Uddeshya Upadhyay
Looking Beyond Language Priors: Enhancing Visual Comprehension and Attention in Multimodal Models
May 08, 2025



Abstract:Achieving deep alignment between vision and language remains a central challenge for Multimodal Large Language Models (MLLMs). These models often fail to fully leverage visual input, defaulting to strong language priors. Our approach first provides insights into how MLLMs internally build visual understanding of image regions and then introduces techniques to amplify this capability. Specifically, we explore techniques designed both to deepen the model's understanding of visual content and to ensure that these visual insights actively guide language generation. We demonstrate the superior multimodal understanding of our resultant model through a detailed upstream analysis quantifying its ability to predict visually-dependent tokens as well as 10 pt boost on visually challenging tasks.
HypUC: Hyperfine Uncertainty Calibration with Gradient-boosted Corrections for Reliable Regression on Imbalanced Electrocardiograms
Nov 23, 2023



Abstract:The automated analysis of medical time series, such as the electrocardiogram (ECG), electroencephalogram (EEG), pulse oximetry, etc, has the potential to serve as a valuable tool for diagnostic decisions, allowing for remote monitoring of patients and more efficient use of expensive and time-consuming medical procedures. Deep neural networks (DNNs) have been demonstrated to process such signals effectively. However, previous research has primarily focused on classifying medical time series rather than attempting to regress the continuous-valued physiological parameters central to diagnosis. One significant challenge in this regard is the imbalanced nature of the dataset, as a low prevalence of abnormal conditions can lead to heavily skewed data that results in inaccurate predictions and a lack of certainty in such predictions when deployed. To address these challenges, we propose HypUC, a framework for imbalanced probabilistic regression in medical time series, making several contributions. (i) We introduce a simple kernel density-based technique to tackle the imbalanced regression problem with medical time series. (ii) Moreover, we employ a probabilistic regression framework that allows uncertainty estimation for the predicted continuous values. (iii) We also present a new approach to calibrate the predicted uncertainty further. (iv) Finally, we demonstrate a technique to use calibrated uncertainty estimates to improve the predicted continuous value and show the efficacy of the calibrated uncertainty estimates to flag unreliable predictions. HypUC is evaluated on a large, diverse, real-world dataset of ECGs collected from millions of patients, outperforming several conventional baselines on various diagnostic tasks, suggesting a potential use-case for the reliable clinical deployment of deep learning models.
* Published at TMLR
ProbVLM: Probabilistic Adapter for Frozen Vison-Language Models
Jul 01, 2023



Abstract:Large-scale vision-language models (VLMs) like CLIP successfully find correspondences between images and text. Through the standard deterministic mapping process, an image or a text sample is mapped to a single vector in the embedding space. This is problematic: as multiple samples (images or text) can abstract the same concept in the physical world, deterministic embeddings do not reflect the inherent ambiguity in the embedding space. We propose ProbVLM, a probabilistic adapter that estimates probability distributions for the embeddings of pre-trained VLMs via inter/intra-modal alignment in a post-hoc manner without needing large-scale datasets or computing. On four challenging datasets, i.e., COCO, Flickr, CUB, and Oxford-flowers, we estimate the multi-modal embedding uncertainties for two VLMs, i.e., CLIP and BLIP, quantify the calibration of embedding uncertainties in retrieval tasks and show that ProbVLM outperforms other methods. Furthermore, we propose active learning and model selection as two real-world downstream tasks for VLMs and show that the estimated uncertainty aids both tasks. Lastly, we present a novel technique for visualizing the embedding distributions using a large-scale pre-trained latent diffusion model.
USIM-DAL: Uncertainty-aware Statistical Image Modeling-based Dense Active Learning for Super-resolution
May 27, 2023



Abstract:Dense regression is a widely used approach in computer vision for tasks such as image super-resolution, enhancement, depth estimation, etc. However, the high cost of annotation and labeling makes it challenging to achieve accurate results. We propose incorporating active learning into dense regression models to address this problem. Active learning allows models to select the most informative samples for labeling, reducing the overall annotation cost while improving performance. Despite its potential, active learning has not been widely explored in high-dimensional computer vision regression tasks like super-resolution. We address this research gap and propose a new framework called USIM-DAL that leverages the statistical properties of colour images to learn informative priors using probabilistic deep neural networks that model the heteroscedastic predictive distribution allowing uncertainty quantification. Moreover, the aleatoric uncertainty from the network serves as a proxy for error that is used for active learning. Our experiments on a wide variety of datasets spanning applications in natural images (visual genome, BSD100), medical imaging (histopathology slides), and remote sensing (satellite images) demonstrate the efficacy of the newly proposed USIM-DAL and superiority over several dense regression active learning methods.
Posterior Annealing: Fast Calibrated Uncertainty for Regression
Feb 21, 2023Abstract:Bayesian deep learning approaches that allow uncertainty estimation for regression problems often converge slowly and yield poorly calibrated uncertainty estimates that can not be effectively used for quantification. Recently proposed post hoc calibration techniques are seldom applicable to regression problems and often add overhead to an already slow model training phase. This work presents a fast calibrated uncertainty estimation method for regression tasks, called posterior annealing, that consistently improves the convergence of deep regression models and yields calibrated uncertainty without any post hoc calibration phase. Unlike previous methods for calibrated uncertainty in regression that focus only on low-dimensional regression problems, our method works well on a wide spectrum of regression problems. Our empirical analysis shows that our approach is generalizable to various network architectures including, multilayer perceptrons, 1D/2D convolutional networks, and graph neural networks, on five vastly diverse tasks, i.e., chaotic particle trajectory denoising, physical property prediction of molecules using 3D atomistic representation, natural image super-resolution, and medical image translation using MRI images.
Towards Automating Retinoscopy for Refractive Error Diagnosis
Aug 10, 2022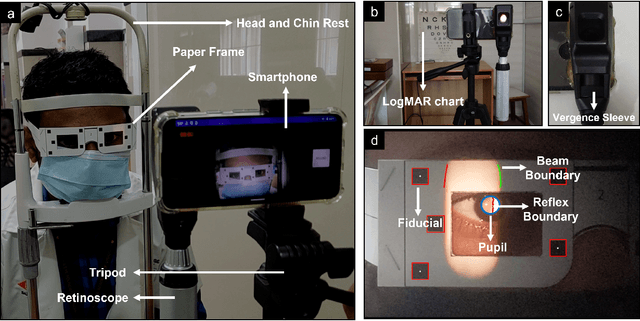
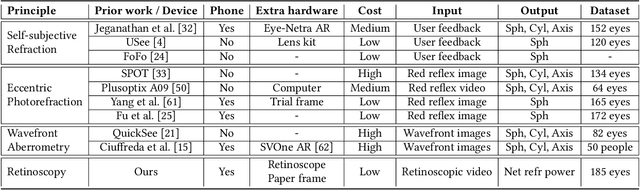
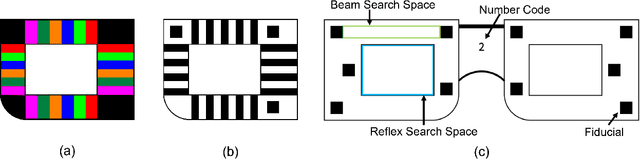

Abstract:Refractive error is the most common eye disorder and is the key cause behind correctable visual impairment, responsible for nearly 80% of the visual impairment in the US. Refractive error can be diagnosed using multiple methods, including subjective refraction, retinoscopy, and autorefractors. Although subjective refraction is the gold standard, it requires cooperation from the patient and hence is not suitable for infants, young children, and developmentally delayed adults. Retinoscopy is an objective refraction method that does not require any input from the patient. However, retinoscopy requires a lens kit and a trained examiner, which limits its use for mass screening. In this work, we automate retinoscopy by attaching a smartphone to a retinoscope and recording retinoscopic videos with the patient wearing a custom pair of paper frames. We develop a video processing pipeline that takes retinoscopic videos as input and estimates the net refractive error based on our proposed extension of the retinoscopy mathematical model. Our system alleviates the need for a lens kit and can be performed by an untrained examiner. In a clinical trial with 185 eyes, we achieved a sensitivity of 91.0% and specificity of 74.0% on refractive error diagnosis. Moreover, the mean absolute error of our approach was 0.75$\pm$0.67D on net refractive error estimation compared to subjective refraction measurements. Our results indicate that our approach has the potential to be used as a retinoscopy-based refractive error screening tool in real-world medical settings.
BayesCap: Bayesian Identity Cap for Calibrated Uncertainty in Frozen Neural Networks
Jul 14, 2022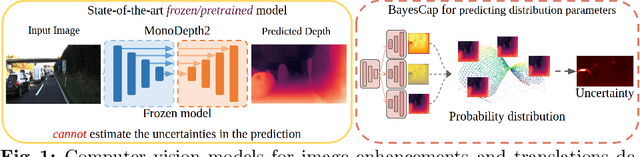
Abstract:High-quality calibrated uncertainty estimates are crucial for numerous real-world applications, especially for deep learning-based deployed ML systems. While Bayesian deep learning techniques allow uncertainty estimation, training them with large-scale datasets is an expensive process that does not always yield models competitive with non-Bayesian counterparts. Moreover, many of the high-performing deep learning models that are already trained and deployed are non-Bayesian in nature and do not provide uncertainty estimates. To address these issues, we propose BayesCap that learns a Bayesian identity mapping for the frozen model, allowing uncertainty estimation. BayesCap is a memory-efficient method that can be trained on a small fraction of the original dataset, enhancing pretrained non-Bayesian computer vision models by providing calibrated uncertainty estimates for the predictions without (i) hampering the performance of the model and (ii) the need for expensive retraining the model from scratch. The proposed method is agnostic to various architectures and tasks. We show the efficacy of our method on a wide variety of tasks with a diverse set of architectures, including image super-resolution, deblurring, inpainting, and crucial application such as medical image translation. Moreover, we apply the derived uncertainty estimates to detect out-of-distribution samples in critical scenarios like depth estimation in autonomous driving. Code is available at https://github.com/ExplainableML/BayesCap.
The Manifold Hypothesis for Gradient-Based Explanations
Jun 15, 2022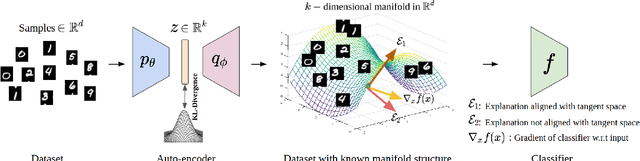
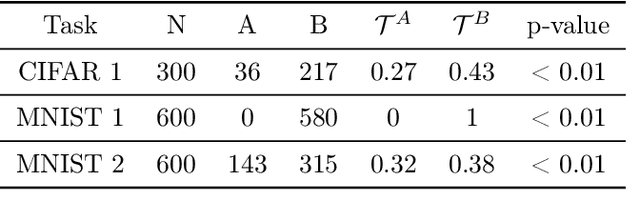
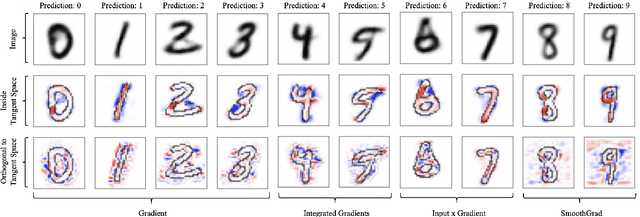
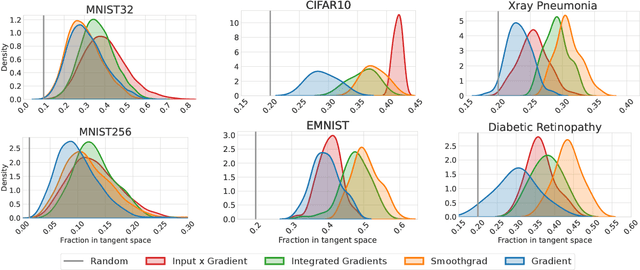
Abstract:When do gradient-based explanation algorithms provide meaningful explanations? We propose a necessary criterion: their feature attributions need to be aligned with the tangent space of the data manifold. To provide evidence for this hypothesis, we introduce a framework based on variational autoencoders that allows to estimate and generate image manifolds. Through experiments across a range of different datasets -- MNIST, EMNIST, CIFAR10, X-ray pneumonia and Diabetic Retinopathy detection -- we demonstrate that the more a feature attribution is aligned with the tangent space of the data, the more structured and explanatory it tends to be. In particular, the attributions provided by popular post-hoc methods such as Integrated Gradients, SmoothGrad and Input $\times$ Gradient tend to be more strongly aligned with the data manifold than the raw gradient. As a consequence, we suggest that explanation algorithms should actively strive to align their explanations with the data manifold. In part, this can be achieved by adversarial training, which leads to better alignment across all datasets. Some form of adjustment to the model architecture or training algorithm is necessary, since we show that generalization of neural networks alone does not imply the alignment of model gradients with the data manifold.
Robustness via Uncertainty-aware Cycle Consistency
Oct 24, 2021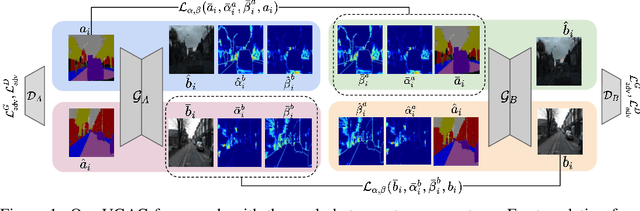
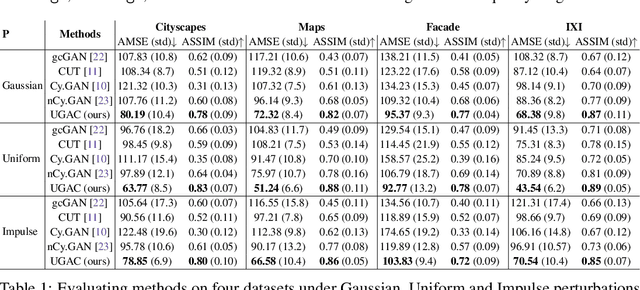
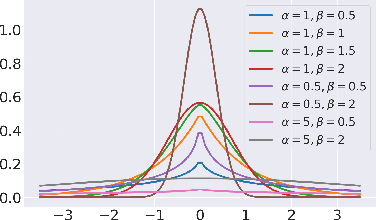
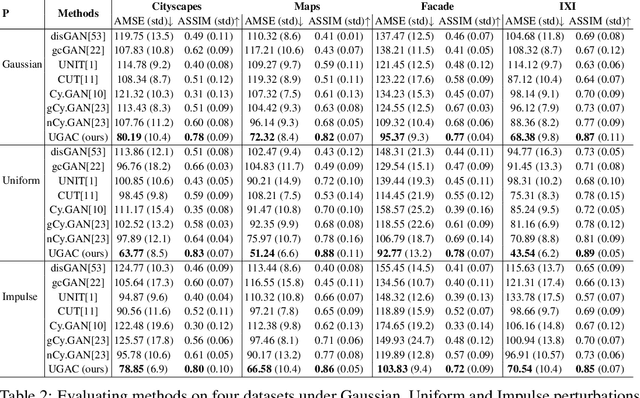
Abstract:Unpaired image-to-image translation refers to learning inter-image-domain mapping without corresponding image pairs. Existing methods learn deterministic mappings without explicitly modelling the robustness to outliers or predictive uncertainty, leading to performance degradation when encountering unseen perturbations at test time. To address this, we propose a novel probabilistic method based on Uncertainty-aware Generalized Adaptive Cycle Consistency (UGAC), which models the per-pixel residual by generalized Gaussian distribution, capable of modelling heavy-tailed distributions. We compare our model with a wide variety of state-of-the-art methods on various challenging tasks including unpaired image translation of natural images, using standard datasets, spanning autonomous driving, maps, facades, and also in medical imaging domain consisting of MRI. Experimental results demonstrate that our method exhibits stronger robustness towards unseen perturbations in test data. Code is released here: https://github.com/ExplainableML/UncertaintyAwareCycleConsistency.
Uncertainty-aware GAN with Adaptive Loss for Robust MRI Image Enhancement
Oct 07, 2021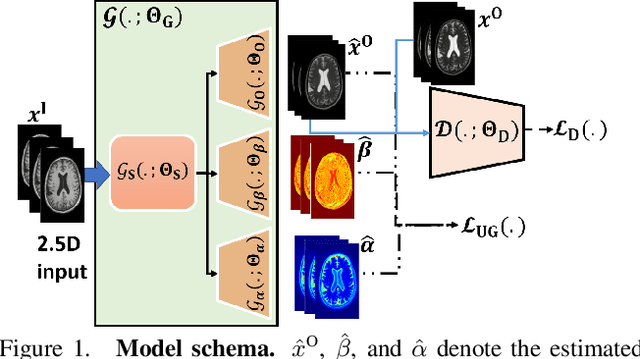
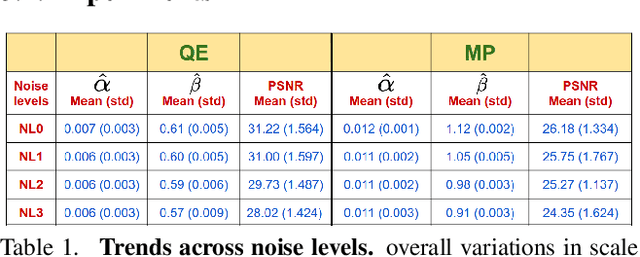
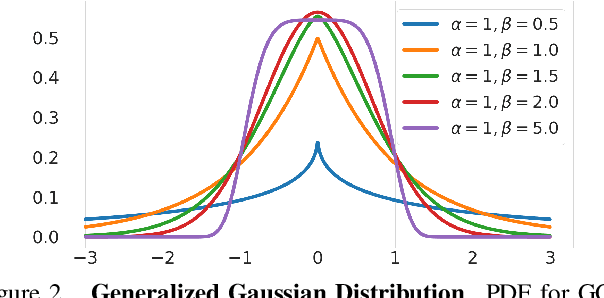
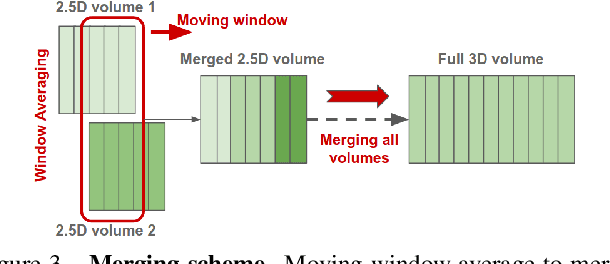
Abstract:Image-to-image translation is an ill-posed problem as unique one-to-one mapping may not exist between the source and target images. Learning-based methods proposed in this context often evaluate the performance on test data that is similar to the training data, which may be impractical. This demands robust methods that can quantify uncertainty in the prediction for making informed decisions, especially for critical areas such as medical imaging. Recent works that employ conditional generative adversarial networks (GANs) have shown improved performance in learning photo-realistic image-to-image mappings between the source and the target images. However, these methods do not focus on (i)~robustness of the models to out-of-distribution (OOD)-noisy data and (ii)~uncertainty quantification. This paper proposes a GAN-based framework that (i)~models an adaptive loss function for robustness to OOD-noisy data that automatically tunes the spatially varying norm for penalizing the residuals and (ii)~estimates the per-voxel uncertainty in the predictions. We demonstrate our method on two key applications in medical imaging: (i)~undersampled magnetic resonance imaging (MRI) reconstruction (ii)~MRI modality propagation. Our experiments with two different real-world datasets show that the proposed method (i)~is robust to OOD-noisy test data and provides improved accuracy and (ii)~quantifies voxel-level uncertainty in the predictions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge