Sotirios A. Tsaftaris
Parameter-Efficient Fine-Tuning for Medical Image Analysis: The Missed Opportunity
May 24, 2023



Abstract:We present a comprehensive evaluation of Parameter-Efficient Fine-Tuning (PEFT) techniques for diverse medical image analysis tasks. PEFT is increasingly exploited as a valuable approach for knowledge transfer from pre-trained models in natural language processing, vision, speech, and cross-modal tasks, such as vision-language and text-to-image generation. However, its application in medical image analysis remains relatively unexplored. As foundation models are increasingly exploited in the medical domain, it is crucial to investigate and comparatively assess various strategies for knowledge transfer that can bolster a range of downstream tasks. Our study, the first of its kind (to the best of our knowledge), evaluates 16 distinct PEFT methodologies proposed for convolutional and transformer-based networks, focusing on image classification and text-to-image generation tasks across six medical datasets ranging in size, modality, and complexity. Through a battery of more than 600 controlled experiments, we demonstrate performance gains of up to 22% under certain scenarios and demonstrate the efficacy of PEFT for medical text-to-image generation. Further, we reveal the instances where PEFT methods particularly dominate over conventional fine-tuning approaches by studying their relationship with downstream data volume.
Understanding metric-related pitfalls in image analysis validation
Feb 09, 2023Abstract:Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, the present work provides the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
The role of noise in denoising models for anomaly detection in medical images
Jan 19, 2023



Abstract:Pathological brain lesions exhibit diverse appearance in brain images, in terms of intensity, texture, shape, size, and location. Comprehensive sets of data and annotations are difficult to acquire. Therefore, unsupervised anomaly detection approaches have been proposed using only normal data for training, with the aim of detecting outlier anomalous voxels at test time. Denoising methods, for instance classical denoising autoencoders (DAEs) and more recently emerging diffusion models, are a promising approach, however naive application of pixelwise noise leads to poor anomaly detection performance. We show that optimization of the spatial resolution and magnitude of the noise improves the performance of different model training regimes, with similar noise parameter adjustments giving good performance for both DAEs and diffusion models. Visual inspection of the reconstructions suggests that the training noise influences the trade-off between the extent of the detail that is reconstructed and the extent of erasure of anomalies, both of which contribute to better anomaly detection performance. We validate our findings on two real-world datasets (tumor detection in brain MRI and hemorrhage/ischemia/tumor detection in brain CT), showing good detection on diverse anomaly appearances. Overall, we find that a DAE trained with coarse noise is a fast and simple method that gives state-of-the-art accuracy. Diffusion models applied to anomaly detection are as yet in their infancy and provide a promising avenue for further research.
Clinically Plausible Pathology-Anatomy Disentanglement in Patient Brain MRI with Structured Variational Priors
Nov 16, 2022



Abstract:We propose a hierarchically structured variational inference model for accurately disentangling observable evidence of disease (e.g. brain lesions or atrophy) from subject-specific anatomy in brain MRIs. With flexible, partially autoregressive priors, our model (1) addresses the subtle and fine-grained dependencies that typically exist between anatomical and pathological generating factors of an MRI to ensure the clinical validity of generated samples; (2) preserves and disentangles finer pathological details pertaining to a patient's disease state. Additionally, we experiment with an alternative training configuration where we provide supervision to a subset of latent units. It is shown that (1) a partially supervised latent space achieves a higher degree of disentanglement between evidence of disease and subject-specific anatomy; (2) when the prior is formulated with an autoregressive structure, knowledge from the supervision can propagate to the unsupervised latent units, resulting in more informative latent representations capable of modelling anatomy-pathology interdependencies.
Rethinking Generalization: The Impact of Annotation Style on Medical Image Segmentation
Oct 31, 2022Abstract:Generalization is an important attribute of machine learning models, particularly for those that are to be deployed in a medical context, where unreliable predictions can have real world consequences. While the failure of models to generalize across datasets is typically attributed to a mismatch in the data distributions, performance gaps are often a consequence of biases in the ``ground-truth" label annotations. This is particularly important in the context of medical image segmentation of pathological structures (e.g. lesions), where the annotation process is much more subjective, and affected by a number underlying factors, including the annotation protocol, rater education/experience, and clinical aims, among others. In this paper, we show that modeling annotation biases, rather than ignoring them, poses a promising way of accounting for differences in annotation style across datasets. To this end, we propose a generalized conditioning framework to (1) learn and account for different annotation styles across multiple datasets using a single model, (2) identify similar annotation styles across different datasets in order to permit their effective aggregation, and (3) fine-tune a fully trained model to a new annotation style with just a few samples. Next, we present an image-conditioning approach to model annotation styles that correlate with specific image features, potentially enabling detection biases to be more easily identified.
Diffusion Models for Causal Discovery via Topological Ordering
Oct 12, 2022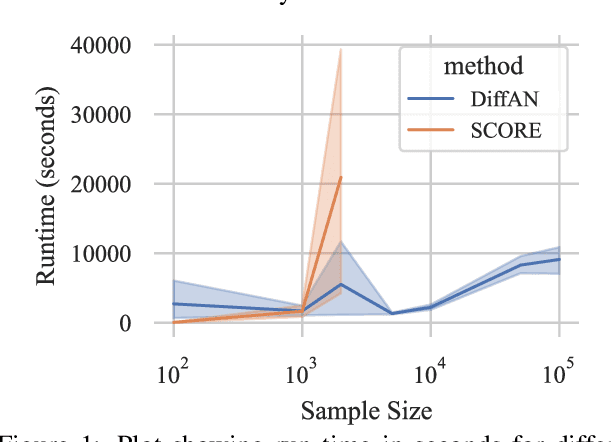
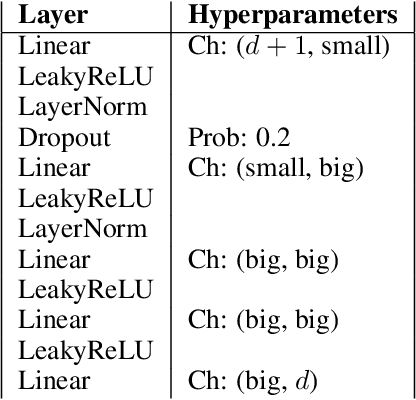
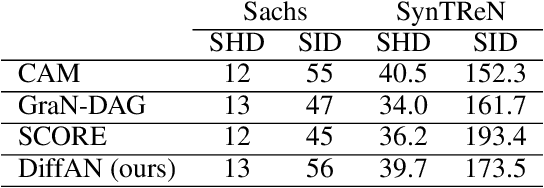
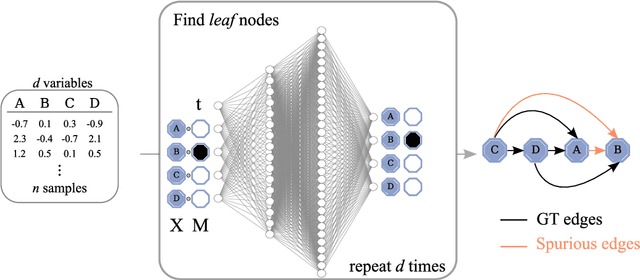

Abstract:Discovering causal relations from observational data becomes possible with additional assumptions such as considering the functional relations to be constrained as nonlinear with additive noise. In this case, the Hessian of the data log-likelihood can be used for finding leaf nodes in a causal graph. Topological ordering approaches for causal discovery exploit this by performing graph discovery in two steps, first sequentially identifying nodes in reverse order of depth (topological ordering), and secondly pruning the potential relations. This is more efficient since the search is performed over a permutation rather than a graph space. However, existing computational methods for obtaining the Hessian still do not scale as the number of variables and the number of samples are increased. Therefore, inspired by recent innovations in diffusion probabilistic models (DPMs), we propose DiffAN, a topological ordering algorithm that leverages DPMs. Further, we introduce theory for updating the learned Hessian without re-training the neural network, and we show that computing with a subset of samples gives an accurate approximation of the ordering, which allows scaling to datasets with more samples and variables. We show empirically that our method scales exceptionally well to datasets with up to $500$ nodes and up to $10^5$ samples while still performing on par over small datasets with state-of-the-art causal discovery methods. Implementation is available at https://github.com/vios-s/DiffAN .
HSIC-InfoGAN: Learning Unsupervised Disentangled Representations by Maximising Approximated Mutual Information
Aug 06, 2022

Abstract:Learning disentangled representations requires either supervision or the introduction of specific model designs and learning constraints as biases. InfoGAN is a popular disentanglement framework that learns unsupervised disentangled representations by maximising the mutual information between latent representations and their corresponding generated images. Maximisation of mutual information is achieved by introducing an auxiliary network and training with a latent regression loss. In this short exploratory paper, we study the use of the Hilbert-Schmidt Independence Criterion (HSIC) to approximate mutual information between latent representation and image, termed HSIC-InfoGAN. Directly optimising the HSIC loss avoids the need for an additional auxiliary network. We qualitatively compare the level of disentanglement in each model, suggest a strategy to tune the hyperparameters of HSIC-InfoGAN, and discuss the potential of HSIC-InfoGAN for medical applications.
What is Healthy? Generative Counterfactual Diffusion for Lesion Localization
Jul 25, 2022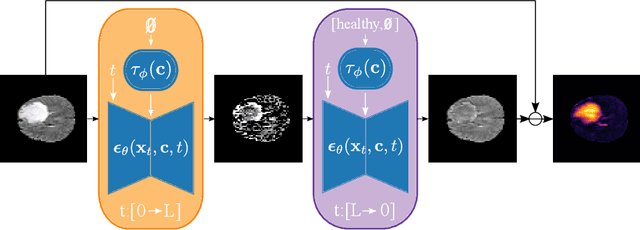
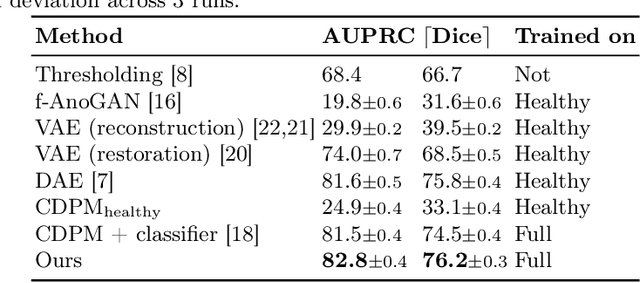
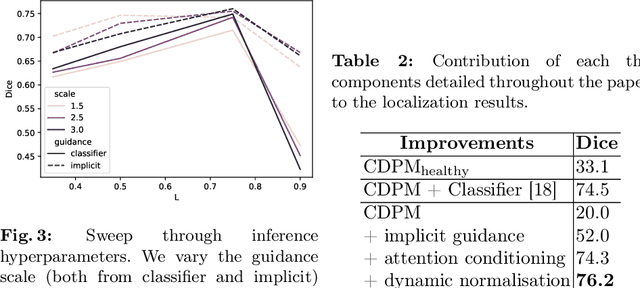
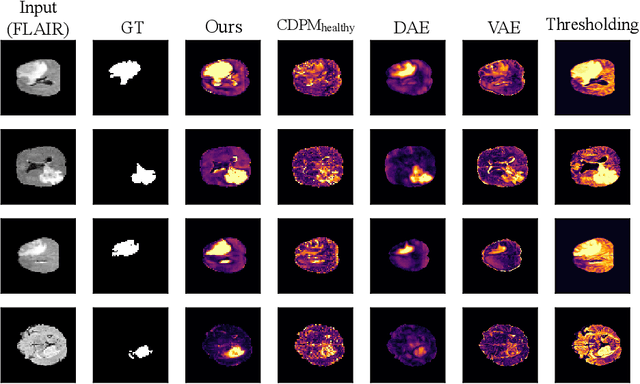
Abstract:Reducing the requirement for densely annotated masks in medical image segmentation is important due to cost constraints. In this paper, we consider the problem of inferring pixel-level predictions of brain lesions by only using image-level labels for training. By leveraging recent advances in generative diffusion probabilistic models (DPM), we synthesize counterfactuals of "How would a patient appear if X pathology was not present?". The difference image between the observed patient state and the healthy counterfactual can be used for inferring the location of pathology. We generate counterfactuals that correspond to the minimal change of the input such that it is transformed to healthy domain. This requires training with healthy and unhealthy data in DPMs. We improve on previous counterfactual DPMs by manipulating the generation process with implicit guidance along with attention conditioning instead of using classifiers. Code is available at https://github.com/vios-s/Diff-SCM.
Why patient data cannot be easily forgotten?
Jun 29, 2022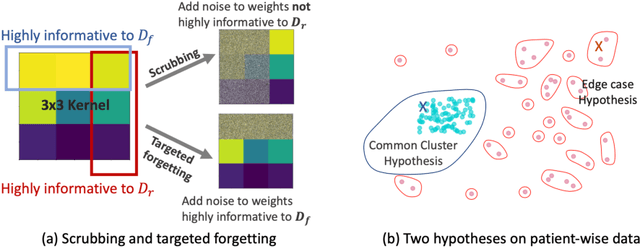
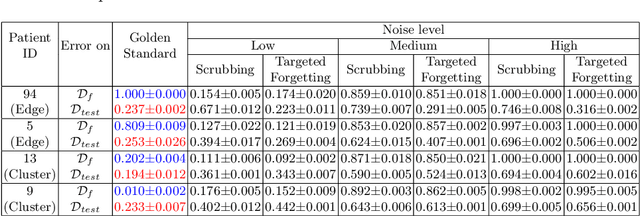
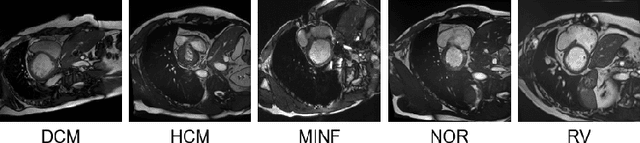
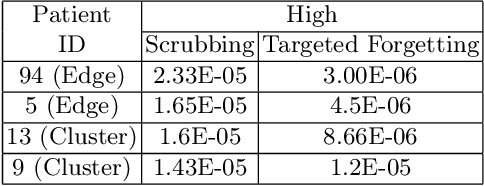
Abstract:Rights provisioned within data protection regulations, permit patients to request that knowledge about their information be eliminated by data holders. With the advent of AI learned on data, one can imagine that such rights can extent to requests for forgetting knowledge of patient's data within AI models. However, forgetting patients' imaging data from AI models, is still an under-explored problem. In this paper, we study the influence of patient data on model performance and formulate two hypotheses for a patient's data: either they are common and similar to other patients or form edge cases, i.e. unique and rare cases. We show that it is not possible to easily forget patient data. We propose a targeted forgetting approach to perform patient-wise forgetting. Extensive experiments on the benchmark Automated Cardiac Diagnosis Challenge dataset showcase the improved performance of the proposed targeted forgetting approach as opposed to a state-of-the-art method.
vMFNet: Compositionality Meets Domain-generalised Segmentation
Jun 29, 2022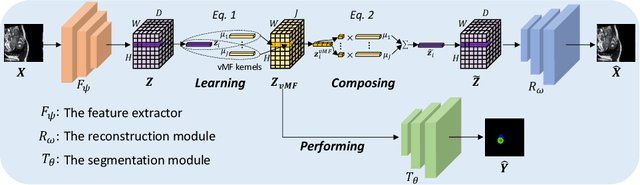
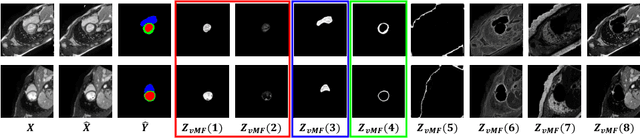
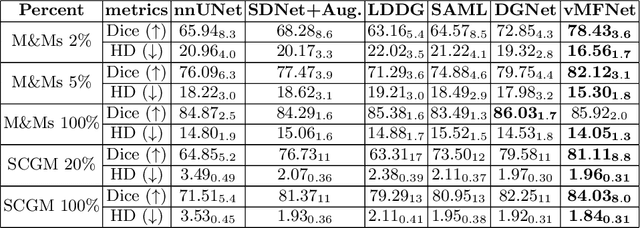

Abstract:Training medical image segmentation models usually requires a large amount of labeled data. By contrast, humans can quickly learn to accurately recognise anatomy of interest from medical (e.g. MRI and CT) images with some limited guidance. Such recognition ability can easily generalise to new images from different clinical centres. This rapid and generalisable learning ability is mostly due to the compositional structure of image patterns in the human brain, which is less incorporated in medical image segmentation. In this paper, we model the compositional components (i.e. patterns) of human anatomy as learnable von-Mises-Fisher (vMF) kernels, which are robust to images collected from different domains (e.g. clinical centres). The image features can be decomposed to (or composed by) the components with the composing operations, i.e. the vMF likelihoods. The vMF likelihoods tell how likely each anatomical part is at each position of the image. Hence, the segmentation mask can be predicted based on the vMF likelihoods. Moreover, with a reconstruction module, unlabeled data can also be used to learn the vMF kernels and likelihoods by recombining them to reconstruct the input image. Extensive experiments show that the proposed vMFNet achieves improved generalisation performance on two benchmarks, especially when annotations are limited. Code is publicly available at: https://github.com/vios-s/vMFNet.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge