Sunny Virmani
MedGemma 1.5 Technical Report
Apr 06, 2026Abstract:We introduce MedGemma 1.5 4B, the latest model in the MedGemma collection. MedGemma 1.5 expands on MedGemma 1 by integrating additional capabilities: high-dimensional medical imaging (CT/MRI volumes and histopathology whole slide images), anatomical localization via bounding boxes, multi-timepoint chest X-ray analysis, and improved medical document understanding (lab reports, electronic health records). We detail the innovations required to enable these modalities within a single architecture, including new training data, long-context 3D volume slicing, and whole-slide pathology sampling. Compared to MedGemma 1 4B, MedGemma 1.5 4B demonstrates significant gains in these new areas, improving 3D MRI condition classification accuracy by 11% and 3D CT condition classification by 3% (absolute improvements). In whole slide pathology imaging, MedGemma 1.5 4B achieves a 47% macro F1 gain. Additionally, it improves anatomical localization with a 35% increase in Intersection over Union on chest X-rays and achieves a 4% macro accuracy for longitudinal (multi-timepoint) chest x-ray analysis. Beyond its improved multimodal performance over MedGemma 1, MedGemma 1.5 improves on text-based clinical knowledge and reasoning, improving by 5% on MedQA accuracy and 22% on EHRQA accuracy. It also achieves an average of 18% macro F1 on 4 different lab report information extraction datasets (EHR Datasets 2, 3, 4, and Mendeley Clinical Laboratory Test Reports). Taken together, MedGemma 1.5 serves as a robust, open resource for the community, designed as an improved foundation on which developers can create the next generation of medical AI systems. Resources and tutorials for building upon MedGemma 1.5 can be found at https://goo.gle/MedGemma.
Towards Generalist Biomedical AI
Jul 26, 2023



Abstract:Medicine is inherently multimodal, with rich data modalities spanning text, imaging, genomics, and more. Generalist biomedical artificial intelligence (AI) systems that flexibly encode, integrate, and interpret this data at scale can potentially enable impactful applications ranging from scientific discovery to care delivery. To enable the development of these models, we first curate MultiMedBench, a new multimodal biomedical benchmark. MultiMedBench encompasses 14 diverse tasks such as medical question answering, mammography and dermatology image interpretation, radiology report generation and summarization, and genomic variant calling. We then introduce Med-PaLM Multimodal (Med-PaLM M), our proof of concept for a generalist biomedical AI system. Med-PaLM M is a large multimodal generative model that flexibly encodes and interprets biomedical data including clinical language, imaging, and genomics with the same set of model weights. Med-PaLM M reaches performance competitive with or exceeding the state of the art on all MultiMedBench tasks, often surpassing specialist models by a wide margin. We also report examples of zero-shot generalization to novel medical concepts and tasks, positive transfer learning across tasks, and emergent zero-shot medical reasoning. To further probe the capabilities and limitations of Med-PaLM M, we conduct a radiologist evaluation of model-generated (and human) chest X-ray reports and observe encouraging performance across model scales. In a side-by-side ranking on 246 retrospective chest X-rays, clinicians express a pairwise preference for Med-PaLM M reports over those produced by radiologists in up to 40.50% of cases, suggesting potential clinical utility. While considerable work is needed to validate these models in real-world use cases, our results represent a milestone towards the development of generalist biomedical AI systems.
Predicting Risk of Developing Diabetic Retinopathy using Deep Learning
Aug 10, 2020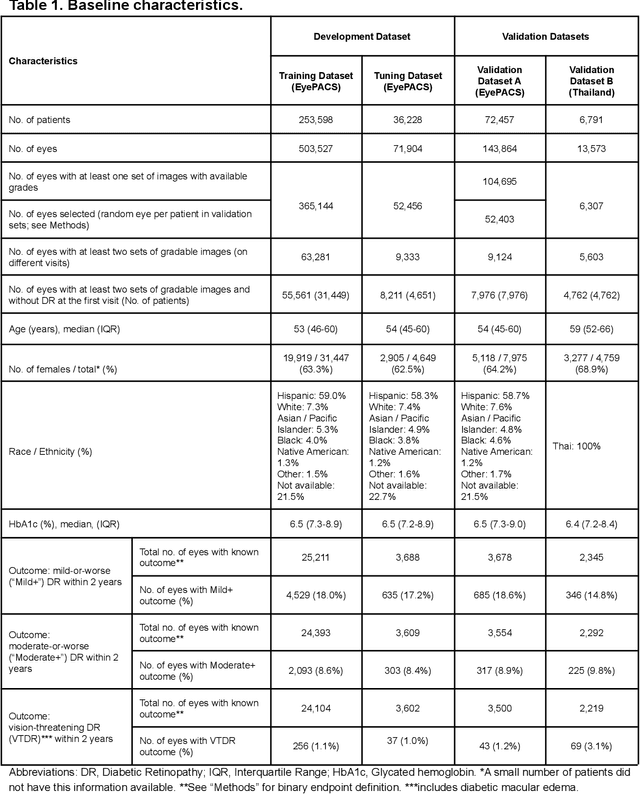
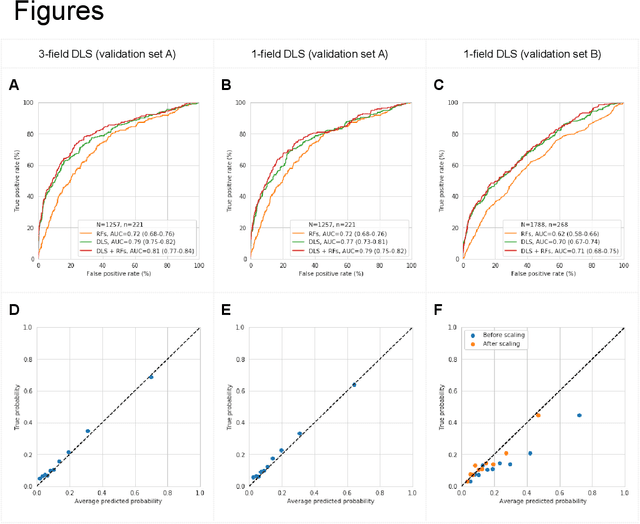
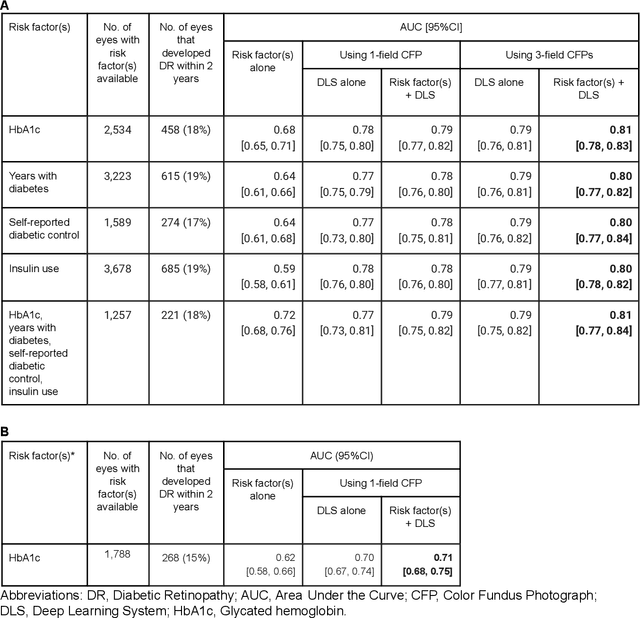
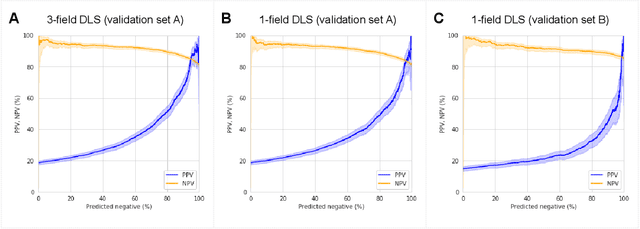
Abstract:Diabetic retinopathy (DR) screening is instrumental in preventing blindness, but faces a scaling challenge as the number of diabetic patients rises. Risk stratification for the development of DR may help optimize screening intervals to reduce costs while improving vision-related outcomes. We created and validated two versions of a deep learning system (DLS) to predict the development of mild-or-worse ("Mild+") DR in diabetic patients undergoing DR screening. The two versions used either three-fields or a single field of color fundus photographs (CFPs) as input. The training set was derived from 575,431 eyes, of which 28,899 had known 2-year outcome, and the remaining were used to augment the training process via multi-task learning. Validation was performed on both an internal validation set (set A; 7,976 eyes; 3,678 with known outcome) and an external validation set (set B; 4,762 eyes; 2,345 with known outcome). For predicting 2-year development of DR, the 3-field DLS had an area under the receiver operating characteristic curve (AUC) of 0.79 (95%CI, 0.78-0.81) on validation set A. On validation set B (which contained only a single field), the 1-field DLS's AUC was 0.70 (95%CI, 0.67-0.74). The DLS was prognostic even after adjusting for available risk factors (p<0.001). When added to the risk factors, the 3-field DLS improved the AUC from 0.72 (95%CI, 0.68-0.76) to 0.81 (95%CI, 0.77-0.84) in validation set A, and the 1-field DLS improved the AUC from 0.62 (95%CI, 0.58-0.66) to 0.71 (95%CI, 0.68-0.75) in validation set B. The DLSs in this study identified prognostic information for DR development from CFPs. This information is independent of and more informative than the available risk factors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge