Daniel M. Lang
The Mean is the Mirage: Entropy-Adaptive Model Merging under Heterogeneous Domain Shifts in Medical Imaging
Feb 24, 2026Abstract:Model merging under unseen test-time distribution shifts often renders naive strategies, such as mean averaging unreliable. This challenge is especially acute in medical imaging, where models are fine-tuned locally at clinics on private data, producing domain-specific models that differ by scanner, protocol, and population. When deployed at an unseen clinical site, test cases arrive in unlabeled, non-i.i.d. batches, and the model must adapt immediately without labels. In this work, we introduce an entropy-adaptive, fully online model-merging method that yields a batch-specific merged model via only forward passes, effectively leveraging target information. We further demonstrate why mean merging is prone to failure and misaligned under heterogeneous domain shifts. Next, we mitigate encoder classifier mismatch by decoupling the encoder and classification head, merging with separate merging coefficients. We extensively evaluate our method with state-of-the-art baselines using two backbones across nine medical and natural-domain generalization image classification datasets, showing consistent gains across standard evaluation and challenging scenarios. These performance gains are achieved while retaining single-model inference at test-time, thereby demonstrating the effectiveness of our method.
TomoGraphView: 3D Medical Image Classification with Omnidirectional Slice Representations and Graph Neural Networks
Nov 12, 2025Abstract:The growing number of medical tomography examinations has necessitated the development of automated methods capable of extracting comprehensive imaging features to facilitate downstream tasks such as tumor characterization, while assisting physicians in managing their growing workload. However, 3D medical image classification remains a challenging task due to the complex spatial relationships and long-range dependencies inherent in volumetric data. Training models from scratch suffers from low data regimes, and the absence of 3D large-scale multimodal datasets has limited the development of 3D medical imaging foundation models. Recent studies, however, have highlighted the potential of 2D vision foundation models, originally trained on natural images, as powerful feature extractors for medical image analysis. Despite these advances, existing approaches that apply 2D models to 3D volumes via slice-based decomposition remain suboptimal. Conventional volume slicing strategies, which rely on canonical planes such as axial, sagittal, or coronal, may inadequately capture the spatial extent of target structures when these are misaligned with standardized viewing planes. Furthermore, existing slice-wise aggregation strategies rarely account for preserving the volumetric structure, resulting in a loss of spatial coherence across slices. To overcome these limitations, we propose TomoGraphView, a novel framework that integrates omnidirectional volume slicing with spherical graph-based feature aggregation. We publicly share our accessible code base at http://github.com/compai-lab/2025-MedIA-kiechle and provide a user-friendly library for omnidirectional volume slicing at https://pypi.org/project/OmniSlicer.
Simulating Dynamic Tumor Contrast Enhancement in Breast MRI using Conditional Generative Adversarial Networks
Sep 27, 2024Abstract:This paper presents a method for virtual contrast enhancement in breast MRI, offering a promising non-invasive alternative to traditional contrast agent-based DCE-MRI acquisition. Using a conditional generative adversarial network, we predict DCE-MRI images, including jointly-generated sequences of multiple corresponding DCE-MRI timepoints, from non-contrast-enhanced MRIs, enabling tumor localization and characterization without the associated health risks. Furthermore, we qualitatively and quantitatively evaluate the synthetic DCE-MRI images, proposing a multi-metric Scaled Aggregate Measure (SAMe), assessing their utility in a tumor segmentation downstream task, and conclude with an analysis of the temporal patterns in multi-sequence DCE-MRI generation. Our approach demonstrates promising results in generating realistic and useful DCE-MRI sequences, highlighting the potential of virtual contrast enhancement for improving breast cancer diagnosis and treatment, particularly for patients where contrast agent administration is contraindicated.
Graph Neural Networks: A suitable Alternative to MLPs in Latent 3D Medical Image Classification?
Jul 24, 2024



Abstract:Recent studies have underscored the capabilities of natural imaging foundation models to serve as powerful feature extractors, even in a zero-shot setting for medical imaging data. Most commonly, a shallow multi-layer perceptron (MLP) is appended to the feature extractor to facilitate end-to-end learning and downstream prediction tasks such as classification, thus representing the de facto standard. However, as graph neural networks (GNNs) have become a practicable choice for various tasks in medical research in the recent past, we direct attention to the question of how effective GNNs are compared to MLP prediction heads for the task of 3D medical image classification, proposing them as a potential alternative. In our experiments, we devise a subject-level graph for each volumetric dataset instance. Therein latent representations of all slices in the volume, encoded through a DINOv2 pretrained vision transformer (ViT), constitute the nodes and their respective node features. We use public datasets to compare the classification heads numerically and evaluate various graph construction and graph convolution methods in our experiments. Our findings show enhancements of the GNN in classification performance and substantial improvements in runtime compared to an MLP prediction head. Additional robustness evaluations further validate the promising performance of the GNN, promoting them as a suitable alternative to traditional MLP classification heads. Our code is publicly available at: https://github.com/compai-lab/2024-miccai-grail-kiechle
MedEdit: Counterfactual Diffusion-based Image Editing on Brain MRI
Jul 21, 2024


Abstract:Denoising diffusion probabilistic models enable high-fidelity image synthesis and editing. In biomedicine, these models facilitate counterfactual image editing, producing pairs of images where one is edited to simulate hypothetical conditions. For example, they can model the progression of specific diseases, such as stroke lesions. However, current image editing techniques often fail to generate realistic biomedical counterfactuals, either by inadequately modeling indirect pathological effects like brain atrophy or by excessively altering the scan, which disrupts correspondence to the original images. Here, we propose MedEdit, a conditional diffusion model for medical image editing. MedEdit induces pathology in specific areas while balancing the modeling of disease effects and preserving the integrity of the original scan. We evaluated MedEdit on the Atlas v2.0 stroke dataset using Frechet Inception Distance and Dice scores, outperforming state-of-the-art diffusion-based methods such as Palette (by 45%) and SDEdit (by 61%). Additionally, clinical evaluations by a board-certified neuroradiologist confirmed that MedEdit generated realistic stroke scans indistinguishable from real ones. We believe this work will enable counterfactual image editing research to further advance the development of realistic and clinically useful imaging tools.
Enhancing the Utility of Privacy-Preserving Cancer Classification using Synthetic Data
Jul 17, 2024Abstract:Deep learning holds immense promise for aiding radiologists in breast cancer detection. However, achieving optimal model performance is hampered by limitations in availability and sharing of data commonly associated to patient privacy concerns. Such concerns are further exacerbated, as traditional deep learning models can inadvertently leak sensitive training information. This work addresses these challenges exploring and quantifying the utility of privacy-preserving deep learning techniques, concretely, (i) differentially private stochastic gradient descent (DP-SGD) and (ii) fully synthetic training data generated by our proposed malignancy-conditioned generative adversarial network. We assess these methods via downstream malignancy classification of mammography masses using a transformer model. Our experimental results depict that synthetic data augmentation can improve privacy-utility tradeoffs in differentially private model training. Further, model pretraining on synthetic data achieves remarkable performance, which can be further increased with DP-SGD fine-tuning across all privacy guarantees. With this first in-depth exploration of privacy-preserving deep learning in breast imaging, we address current and emerging clinical privacy requirements and pave the way towards the adoption of private high-utility deep diagnostic models. Our reproducible codebase is publicly available at https://github.com/RichardObi/mammo_dp.
Progressive Growing of Patch Size: Resource-Efficient Curriculum Learning for Dense Prediction Tasks
Jul 11, 2024Abstract:In this work, we introduce Progressive Growing of Patch Size, a resource-efficient implicit curriculum learning approach for dense prediction tasks. Our curriculum approach is defined by growing the patch size during model training, which gradually increases the task's difficulty. We integrated our curriculum into the nnU-Net framework and evaluated the methodology on all 10 tasks of the Medical Segmentation Decathlon. With our approach, we are able to substantially reduce runtime, computational costs, and CO2 emissions of network training compared to classical constant patch size training. In our experiments, the curriculum approach resulted in improved convergence. We are able to outperform standard nnU-Net training, which is trained with constant patch size, in terms of Dice Score on 7 out of 10 MSD tasks while only spending roughly 50% of the original training runtime. To the best of our knowledge, our Progressive Growing of Patch Size is the first successful employment of a sample-length curriculum in the form of patch size in the field of computer vision. Our code is publicly available at https://github.com/compai-lab/2024-miccai-fischer.
Mask the Unknown: Assessing Different Strategies to Handle Weak Annotations in the MICCAI2023 Mediastinal Lymph Node Quantification Challenge
Jun 20, 2024



Abstract:Pathological lymph node delineation is crucial in cancer diagnosis, progression assessment, and treatment planning. The MICCAI 2023 Lymph Node Quantification Challenge published the first public dataset for pathological lymph node segmentation in the mediastinum. As lymph node annotations are expensive, the challenge was formed as a weakly supervised learning task, where only a subset of all lymph nodes in the training set have been annotated. For the challenge submission, multiple methods for training on these weakly supervised data were explored, including noisy label training, loss masking of unlabeled data, and an approach that integrated the TotalSegmentator toolbox as a form of pseudo labeling in order to reduce the number of unknown voxels. Furthermore, multiple public TCIA datasets were incorporated into the training to improve the performance of the deep learning model. Our submitted model achieved a Dice score of 0.628 and an average symmetric surface distance of 5.8~mm on the challenge test set. With our submitted model, we accomplished third rank in the MICCAI2023 LNQ challenge. A finding of our analysis was that the integration of all visible, including non-pathological, lymph nodes improved the overall segmentation performance on pathological lymph nodes of the test set. Furthermore, segmentation models trained only on clinically enlarged lymph nodes, as given in the challenge scenario, could not generalize to smaller pathological lymph nodes. The code and model for the challenge submission are available at \url{https://gitlab.lrz.de/compai/MediastinalLymphNodeSegmentation}.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2024:008
3D Masked Autoencoders with Application to Anomaly Detection in Non-Contrast Enhanced Breast MRI
Mar 10, 2023Abstract:Self-supervised models allow (pre-)training on unlabeled data and therefore have the potential to overcome the need for large annotated cohorts. One leading self-supervised model is the masked autoencoder (MAE) which was developed on natural imaging data. The MAE is masking out a high fraction of visual transformer (ViT) input patches, to then recover the uncorrupted images as a pretraining task. In this work, we extend MAE to perform anomaly detection on breast magnetic resonance imaging (MRI). This new model, coined masked autoencoder for medical imaging (MAEMI) is trained on two non-contrast enhanced MRI sequences, aiming at lesion detection without the need for intravenous injection of contrast media and temporal image acquisition. During training, only non-cancerous images are presented to the model, with the purpose of localizing anomalous tumor regions during test time. We use a public dataset for model development. Performance of the architecture is evaluated in reference to subtraction images created from dynamic contrast enhanced (DCE)-MRI.
Deep Learning Based HPV Status Prediction for Oropharyngeal Cancer Patients
Nov 17, 2020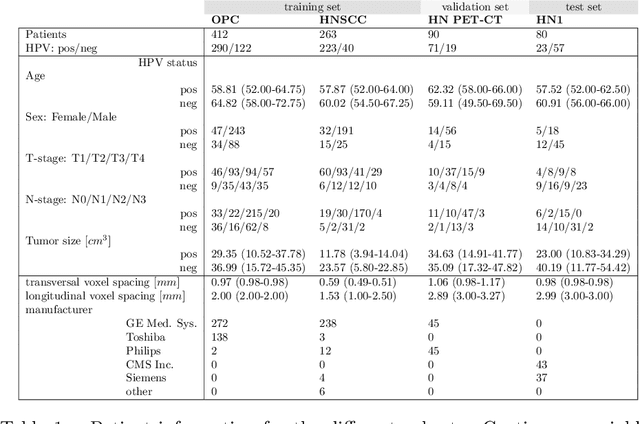
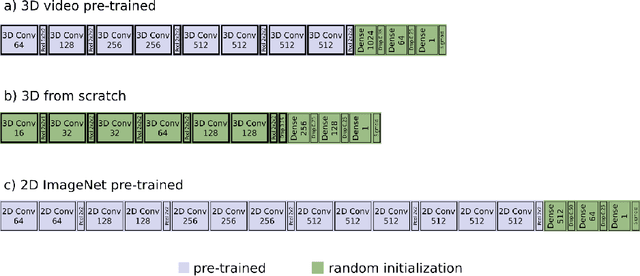
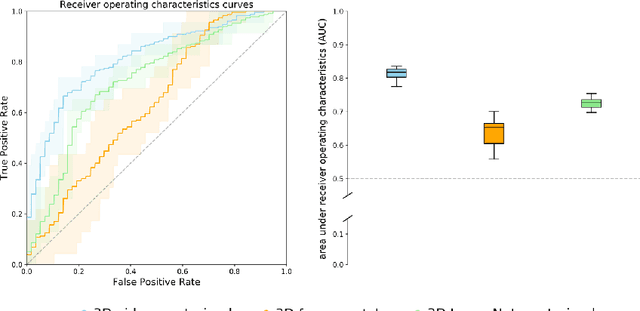
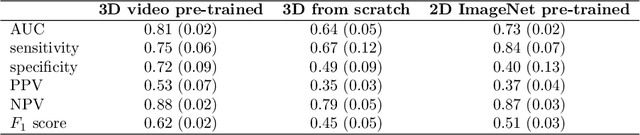
Abstract:We investigated the ability of deep learning models for imaging based HPV status detection. To overcome the problem of small medical datasets we used a transfer learning approach. A 3D convolutional network pre-trained on sports video clips was fine tuned such that full 3D information in the CT images could be exploited. The video pre-trained model was able to differentiate HPV-positive from HPV-negative cases with an area under the receiver operating characteristic curve (AUC) of 0.81 for an external test set. In comparison to a 3D convolutional neural network (CNN) trained from scratch and a 2D architecture pre-trained on ImageNet the video pre-trained model performed best.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge