connected by novel 3D-to-2D blocks. The SSL method consists of reconstructing image pairs of modalities with different dimensionality. The approach has been validated in two tasks with clinical relevance: the en-face segmentation of geographic atrophy and reticular pseudodrusen in optical coherence tomography. Results on different datasets demonstrate that the proposed CNN significantly improves the state of the art in scenarios with limited labeled data by up to 8% in Dice score. Moreover, the proposed SSL method allows further improvement of this performance by up to 23%, and we show that the SSL is beneficial regardless of the network architecture.
Morph-SSL: Self-Supervision with Longitudinal Morphing to Predict AMD Progression from OCT
Apr 17, 2023



Abstract:The lack of reliable biomarkers makes predicting the conversion from intermediate to neovascular age-related macular degeneration (iAMD, nAMD) a challenging task. We develop a Deep Learning (DL) model to predict the future risk of conversion of an eye from iAMD to nAMD from its current OCT scan. Although eye clinics generate vast amounts of longitudinal OCT scans to monitor AMD progression, only a small subset can be manually labeled for supervised DL. To address this issue, we propose Morph-SSL, a novel Self-supervised Learning (SSL) method for longitudinal data. It uses pairs of unlabelled OCT scans from different visits and involves morphing the scan from the previous visit to the next. The Decoder predicts the transformation for morphing and ensures a smooth feature manifold that can generate intermediate scans between visits through linear interpolation. Next, the Morph-SSL trained features are input to a Classifier which is trained in a supervised manner to model the cumulative probability distribution of the time to conversion with a sigmoidal function. Morph-SSL was trained on unlabelled scans of 399 eyes (3570 visits). The Classifier was evaluated with a five-fold cross-validation on 2418 scans from 343 eyes with clinical labels of the conversion date. The Morph-SSL features achieved an AUC of 0.766 in predicting the conversion to nAMD within the next 6 months, outperforming the same network when trained end-to-end from scratch or pre-trained with popular SSL methods. Automated prediction of the future risk of nAMD onset can enable timely treatment and individualized AMD management.
Clustering disease trajectories in contrastive feature space for biomarker discovery in age-related macular degeneration
Jan 11, 2023Abstract:Age-related macular degeneration (AMD) is the leading cause of blindness in the elderly. Despite this, the exact dynamics of disease progression are poorly understood. There is a clear need for imaging biomarkers in retinal optical coherence tomography (OCT) that aid the diagnosis, prognosis and management of AMD. However, current grading systems, which coarsely group disease stage into broad categories describing early and intermediate AMD, have very limited prognostic value for the conversion to late AMD. In this paper, we are the first to analyse disease progression as clustered trajectories in a self-supervised feature space. Our method first pretrains an encoder with contrastive learning to project images from longitudinal time series to points in feature space. This enables the creation of disease trajectories, which are then denoised, partitioned and grouped into clusters. These clusters, found in two datasets containing time series of 7,912 patients imaged over eight years, were correlated with known OCT biomarkers. This reinforced efforts by four expert ophthalmologists to investigate clusters, during a clinical comparison and interpretation task, as candidates for time-dependent biomarkers that describe progression of AMD.
Learning Spatio-Temporal Model of Disease Progression with NeuralODEs from Longitudinal Volumetric Data
Nov 08, 2022



Abstract:Robust forecasting of the future anatomical changes inflicted by an ongoing disease is an extremely challenging task that is out of grasp even for experienced healthcare professionals. Such a capability, however, is of great importance since it can improve patient management by providing information on the speed of disease progression already at the admission stage, or it can enrich the clinical trials with fast progressors and avoid the need for control arms by the means of digital twins. In this work, we develop a deep learning method that models the evolution of age-related disease by processing a single medical scan and providing a segmentation of the target anatomy at a requested future point in time. Our method represents a time-invariant physical process and solves a large-scale problem of modeling temporal pixel-level changes utilizing NeuralODEs. In addition, we demonstrate the approaches to incorporate the prior domain-specific constraints into our method and define temporal Dice loss for learning temporal objectives. To evaluate the applicability of our approach across different age-related diseases and imaging modalities, we developed and tested the proposed method on the datasets with 967 retinal OCT volumes of 100 patients with Geographic Atrophy, and 2823 brain MRI volumes of 633 patients with Alzheimer's Disease. For Geographic Atrophy, the proposed method outperformed the related baseline models in the atrophy growth prediction. For Alzheimer's Disease, the proposed method demonstrated remarkable performance in predicting the brain ventricle changes induced by the disease, achieving the state-of-the-art result on TADPOLE challenge.
Segmentation of Bruch's Membrane in retinal OCT with AMD using anatomical priors and uncertainty quantification
Oct 30, 2022Abstract:Bruch's membrane (BM) segmentation on optical coherence tomography (OCT) is a pivotal step for the diagnosis and follow-up of age-related macular degeneration (AMD), one of the leading causes of blindness in the developed world. Automated BM segmentation methods exist, but they usually do not account for the anatomical coherence of the results, neither provide feedback on the confidence of the prediction. These factors limit the applicability of these systems in real-world scenarios. With this in mind, we propose an end-to-end deep learning method for automated BM segmentation in AMD patients. An Attention U-Net is trained to output a probability density function of the BM position, while taking into account the natural curvature of the surface. Besides the surface position, the method also estimates an A-scan wise uncertainty measure of the segmentation output. Subsequently, the A-scans with high uncertainty are interpolated using thin plate splines (TPS). We tested our method with ablation studies on an internal dataset with 138 patients covering all three AMD stages, and achieved a mean absolute localization error of 4.10 um. In addition, the proposed segmentation method was compared against the state-of-the-art methods and showed a superior performance on an external publicly available dataset from a different patient cohort and OCT device, demonstrating strong generalization ability.
Data-centric AI approach to improve optic nerve head segmentation and localization in OCT en face images
Aug 08, 2022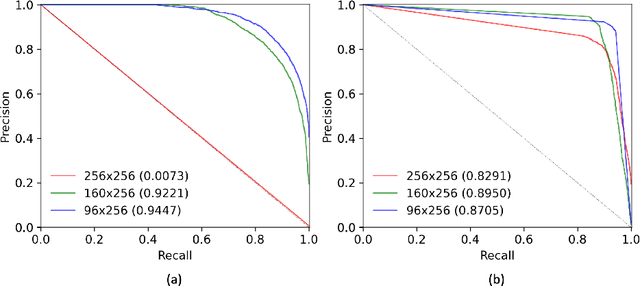
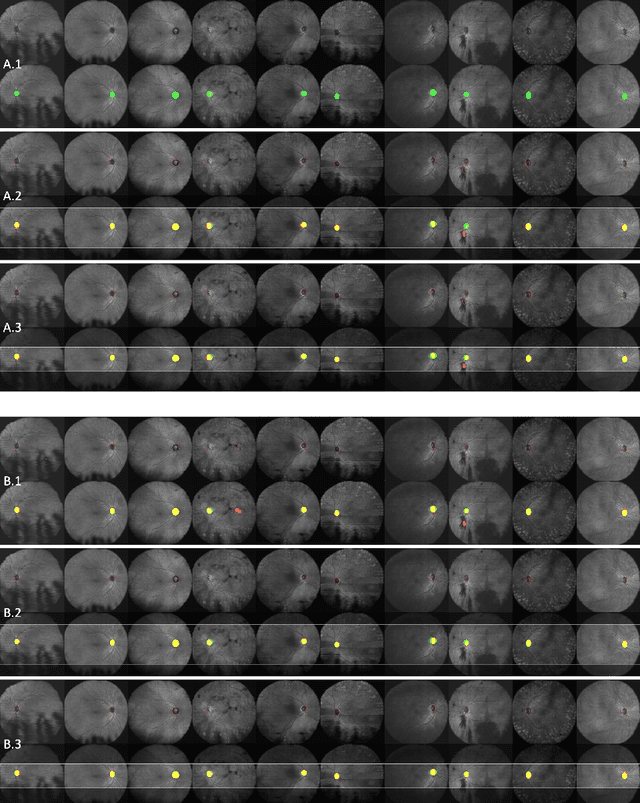
Abstract:The automatic detection and localization of anatomical features in retinal imaging data are relevant for many aspects. In this work, we follow a data-centric approach to optimize classifier training for optic nerve head detection and localization in optical coherence tomography en face images of the retina. We examine the effect of domain knowledge driven spatial complexity reduction on the resulting optic nerve head segmentation and localization performance. We present a machine learning approach for segmenting optic nerve head in 2D en face projections of 3D widefield swept source optical coherence tomography scans that enables the automated assessment of large amounts of data. Evaluation on manually annotated 2D en face images of the retina demonstrates that training of a standard U-Net can yield improved optic nerve head segmentation and localization performance when the underlying pixel-level binary classification task is spatially relaxed through domain knowledge.
Metadata-enhanced contrastive learning from retinal optical coherence tomography images
Aug 04, 2022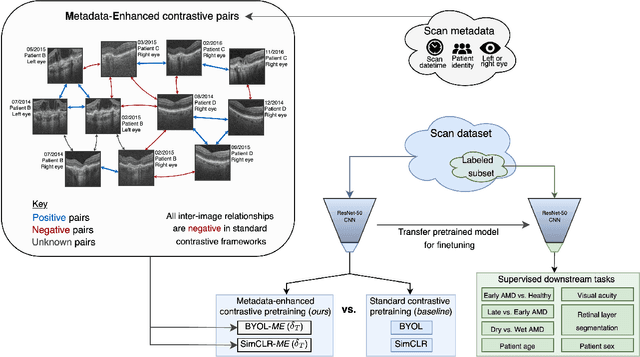
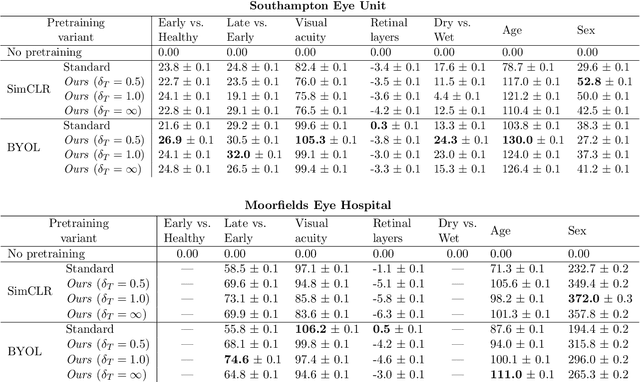
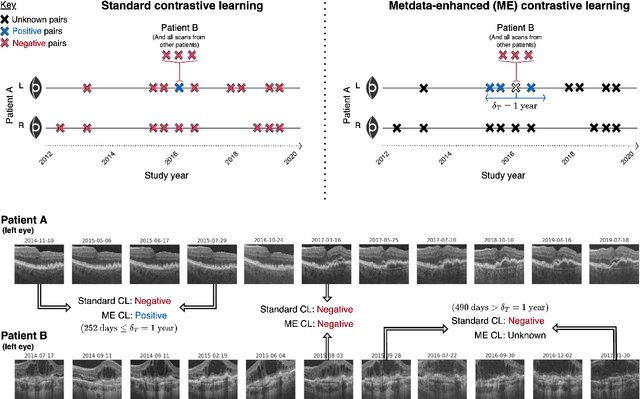
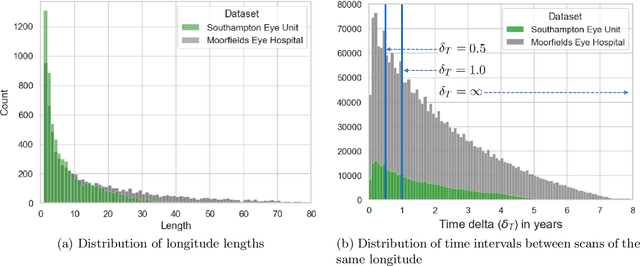
Abstract:Supervised deep learning algorithms hold great potential to automate screening, monitoring and grading of medical images. However, training performant models has typically required vast quantities of labelled data, which is scarcely available in the medical domain. Self-supervised contrastive frameworks relax this dependency by first learning from unlabelled images. In this work we show that pretraining with two contrastive methods, SimCLR and BYOL, improves the utility of deep learning with regard to the clinical assessment of age-related macular degeneration (AMD). In experiments using two large clinical datasets containing 170,427 optical coherence tomography (OCT) images of 7,912 patients, we evaluate benefits attributed to pretraining across seven downstream tasks ranging from AMD stage and type classification to prediction of functional endpoints to segmentation of retinal layers, finding performance significantly increased in six out of seven tasks with fewer labels. However, standard contrastive frameworks have two known weaknesses that are detrimental to pretraining in the medical domain. Several of the image transformations used to create positive contrastive pairs are not applicable to greyscale medical scans. Furthermore, medical images often depict the same anatomical region and disease severity, resulting in numerous misleading negative pairs. To address these issues we develop a novel metadata-enhanced approach that exploits the rich set of inherently available patient information. To this end we employ records for patient identity, eye position (i.e. left or right) and time series data to indicate the typically unknowable set of inter-image contrastive relationships. By leveraging this often neglected information our metadata-enhanced contrastive pretraining leads to further benefits and outperforms conventional contrastive methods in five out of seven downstream tasks.
SD-LayerNet: Semi-supervised retinal layer segmentation in OCT using disentangled representation with anatomical priors
Jul 01, 2022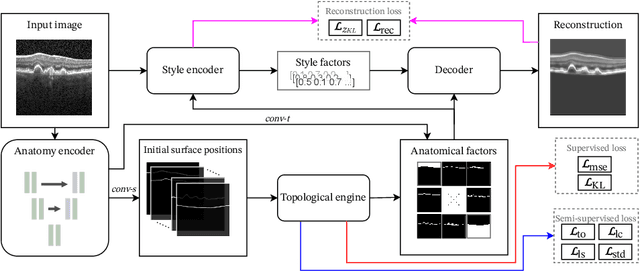
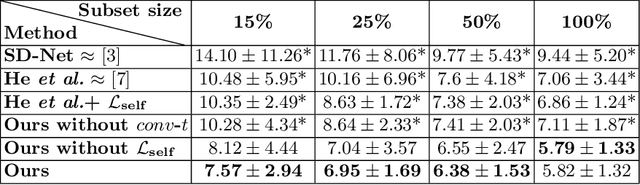
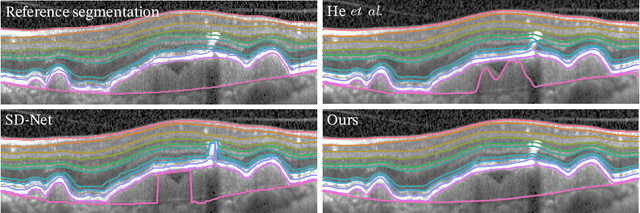
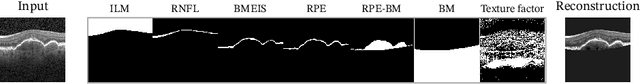
Abstract:Optical coherence tomography (OCT) is a non-invasive 3D modality widely used in ophthalmology for imaging the retina. Achieving automated, anatomically coherent retinal layer segmentation on OCT is important for the detection and monitoring of different retinal diseases, like Age-related Macular Disease (AMD) or Diabetic Retinopathy. However, the majority of state-of-the-art layer segmentation methods are based on purely supervised deep-learning, requiring a large amount of pixel-level annotated data that is expensive and hard to obtain. With this in mind, we introduce a semi-supervised paradigm into the retinal layer segmentation task that makes use of the information present in large-scale unlabeled datasets as well as anatomical priors. In particular, a novel fully differentiable approach is used for converting surface position regression into a pixel-wise structured segmentation, allowing to use both 1D surface and 2D layer representations in a coupled fashion to train the model. In particular, these 2D segmentations are used as anatomical factors that, together with learned style factors, compose disentangled representations used for reconstructing the input image. In parallel, we propose a set of anatomical priors to improve network training when a limited amount of labeled data is available. We demonstrate on the real-world dataset of scans with intermediate and wet-AMD that our method outperforms state-of-the-art when using our full training set, but more importantly largely exceeds state-of-the-art when it is trained with a fraction of the labeled data.
TINC: Temporally Informed Non-Contrastive Learning for Disease Progression Modeling in Retinal OCT Volumes
Jun 30, 2022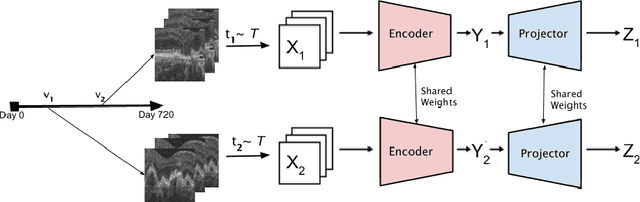
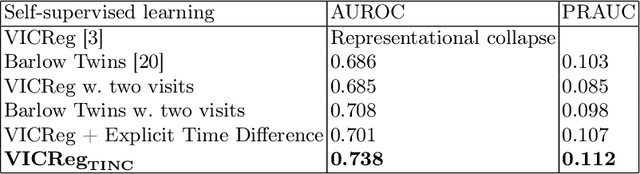

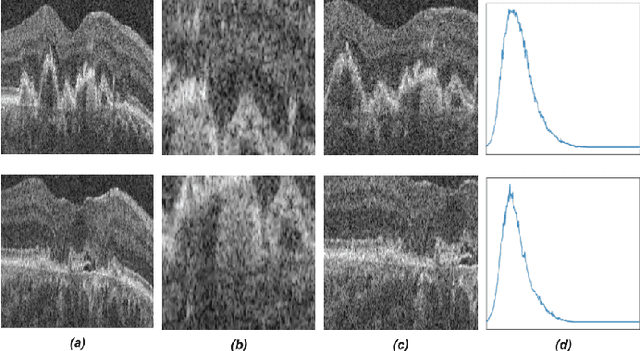

Abstract:Recent contrastive learning methods achieved state-of-the-art in low label regimes. However, the training requires large batch sizes and heavy augmentations to create multiple views of an image. With non-contrastive methods, the negatives are implicitly incorporated in the loss, allowing different images and modalities as pairs. Although the meta-information (i.e., age, sex) in medical imaging is abundant, the annotations are noisy and prone to class imbalance. In this work, we exploited already existing temporal information (different visits from a patient) in a longitudinal optical coherence tomography (OCT) dataset using temporally informed non-contrastive loss (TINC) without increasing complexity and need for negative pairs. Moreover, our novel pair-forming scheme can avoid heavy augmentations and implicitly incorporates the temporal information in the pairs. Finally, these representations learned from the pretraining are more successful in predicting disease progression where the temporal information is crucial for the downstream task. More specifically, our model outperforms existing models in predicting the risk of conversion within a time frame from intermediate age-related macular degeneration (AMD) to the late wet-AMD stage.
Blood vessel segmentation in en-face OCTA images: a frequency based method
Sep 13, 2021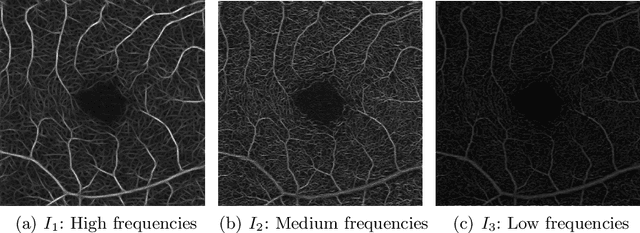
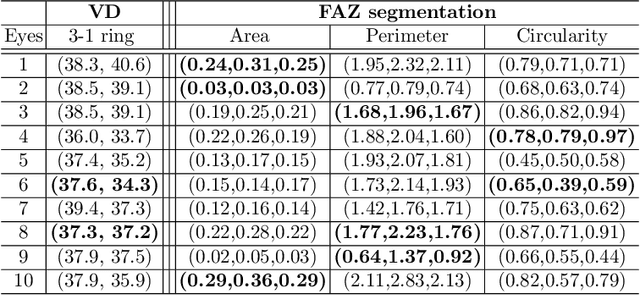
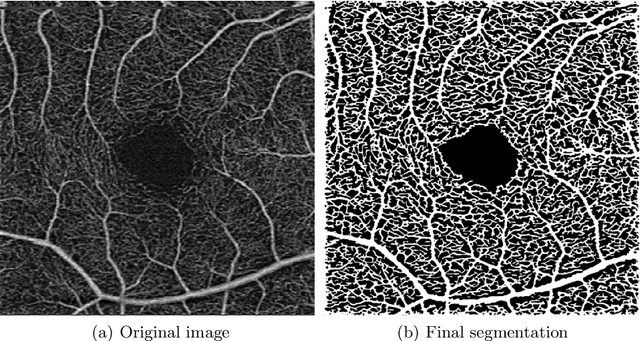
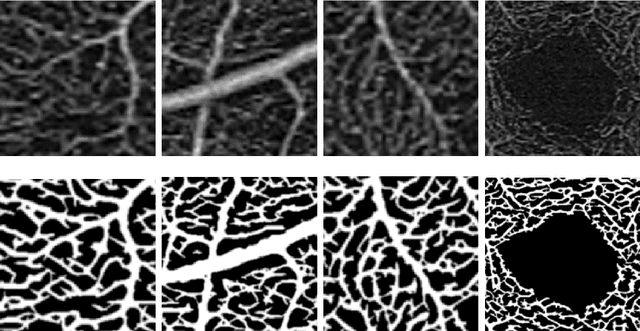
Abstract:Optical coherence tomography angiography (OCTA) is a novel noninvasive imaging modality for visualization of retinal blood flow in the human retina. Using specific OCTA imaging biomarkers for the identification of pathologies, automated image segmentations of the blood vessels can improve subsequent analysis and diagnosis. We present a novel method for the vessel identification based on frequency representations of the image, in particular, using so-called Gabor filter banks. The algorithm is evaluated on an OCTA image data set from $10$ eyes acquired by a Cirrus HD-OCT device. The segmentation outcomes received very good qualitative visual evaluation feedback and coincide well with device-specific values concerning vessel density. Concerning locality our segmentations are even more reliable and accurate. Therefore, we suggest the computation of adaptive local vessel density maps that allow straightforward analysis of retinal blood flow.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge