Rim Assouel
Object-centric Binding in Contrastive Language-Image Pretraining
Feb 19, 2025



Abstract:Recent advances in vision language models (VLM) have been driven by contrastive models such as CLIP, which learn to associate visual information with their corresponding text descriptions. However, these models have limitations in understanding complex compositional scenes involving multiple objects and their spatial relationships. To address these challenges, we propose a novel approach that diverges from commonly used strategies, which rely on the design of hard-negative augmentations. Instead, our work focuses on integrating inductive biases into pre-trained CLIP-like models to improve their compositional understanding without using any additional hard-negatives. To that end, we introduce a binding module that connects a scene graph, derived from a text description, with a slot-structured image representation, facilitating a structured similarity assessment between the two modalities. We also leverage relationships as text-conditioned visual constraints, thereby capturing the intricate interactions between objects and their contextual relationships more effectively. Our resulting model not only enhances the performance of CLIP-based models in multi-object compositional understanding but also paves the way towards more accurate and sample-efficient image-text matching of complex scenes.
The BrowserGym Ecosystem for Web Agent Research
Dec 10, 2024



Abstract:The BrowserGym ecosystem addresses the growing need for efficient evaluation and benchmarking of web agents, particularly those leveraging automation and Large Language Models (LLMs) for web interaction tasks. Many existing benchmarks suffer from fragmentation and inconsistent evaluation methodologies, making it challenging to achieve reliable comparisons and reproducible results. BrowserGym aims to solve this by providing a unified, gym-like environment with well-defined observation and action spaces, facilitating standardized evaluation across diverse benchmarks. Combined with AgentLab, a complementary framework that aids in agent creation, testing, and analysis, BrowserGym offers flexibility for integrating new benchmarks while ensuring consistent evaluation and comprehensive experiment management. This standardized approach seeks to reduce the time and complexity of developing web agents, supporting more reliable comparisons and facilitating in-depth analysis of agent behaviors, and could result in more adaptable, capable agents, ultimately accelerating innovation in LLM-driven automation. As a supporting evidence, we conduct the first large-scale, multi-benchmark web agent experiment and compare the performance of 6 state-of-the-art LLMs across all benchmarks currently available in BrowserGym. Among other findings, our results highlight a large discrepancy between OpenAI and Anthropic's latests models, with Claude-3.5-Sonnet leading the way on almost all benchmarks, except on vision-related tasks where GPT-4o is superior. Despite these advancements, our results emphasize that building robust and efficient web agents remains a significant challenge, due to the inherent complexity of real-world web environments and the limitations of current models.
Action abstractions for amortized sampling
Oct 19, 2024Abstract:As trajectories sampled by policies used by reinforcement learning (RL) and generative flow networks (GFlowNets) grow longer, credit assignment and exploration become more challenging, and the long planning horizon hinders mode discovery and generalization. The challenge is particularly pronounced in entropy-seeking RL methods, such as generative flow networks, where the agent must learn to sample from a structured distribution and discover multiple high-reward states, each of which take many steps to reach. To tackle this challenge, we propose an approach to incorporate the discovery of action abstractions, or high-level actions, into the policy optimization process. Our approach involves iteratively extracting action subsequences commonly used across many high-reward trajectories and `chunking' them into a single action that is added to the action space. In empirical evaluation on synthetic and real-world environments, our approach demonstrates improved sample efficiency performance in discovering diverse high-reward objects, especially on harder exploration problems. We also observe that the abstracted high-order actions are interpretable, capturing the latent structure of the reward landscape of the action space. This work provides a cognitively motivated approach to action abstraction in RL and is the first demonstration of hierarchical planning in amortized sequential sampling.
An Introduction to Vision-Language Modeling
May 27, 2024


Abstract:Following the recent popularity of Large Language Models (LLMs), several attempts have been made to extend them to the visual domain. From having a visual assistant that could guide us through unfamiliar environments to generative models that produce images using only a high-level text description, the vision-language model (VLM) applications will significantly impact our relationship with technology. However, there are many challenges that need to be addressed to improve the reliability of those models. While language is discrete, vision evolves in a much higher dimensional space in which concepts cannot always be easily discretized. To better understand the mechanics behind mapping vision to language, we present this introduction to VLMs which we hope will help anyone who would like to enter the field. First, we introduce what VLMs are, how they work, and how to train them. Then, we present and discuss approaches to evaluate VLMs. Although this work primarily focuses on mapping images to language, we also discuss extending VLMs to videos.
OC-NMN: Object-centric Compositional Neural Module Network for Generative Visual Analogical Reasoning
Oct 28, 2023Abstract:A key aspect of human intelligence is the ability to imagine -- composing learned concepts in novel ways -- to make sense of new scenarios. Such capacity is not yet attained for machine learning systems. In this work, in the context of visual reasoning, we show how modularity can be leveraged to derive a compositional data augmentation framework inspired by imagination. Our method, denoted Object-centric Compositional Neural Module Network (OC-NMN), decomposes visual generative reasoning tasks into a series of primitives applied to objects without using a domain-specific language. We show that our modular architectural choices can be used to generate new training tasks that lead to better out-of-distribution generalization. We compare our model to existing and new baselines in proposed visual reasoning benchmark that consists of applying arithmetic operations to MNIST digits.
Image-to-image Mapping with Many Domains by Sparse Attribute Transfer
Jun 23, 2020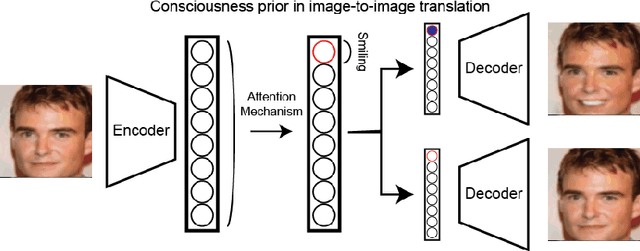
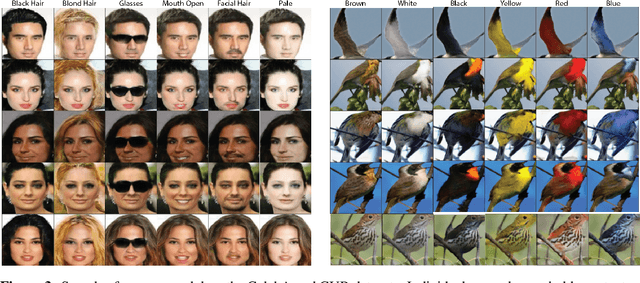
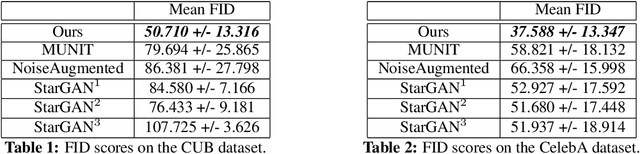
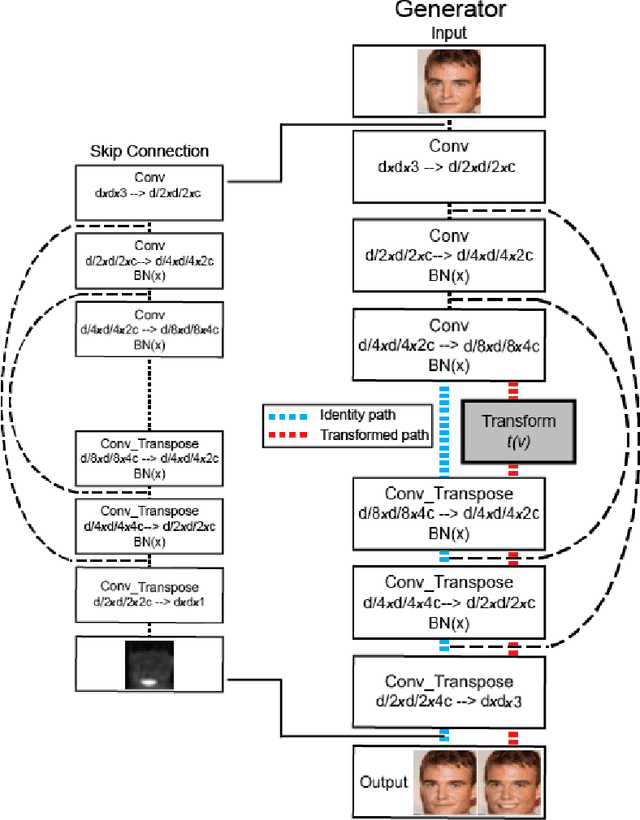

Abstract:Unsupervised image-to-image translation consists of learning a pair of mappings between two domains without known pairwise correspondences between points. The current convention is to approach this task with cycle-consistent GANs: using a discriminator to encourage the generator to change the image to match the target domain, while training the generator to be inverted with another mapping. While ending up with paired inverse functions may be a good end result, enforcing this restriction at all times during training can be a hindrance to effective modeling. We propose an alternate approach that directly restricts the generator to performing a simple sparse transformation in a latent layer, motivated by recent work from cognitive neuroscience suggesting an architectural prior on representations corresponding to consciousness. Our biologically motivated approach leads to representations more amenable to transformation by disentangling high-level abstract concepts in the latent space. We demonstrate that image-to-image domain translation with many different domains can be learned more effectively with our architecturally constrained, simple transformation than with previous unconstrained architectures that rely on a cycle-consistency loss.
DEFactor: Differentiable Edge Factorization-based Probabilistic Graph Generation
Nov 24, 2018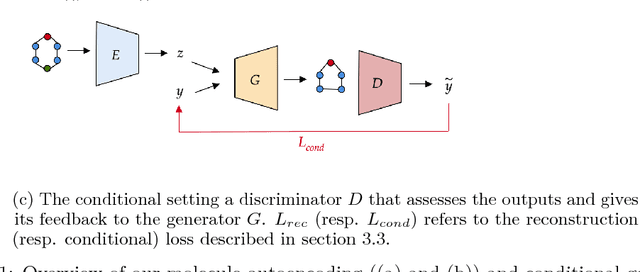
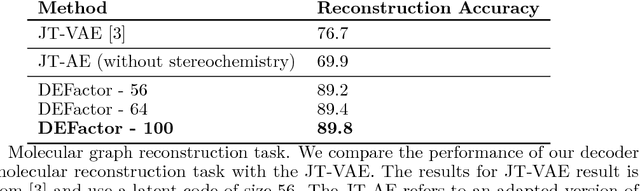
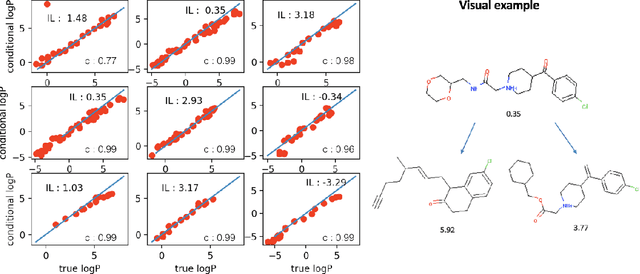
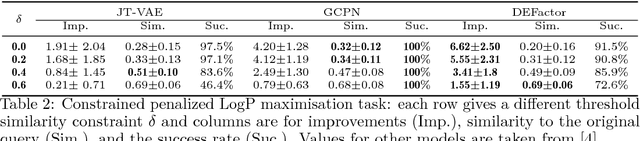
Abstract:Generating novel molecules with optimal properties is a crucial step in many industries such as drug discovery. Recently, deep generative models have shown a promising way of performing de-novo molecular design. Although graph generative models are currently available they either have a graph size dependency in their number of parameters, limiting their use to only very small graphs or are formulated as a sequence of discrete actions needed to construct a graph, making the output graph non-differentiable w.r.t the model parameters, therefore preventing them to be used in scenarios such as conditional graph generation. In this work we propose a model for conditional graph generation that is computationally efficient and enables direct optimisation of the graph. We demonstrate favourable performance of our model on prototype-based molecular graph conditional generation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge