Samuel Kadoury
Polytechnique Montreal, Canada, CHU Sainte-Justine Research Center, Montreal, Canada
A Normalized Fully Convolutional Approach to Head and Neck Cancer Outcome Prediction
May 29, 2020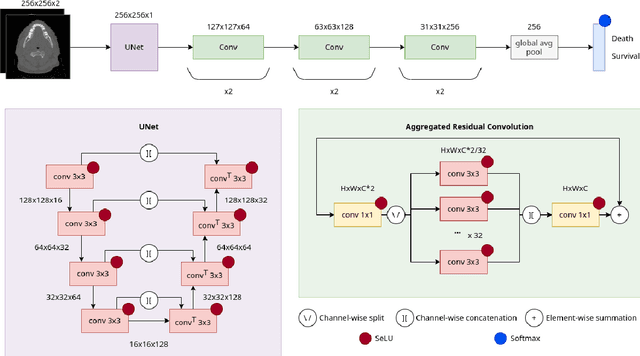
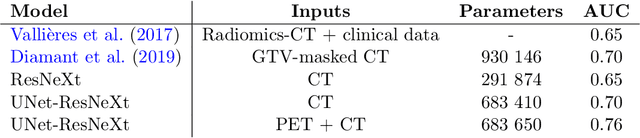

Abstract:In medical imaging, radiological scans of different modalities serve to enhance different sets of features for clinical diagnosis and treatment planning. This variety enriches the source information that could be used for outcome prediction. Deep learning methods are particularly well-suited for feature extraction from high-dimensional inputs such as images. In this work, we apply a CNN classification network augmented with a FCN preprocessor sub-network to a public TCIA head and neck cancer dataset. The training goal is survival prediction of radiotherapy cases based on pre-treatment FDG PET-CT scans, acquired across 4 different hospitals. We show that the preprocessor sub-network in conjunction with aggregated residual connection leads to improvements over state-of-the-art results when combining both CT and PET input images.
Boosting segmentation with weak supervision from image-to-image translation
Apr 04, 2019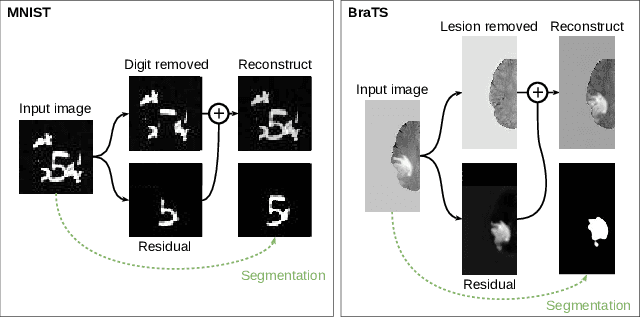

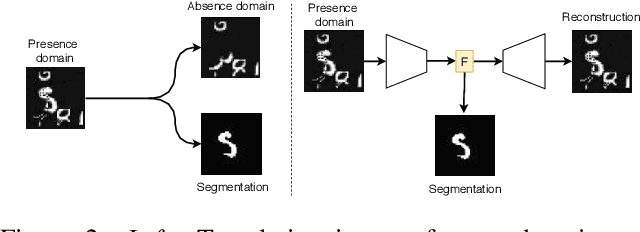
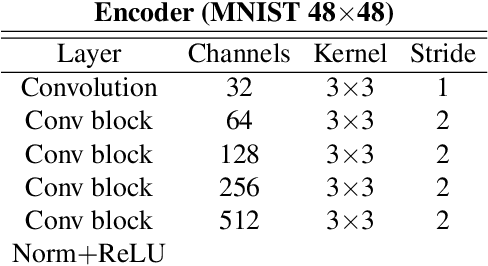
Abstract:In many cases, especially with medical images, it is prohibitively challenging to produce a sufficiently large training sample of pixel-level annotations to train deep neural networks for semantic image segmentation. On the other hand, some information is often known about the contents of images. We leverage information on whether an image presents the segmentation target or whether it is absent from the image to improve segmentation performance by augmenting the amount of data usable for model training. Specifically, we propose a semi-supervised framework that employs image-to-image translation between weak labels (e.g., presence vs. absence of cancer), in addition to fully supervised segmentation on some examples. We conjecture that this translation objective is well aligned with the segmentation objective as both require the same disentangling of image variations. Building on prior image-to-image translation work, we re-use the encoder and decoders for translating in either direction between two domains, employing a strategy of selectively decoding domain-specific variations. For presence vs. absence domains, the encoder produces variations that are common to both and those unique to the presence domain. Furthermore, we successfully re-use one of the decoders used in translation for segmentation. We validate the proposed method on synthetic tasks of varying difficulty as well as on the real task of brain tumor segmentation in magnetic resonance images, where we show significant improvements over standard semi-supervised training with autoencoding.
End-to-End Discriminative Deep Network for Liver Lesion Classification
Jan 28, 2019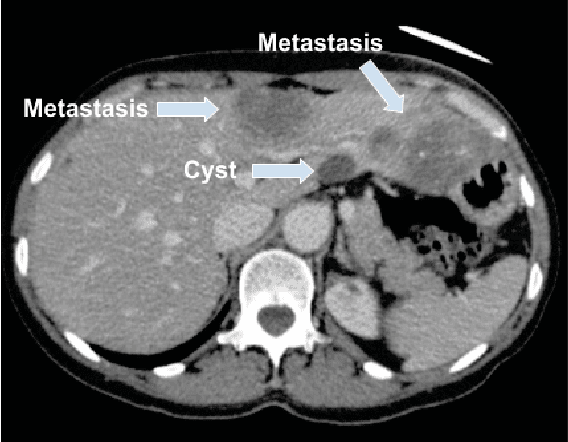
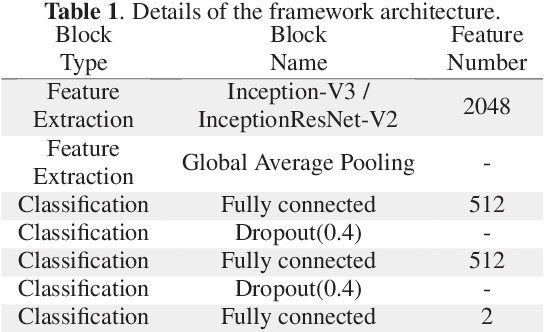
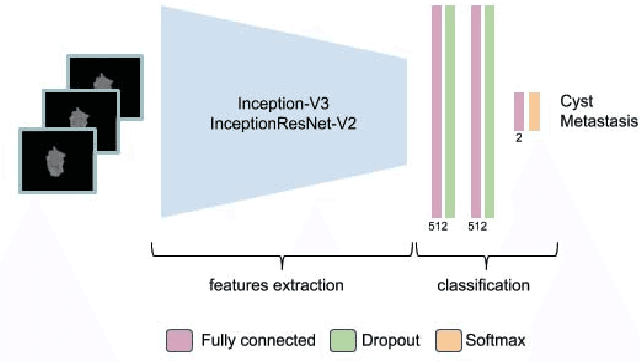
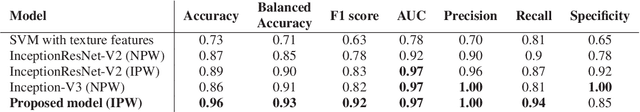
Abstract:Colorectal liver metastasis is one of most aggressive liver malignancies. While the definition of lesion type based on CT images determines the diagnosis and therapeutic strategy, the discrimination between cancerous and non-cancerous lesions are critical and requires highly skilled expertise, experience and time. In the present work we introduce an end-to-end deep learning approach to assist in the discrimination between liver metastases from colorectal cancer and benign cysts in abdominal CT images of the liver. Our approach incorporates the efficient feature extraction of InceptionV3 combined with residual connections and pre-trained weights from ImageNet. The architecture also includes fully connected classification layers to generate a probabilistic output of lesion type. We use an in-house clinical biobank with 230 liver lesions originating from 63 patients. With an accuracy of 0.96 and a F1-score of 0.92, the results obtained with the proposed approach surpasses state of the art methods. Our work provides the basis for incorporating machine learning tools in specialized radiology software to assist physicians in the early detection and treatment of liver lesions.
The Liver Tumor Segmentation Benchmark (LiTS)
Jan 13, 2019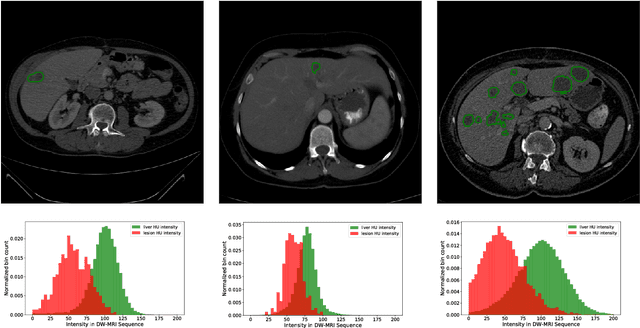
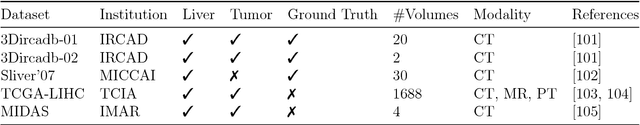

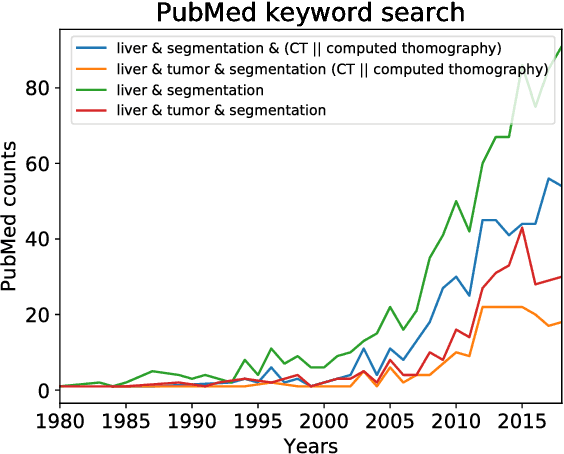
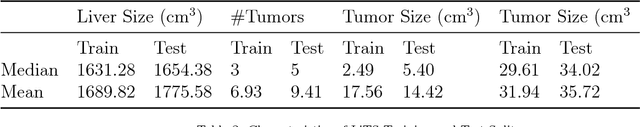
Abstract:In this work, we report the set-up and results of the Liver Tumor Segmentation Benchmark (LITS) organized in conjunction with the IEEE International Symposium on Biomedical Imaging (ISBI) 2016 and International Conference On Medical Image Computing Computer Assisted Intervention (MICCAI) 2017. Twenty four valid state-of-the-art liver and liver tumor segmentation algorithms were applied to a set of 131 computed tomography (CT) volumes with different types of tumor contrast levels (hyper-/hypo-intense), abnormalities in tissues (metastasectomie) size and varying amount of lesions. The submitted algorithms have been tested on 70 undisclosed volumes. The dataset is created in collaboration with seven hospitals and research institutions and manually reviewed by independent three radiologists. We found that not a single algorithm performed best for liver and tumors. The best liver segmentation algorithm achieved a Dice score of 0.96(MICCAI) whereas for tumor segmentation the best algorithm evaluated at 0.67(ISBI) and 0.70(MICCAI). The LITS image data and manual annotations continue to be publicly available through an online evaluation system as an ongoing benchmarking resource.
Multi-Level Batch Normalization In Deep Networks For Invasive Ductal Carcinoma Cell Discrimination In Histopathology Images
Jan 11, 2019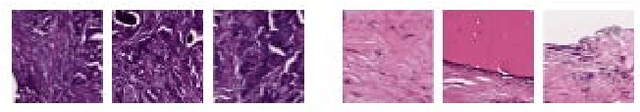
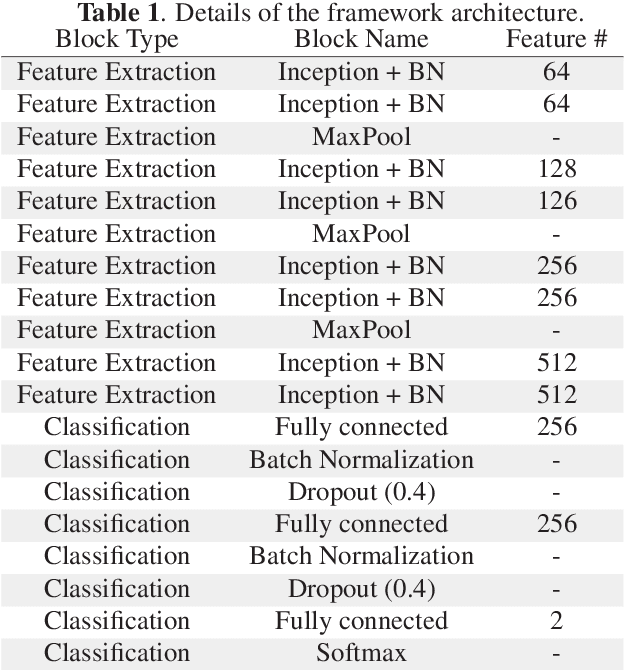
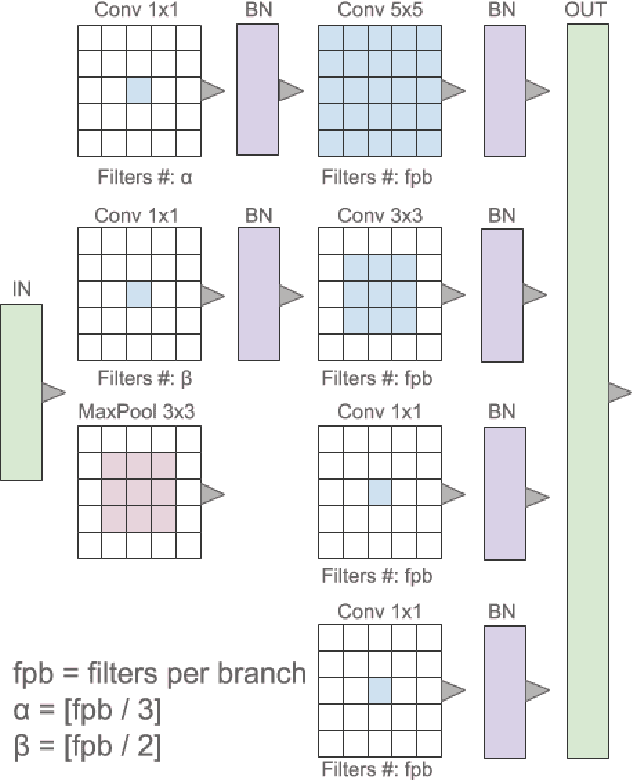
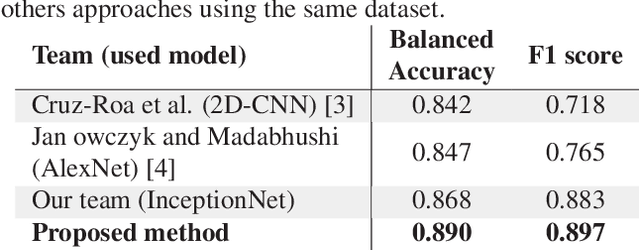
Abstract:Breast cancer is the most diagnosed cancer and the most predominant cause of death in women worldwide. Imaging techniques such as the breast cancer pathology helps in the diagnosis and monitoring of the disease. However identification of malignant cells can be challenging given the high heterogeneity in tissue absorbotion from staining agents. In this work, we present a novel approach for Invasive Ductal Carcinoma (IDC) cells discrimination in histopathology slides. We propose a model derived from the Inception architecture, proposing a multi-level batch normalization module between each convolutional steps. This module was used as a base block for the feature extraction in a CNN architecture. We used the open IDC dataset in which we obtained a balanced accuracy of 0.89 and an F1 score of 0.90, thus surpassing recent state of the art classification algorithms tested on this public dataset.
Liver lesion segmentation informed by joint liver segmentation
Aug 11, 2018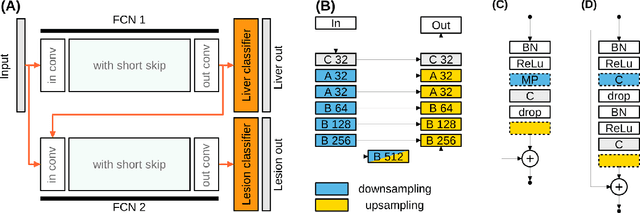
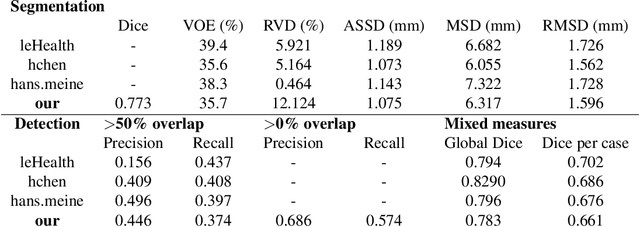
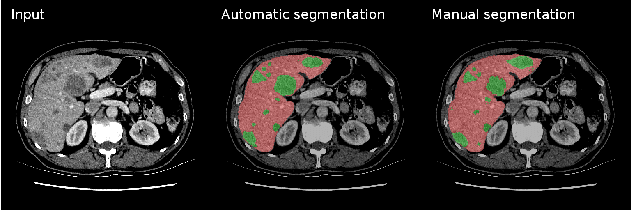
Abstract:We propose a model for the joint segmentation of the liver and liver lesions in computed tomography (CT) volumes. We build the model from two fully convolutional networks, connected in tandem and trained together end-to-end. We evaluate our approach on the 2017 MICCAI Liver Tumour Segmentation Challenge, attaining competitive liver and liver lesion detection and segmentation scores across a wide range of metrics. Unlike other top performing methods, our model output post-processing is trivial, we do not use data external to the challenge, and we propose a simple single-stage model that is trained end-to-end. However, our method nearly matches the top lesion segmentation performance and achieves the second highest precision for lesion detection while maintaining high recall.
Dilatation of Lateral Ventricles with Brain Volumes in Infants with 3D Transfontanelle US
Jun 06, 2018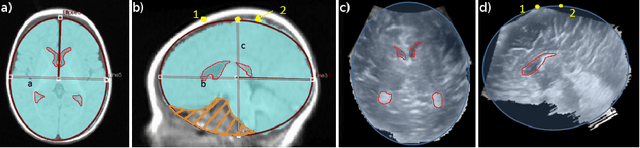
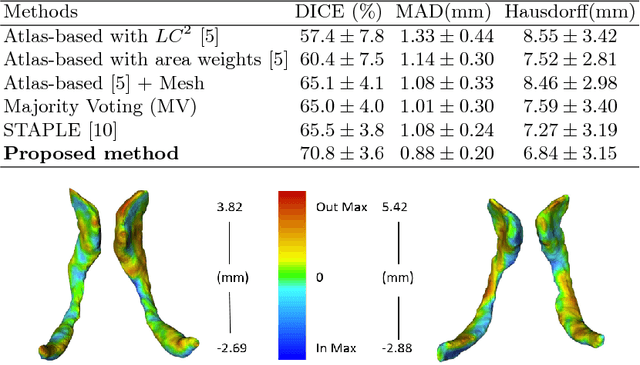
Abstract:Ultrasound (US) can be used to assess brain development in newborns, as MRI is challenging due to immobilization issues, and may require sedation. Dilatation of the lateral ventricles in the brain is a risk factor for poorer neurodevelopment outcomes in infants. Hence, 3D US has the ability to assess the volume of the lateral ventricles similar to clinically standard MRI, but manual segmentation is time consuming. The objective of this study is to develop an approach quantifying the ratio of lateral ventricular dilatation with respect to total brain volume using 3D US, which can assess the severity of macrocephaly. Automatic segmentation of the lateral ventricles is achieved with a multi-atlas deformable registration approach using locally linear correlation metrics for US-MRI fusion, followed by a refinement step using deformable mesh models. Total brain volume is estimated using a 3D ellipsoid modeling approach. Validation was performed on a cohort of 12 infants, ranging from 2 to 8.5 months old, where 3D US and MRI were used to compare brain volumes and segmented lateral ventricles. Automatically extracted volumes from 3D US show a high correlation and no statistically significant difference when compared to ground truth measurements. Differences in volume ratios was 6.0 +/- 4.8% compared to MRI, while lateral ventricular segmentation yielded a mean Dice coefficient of 70.8 +/- 3.6% and a mean absolute distance (MAD) of 0.88 +/- 0.2mm, demonstrating the clinical benefit of this tool in paediatric ultrasound.
Spatiotemporal Manifold Prediction Model for Anterior Vertebral Body Growth Modulation Surgery in Idiopathic Scoliosis
Jun 06, 2018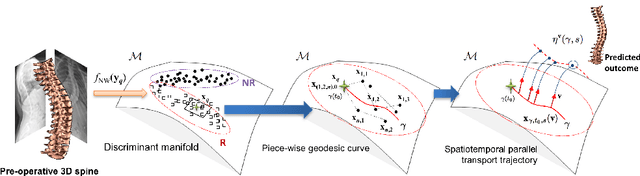
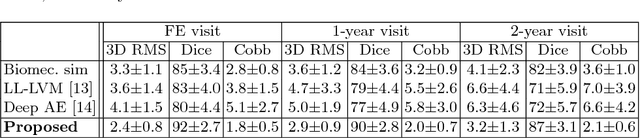
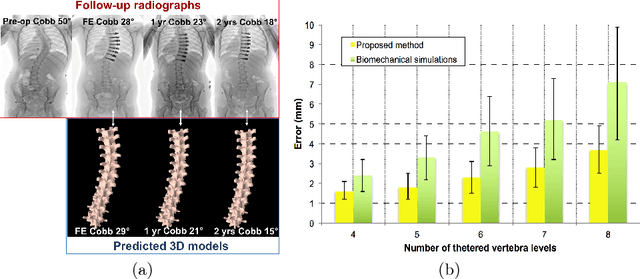
Abstract:Anterior Vertebral Body Growth Modulation (AVBGM) is a minimally invasive surgical technique that gradually corrects spine deformities while preserving lumbar motion. However the selection of potential surgical patients is currently based on clinical judgment and would be facilitated by the identification of patients responding to AVBGM prior to surgery. We introduce a statistical framework for predicting the surgical outcomes following AVBGM in adolescents with idiopathic scoliosis. A discriminant manifold is first constructed to maximize the separation between responsive and non-responsive groups of patients treated with AVBGM for scoliosis. The model then uses subject-specific correction trajectories based on articulated transformations in order to map spine correction profiles to a group-average piecewise-geodesic path. Spine correction trajectories are described in a piecewise-geodesic fashion to account for varying times at follow-up exams, regressing the curve via a quadratic optimization process. To predict the evolution of correction, a baseline reconstruction is projected onto the manifold, from which a spatiotemporal regression model is built from parallel transport curves inferred from neighboring exemplars. The model was trained on 438 reconstructions and tested on 56 subjects using 3D spine reconstructions from follow-up exams, with the probabilistic framework yielding accurate results with differences of 2.1 +/- 0.6deg in main curve angulation, and generating models similar to biomechanical simulations.
On orthogonality and learning recurrent networks with long term dependencies
Oct 12, 2017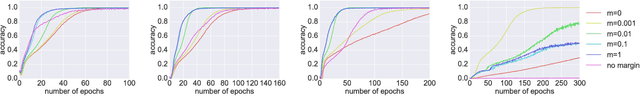
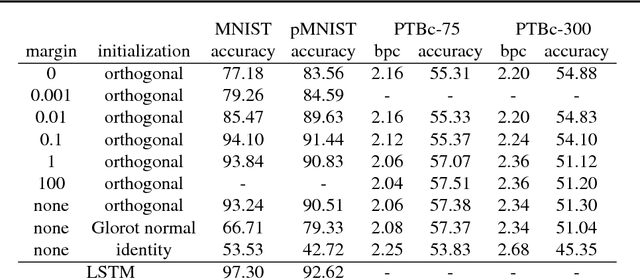
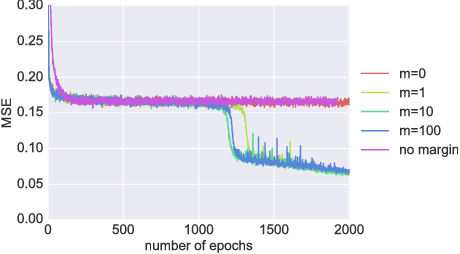
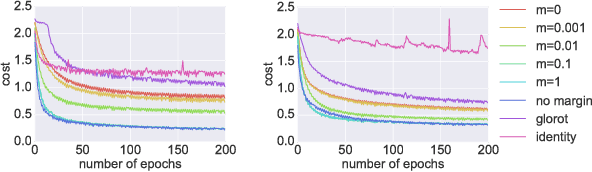
Abstract:It is well known that it is challenging to train deep neural networks and recurrent neural networks for tasks that exhibit long term dependencies. The vanishing or exploding gradient problem is a well known issue associated with these challenges. One approach to addressing vanishing and exploding gradients is to use either soft or hard constraints on weight matrices so as to encourage or enforce orthogonality. Orthogonal matrices preserve gradient norm during backpropagation and may therefore be a desirable property. This paper explores issues with optimization convergence, speed and gradient stability when encouraging or enforcing orthogonality. To perform this analysis, we propose a weight matrix factorization and parameterization strategy through which we can bound matrix norms and therein control the degree of expansivity induced during backpropagation. We find that hard constraints on orthogonality can negatively affect the speed of convergence and model performance.
Learning Normalized Inputs for Iterative Estimation in Medical Image Segmentation
Feb 16, 2017
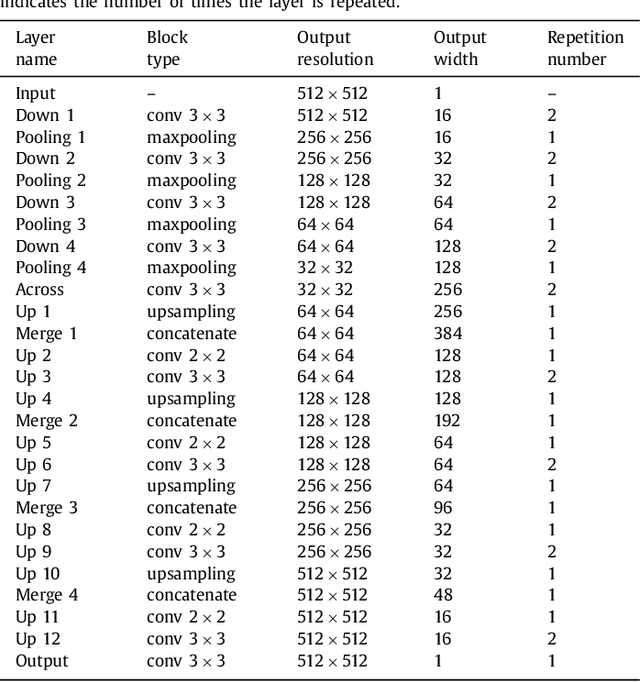
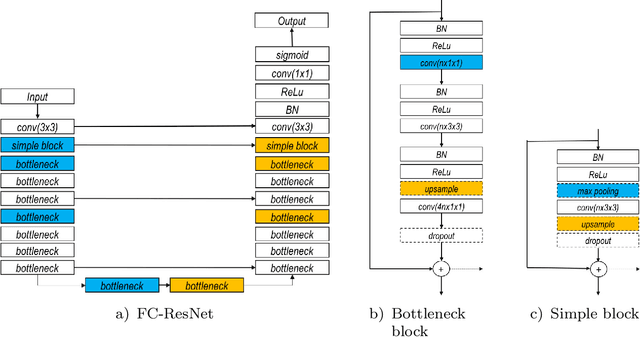
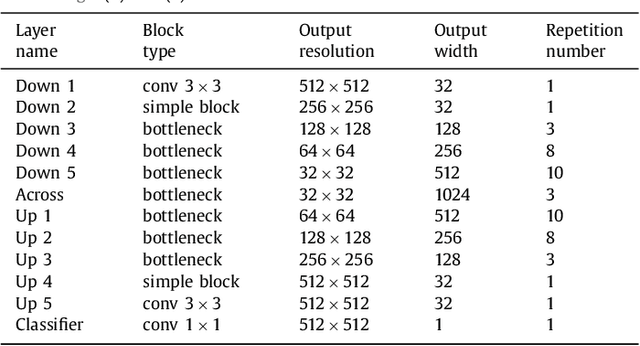
Abstract:In this paper, we introduce a simple, yet powerful pipeline for medical image segmentation that combines Fully Convolutional Networks (FCNs) with Fully Convolutional Residual Networks (FC-ResNets). We propose and examine a design that takes particular advantage of recent advances in the understanding of both Convolutional Neural Networks as well as ResNets. Our approach focuses upon the importance of a trainable pre-processing when using FC-ResNets and we show that a low-capacity FCN model can serve as a pre-processor to normalize medical input data. In our image segmentation pipeline, we use FCNs to obtain normalized images, which are then iteratively refined by means of a FC-ResNet to generate a segmentation prediction. As in other fully convolutional approaches, our pipeline can be used off-the-shelf on different image modalities. We show that using this pipeline, we exhibit state-of-the-art performance on the challenging Electron Microscopy benchmark, when compared to other 2D methods. We improve segmentation results on CT images of liver lesions, when contrasting with standard FCN methods. Moreover, when applying our 2D pipeline on a challenging 3D MRI prostate segmentation challenge we reach results that are competitive even when compared to 3D methods. The obtained results illustrate the strong potential and versatility of the pipeline by achieving highly accurate results on multi-modality images from different anatomical regions and organs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge