Stephanie E. Combs
Atomic Fact-Checking Increases Clinician Trust in Large Language Model Recommendations for Oncology Decision Support: A Randomized Controlled Trial
May 05, 2026Abstract:Question: Does atomic fact-checking, which decomposes AI treatment recommendations into individually verifiable claims linked to source guideline documents, increase clinician trust compared to traditional explainability approaches? Findings: In this randomized trial of 356 clinicians generating 7,476 trust ratings, atomic fact-checking produced a large effect on trust (Cohen's d = 0.94), increasing the proportion of clinicians expressing trust from 26.9% to 66.5%. Traditional transparency mechanisms showed a dose-response gradient of improvement over baseline (d = 0.25 to 0.50). Meaning: Decomposing AI recommendations into individually verifiable claims linked to source guidelines produces substantially higher clinician trust than traditional explainability approaches in high-stakes clinical decisions.
Deep-Learning Atlas Registration for Melanoma Brain Metastases: Preserving Pathology While Enabling Cohort-Level Analyses
Feb 13, 2026Abstract:Melanoma brain metastases (MBM) are common and spatially heterogeneous lesions, complicating cohort-level analyses due to anatomical variability and differing MRI protocols. We propose a fully differentiable, deep-learning-based deformable registration framework that aligns individual pathological brains to a common atlas while preserving metastatic tissue without requiring lesion masks or preprocessing. Missing anatomical correspondences caused by metastases are handled through a forward-model similarity metric based on distance-transformed anatomical labels, combined with a volume-preserving regularization term to ensure deformation plausibility. Registration performance was evaluated using Dice coefficient (DSC), Hausdorff distance (HD), average symmetric surface distance (ASSD), and Jacobian-based measures. The method was applied to 209 MBM patients from three centres, enabling standardized mapping of metastases to anatomical, arterial, and perfusion atlases. The framework achieved high registration accuracy across datasets (DSC 0.89-0.92, HD 6.79-7.60 mm, ASSD 0.63-0.77 mm) while preserving metastatic volumes. Spatial analysis demonstrated significant over-representation of MBM in the cerebral cortex and putamen, under-representation in white matter, and consistent localization near the gray-white matter junction. No arterial territory showed increased metastasis frequency after volume correction. This approach enables robust atlas registration of pathological brain MRI without lesion masks and supports reproducible multi-centre analyses. Applied to MBM, it confirms and refines known spatial predilections, particularly preferential seeding near the gray-white matter junction and cortical regions. The publicly available implementation facilitates reproducible research and extension to other brain tumours and neurological pathologies.
Improving Reliability and Explainability of Medical Question Answering through Atomic Fact Checking in Retrieval-Augmented LLMs
May 30, 2025



Abstract:Large language models (LLMs) exhibit extensive medical knowledge but are prone to hallucinations and inaccurate citations, which pose a challenge to their clinical adoption and regulatory compliance. Current methods, such as Retrieval Augmented Generation, partially address these issues by grounding answers in source documents, but hallucinations and low fact-level explainability persist. In this work, we introduce a novel atomic fact-checking framework designed to enhance the reliability and explainability of LLMs used in medical long-form question answering. This method decomposes LLM-generated responses into discrete, verifiable units called atomic facts, each of which is independently verified against an authoritative knowledge base of medical guidelines. This approach enables targeted correction of errors and direct tracing to source literature, thereby improving the factual accuracy and explainability of medical Q&A. Extensive evaluation using multi-reader assessments by medical experts and an automated open Q&A benchmark demonstrated significant improvements in factual accuracy and explainability. Our framework achieved up to a 40% overall answer improvement and a 50% hallucination detection rate. The ability to trace each atomic fact back to the most relevant chunks from the database provides a granular, transparent explanation of the generated responses, addressing a major gap in current medical AI applications. This work represents a crucial step towards more trustworthy and reliable clinical applications of LLMs, addressing key prerequisites for clinical application and fostering greater confidence in AI-assisted healthcare.
Analysis of clinical, dosimetric and radiomic features for predicting local failure after stereotactic radiotherapy of brain metastases in malignant melanoma
May 31, 2024



Abstract:Background: The aim of this study was to investigate the role of clinical, dosimetric and pretherapeutic magnetic resonance imaging (MRI) features for lesion-specific outcome prediction of stereotactic radiotherapy (SRT) in patients with brain metastases from malignant melanoma (MBM). Methods: In this multicenter, retrospective analysis, we reviewed 517 MBM from 130 patients treated with SRT (single fraction or hypofractionated). For each gross tumor volume (GTV) 1576 radiomic features (RF) were calculated (788 each for the GTV and for a 3 mm margin around the GTV). Clinical parameters, radiation dose and RF from pretherapeutic contrast-enhanced T1-weighted MRI from different institutions were evaluated with a feature processing and elimination pipeline in a nested cross-validation scheme. Results: Seventy-two (72) of 517 lesions (13.9%) showed a local failure (LF) after SRT. The processing pipeline showed clinical, dosimetric and radiomic features providing information for LF prediction. The most prominent ones were the correlation of the gray level co-occurrence matrix of the margin (hazard ratio (HR): 0.37, confidence interval (CI): 0.23-0.58) and systemic therapy before SRT (HR: 0.55, CI: 0.42-0.70). The majority of RF associated with LF was calculated in the margin around the GTV. Conclusions: Pretherapeutic MRI based RF connected with lesion-specific outcome after SRT could be identified, despite multicentric data and minor differences in imaging protocols. Image data analysis of the surrounding metastatic environment may provide therapy-relevant information with the potential to further individualize radiotherapy strategies.
All Sizes Matter: Improving Volumetric Brain Segmentation on Small Lesions
Oct 04, 2023



Abstract:Brain metastases (BMs) are the most frequently occurring brain tumors. The treatment of patients having multiple BMs with stereo tactic radiosurgery necessitates accurate localization of the metastases. Neural networks can assist in this time-consuming and costly task that is typically performed by human experts. Particularly challenging is the detection of small lesions since they are often underrepresented in exist ing approaches. Yet, lesion detection is equally important for all sizes. In this work, we develop an ensemble of neural networks explicitly fo cused on detecting and segmenting small BMs. To accomplish this task, we trained several neural networks focusing on individual aspects of the BM segmentation problem: We use blob loss that specifically addresses the imbalance of lesion instances in terms of size and texture and is, therefore, not biased towards larger lesions. In addition, a model using a subtraction sequence between the T1 and T1 contrast-enhanced sequence focuses on low-contrast lesions. Furthermore, we train additional models only on small lesions. Our experiments demonstrate the utility of the ad ditional blob loss and the subtraction sequence. However, including the specialized small lesion models in the ensemble deteriorates segmentation results. We also find domain-knowledge-inspired postprocessing steps to drastically increase our performance in most experiments. Our approach enables us to submit a competitive challenge entry to the ASNR-MICCAI BraTS Brain Metastasis Challenge 2023.
A unified 3D framework for Organs at Risk Localization and Segmentation for Radiation Therapy Planning
Mar 01, 2022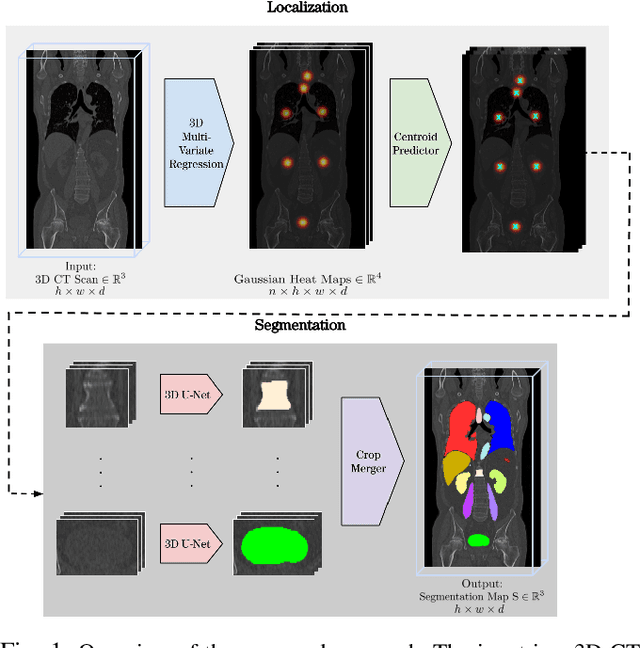
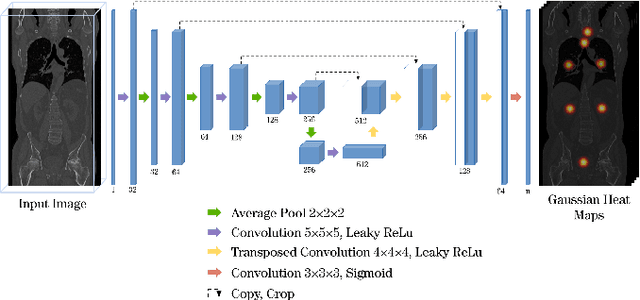
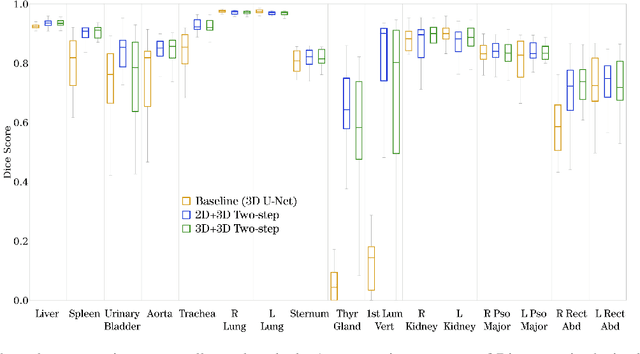
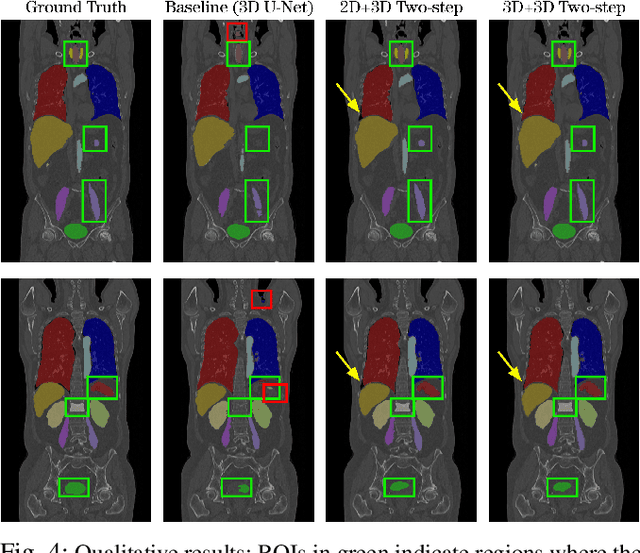
Abstract:Automatic localization and segmentation of organs-at-risk (OAR) in CT are essential pre-processing steps in medical image analysis tasks, such as radiation therapy planning. For instance, the segmentation of OAR surrounding tumors enables the maximization of radiation to the tumor area without compromising the healthy tissues. However, the current medical workflow requires manual delineation of OAR, which is prone to errors and is annotator-dependent. In this work, we aim to introduce a unified 3D pipeline for OAR localization-segmentation rather than novel localization or segmentation architectures. To the best of our knowledge, our proposed framework fully enables the exploitation of 3D context information inherent in medical imaging. In the first step, a 3D multi-variate regression network predicts organs' centroids and bounding boxes. Secondly, 3D organ-specific segmentation networks are leveraged to generate a multi-organ segmentation map. Our method achieved an overall Dice score of $0.9260\pm 0.18 \%$ on the VISCERAL dataset containing CT scans with varying fields of view and multiple organs.
Evaluating the Robustness of Self-Supervised Learning in Medical Imaging
May 14, 2021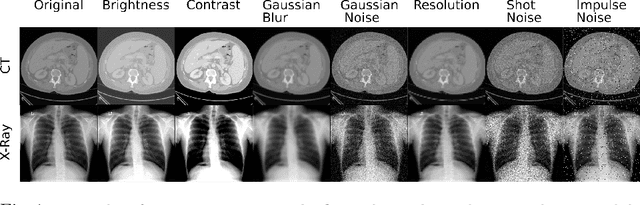
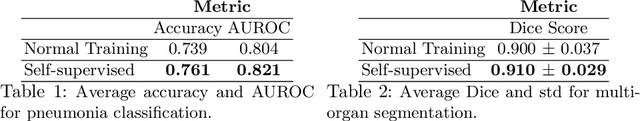

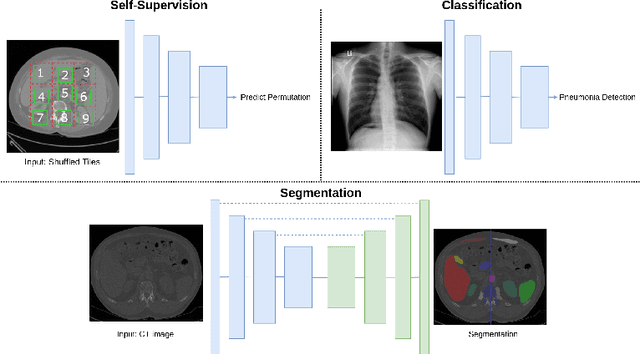
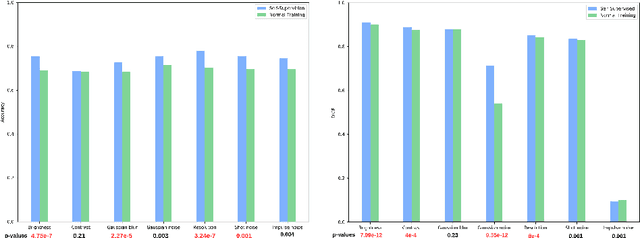
Abstract:Self-supervision has demonstrated to be an effective learning strategy when training target tasks on small annotated data-sets. While current research focuses on creating novel pretext tasks to learn meaningful and reusable representations for the target task, these efforts obtain marginal performance gains compared to fully-supervised learning. Meanwhile, little attention has been given to study the robustness of networks trained in a self-supervised manner. In this work, we demonstrate that networks trained via self-supervised learning have superior robustness and generalizability compared to fully-supervised learning in the context of medical imaging. Our experiments on pneumonia detection in X-rays and multi-organ segmentation in CT yield consistent results exposing the hidden benefits of self-supervision for learning robust feature representations.
Deep Learning Based HPV Status Prediction for Oropharyngeal Cancer Patients
Nov 17, 2020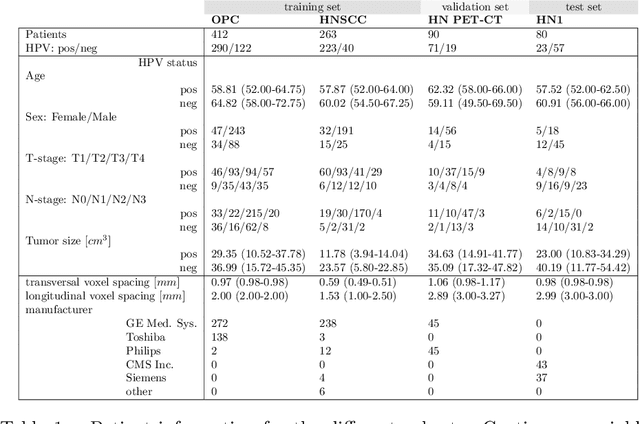
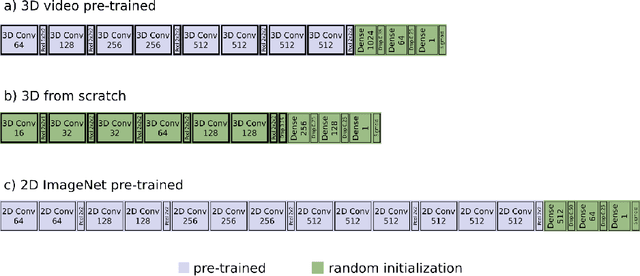
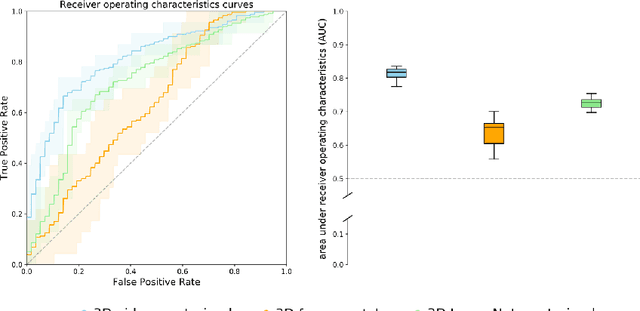
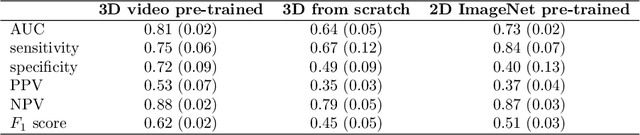
Abstract:We investigated the ability of deep learning models for imaging based HPV status detection. To overcome the problem of small medical datasets we used a transfer learning approach. A 3D convolutional network pre-trained on sports video clips was fine tuned such that full 3D information in the CT images could be exploited. The video pre-trained model was able to differentiate HPV-positive from HPV-negative cases with an area under the receiver operating characteristic curve (AUC) of 0.81 for an external test set. In comparison to a 3D convolutional neural network (CNN) trained from scratch and a 2D architecture pre-trained on ImageNet the video pre-trained model performed best.
Deep Reinforcement Learning for Organ Localization in CT
May 11, 2020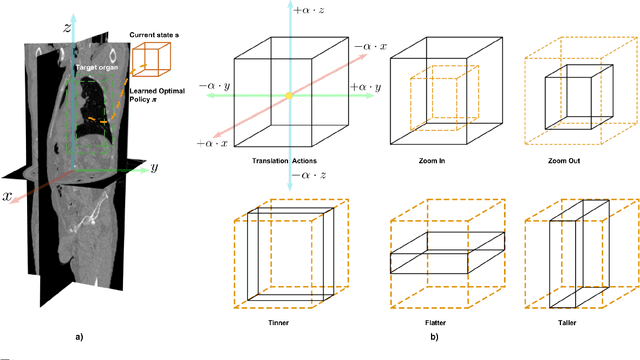
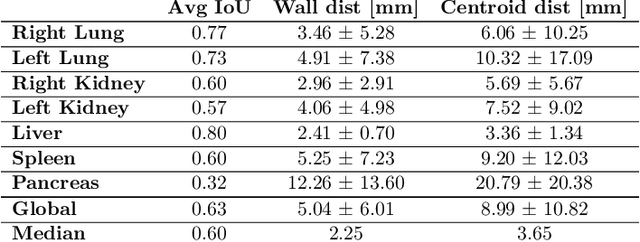
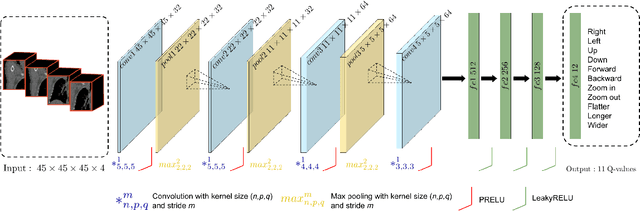
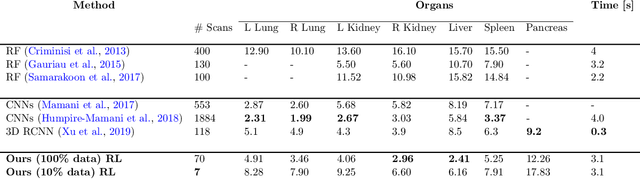
Abstract:Robust localization of organs in computed tomography scans is a constant pre-processing requirement for organ-specific image retrieval, radiotherapy planning, and interventional image analysis. In contrast to current solutions based on exhaustive search or region proposals, which require large amounts of annotated data, we propose a deep reinforcement learning approach for organ localization in CT. In this work, an artificial agent is actively self-taught to localize organs in CT by learning from its asserts and mistakes. Within the context of reinforcement learning, we propose a novel set of actions tailored for organ localization in CT. Our method can use as a plug-and-play module for localizing any organ of interest. We evaluate the proposed solution on the public VISCERAL dataset containing CT scans with varying fields of view and multiple organs. We achieved an overall intersection over union of 0.63, an absolute median wall distance of 2.25 mm, and a median distance between centroids of 3.65 mm.
* Accepted paper in MIDL 2020
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge