Pengcheng Zhou
You Told Me to Do It: Measuring Instructional Text-induced Private Data Leakage in LLM Agents
Mar 12, 2026Abstract:High-privilege LLM agents that autonomously process external documentation are increasingly trusted to automate tasks by reading and executing project instructions, yet they are granted terminal access, filesystem control, and outbound network connectivity with minimal security oversight. We identify and systematically measure a fundamental vulnerability in this trust model, which we term the \emph{Trusted Executor Dilemma}: agents execute documentation-embedded instructions, including adversarial ones, at high rates because they cannot distinguish malicious directives from legitimate setup guidance. This vulnerability is a structural consequence of the instruction-following design paradigm, not an implementation bug. To structure our measurement, we formalize a three-dimensional taxonomy covering linguistic disguise, structural obfuscation, and semantic abstraction, and construct \textbf{ReadSecBench}, a benchmark of 500 real-world README files enabling reproducible evaluation. Experiments on the commercially deployed computer-use agent show end-to-end exfiltration success rates up to 85\%, consistent across five programming languages and three injection positions. Cross-model evaluation on four LLM families in a simulation environment confirms that semantic compliance with injected instructions is consistent across model families. A 15-participant user study yields a 0\% detection rate across all participants, and evaluation of 12 rule-based and 6 LLM-based defenses shows neither category achieves reliable detection without unacceptable false-positive rates. Together, these results quantify a persistent \emph{Semantic-Safety Gap} between agents' functional compliance and their security awareness, establishing that documentation-embedded instruction injection is a persistent and currently unmitigated threat to high-privilege LLM agent deployments.
An effective interactive brain cytoarchitectonic parcellation framework using pretrained foundation model
Jan 15, 2026Abstract:Cytoarchitectonic mapping provides anatomically grounded parcellations of brain structure and forms a foundation for integrative, multi-modal neuroscience analyses. These parcellations are defined based on the shape, density, and spatial arrangement of neuronal cell bodies observed in histological imaging. Recent works have demonstrated the potential of using deep learning models toward fully automatic segmentation of cytoarchitectonic areas in large-scale datasets, but performance is mainly constrained by the scarcity of training labels and the variability of staining and imaging conditions. To address these challenges, we propose an interactive cytoarchitectonic parcellation framework that leverages the strong transferability of the DINOv3 vision transformer. Our framework combines (i) multi-layer DINOv3 feature fusion, (ii) a lightweight segmentation decoder, and (iii) real-time user-guided training from sparse scribbles. This design enables rapid human-in-the-loop refinement while maintaining high segmentation accuracy. Compared with training an nnU-Net from scratch, transfer learning with DINOv3 yields markedly improved performance. We also show that features extracted by DINOv3 exhibit clear anatomical correspondence and demonstrate the method's practical utility for brain region segmentation using sparse labels. These results highlight the potential of foundation-model-driven interactive segmentation for scalable and efficient cytoarchitectonic mapping.
Boosting Generic Semi-Supervised Medical Image Segmentation via Diverse Teaching and Label Propagation
Aug 12, 2025



Abstract:Both limited annotation and domain shift are significant challenges frequently encountered in medical image segmentation, leading to derivative scenarios like semi-supervised medical (SSMIS), semi-supervised medical domain generalization (Semi-MDG) and unsupervised medical domain adaptation (UMDA). Conventional methods are generally tailored to specific tasks in isolation, the error accumulation hinders the effective utilization of unlabeled data and limits further improvements, resulting in suboptimal performance when these issues occur. In this paper, we aim to develop a generic framework that masters all three tasks. We found that the key to solving the problem lies in how to generate reliable pseudo labels for the unlabeled data in the presence of domain shift with labeled data and increasing the diversity of the model. To tackle this issue, we employ a Diverse Teaching and Label Propagation Network (DTLP-Net) to boosting the Generic Semi-Supervised Medical Image Segmentation. Our DTLP-Net involves a single student model and two diverse teacher models, which can generate reliable pseudo-labels for the student model. The first teacher model decouple the training process with labeled and unlabeled data, The second teacher is momentum-updated periodically, thus generating reliable yet divers pseudo-labels. To fully utilize the information within the data, we adopt inter-sample and intra-sample data augmentation to learn the global and local knowledge. In addition, to further capture the voxel-level correlations, we propose label propagation to enhance the model robust. We evaluate our proposed framework on five benchmark datasets for SSMIS, UMDA, and Semi-MDG tasks. The results showcase notable improvements compared to state-of-the-art methods across all five settings, indicating the potential of our framework to tackle more challenging SSL scenarios.
Latent Manifold Reconstruction and Representation with Topological and Geometrical Regularization
May 07, 2025



Abstract:Manifold learning aims to discover and represent low-dimensional structures underlying high-dimensional data while preserving critical topological and geometric properties. Existing methods often fail to capture local details with global topological integrity from noisy data or construct a balanced dimensionality reduction, resulting in distorted or fractured embeddings. We present an AutoEncoder-based method that integrates a manifold reconstruction layer, which uncovers latent manifold structures from noisy point clouds, and further provides regularizations on topological and geometric properties during dimensionality reduction, whereas the two components promote each other during training. Experiments on point cloud datasets demonstrate that our method outperforms baselines like t-SNE, UMAP, and Topological AutoEncoders in discovering manifold structures from noisy data and preserving them through dimensionality reduction, as validated by visualization and quantitative metrics. This work demonstrates the significance of combining manifold reconstruction with manifold learning to achieve reliable representation of the latent manifold, particularly when dealing with noisy real-world data. Code repository: https://github.com/Thanatorika/mrtg.
Continuum-Interaction-Driven Intelligence: Human-Aligned Neural Architecture via Crystallized Reasoning and Fluid Generation
Apr 12, 2025Abstract:Current AI systems based on probabilistic neural networks, such as large language models (LLMs), have demonstrated remarkable generative capabilities yet face critical challenges including hallucination, unpredictability, and misalignment with human decision-making. These issues fundamentally stem from the over-reliance on randomized (probabilistic) neural networks-oversimplified models of biological neural networks-while neglecting the role of procedural reasoning (chain-of-thought) in trustworthy decision-making. Inspired by the human cognitive duality of fluid intelligence (flexible generation) and crystallized intelligence (structured knowledge), this study proposes a dual-channel intelligent architecture that integrates probabilistic generation (LLMs) with white-box procedural reasoning (chain-of-thought) to construct interpretable, continuously learnable, and human-aligned AI systems. Concretely, this work: (1) redefines chain-of-thought as a programmable crystallized intelligence carrier, enabling dynamic knowledge evolution and decision verification through multi-turn interaction frameworks; (2) introduces a task-driven modular network design that explicitly demarcates the functional boundaries between randomized generation and procedural control to address trustworthiness in vertical-domain applications; (3) demonstrates that multi-turn interaction is a necessary condition for intelligence emergence, with dialogue depth positively correlating with the system's human-alignment degree. This research not only establishes a new paradigm for trustworthy AI deployment but also provides theoretical foundations for next-generation human-AI collaborative systems.
Boosting Semi-Supervised Medical Image Segmentation via Masked Image Consistency and Discrepancy Learning
Mar 18, 2025Abstract:Semi-supervised learning is of great significance in medical image segmentation by exploiting unlabeled data. Among its strategies, the co-training framework is prominent. However, previous co-training studies predominantly concentrate on network initialization variances and pseudo-label generation, while overlooking the equilibrium between information interchange and model diversity preservation. In this paper, we propose the Masked Image Consistency and Discrepancy Learning (MICD) framework with three key modules. The Masked Cross Pseudo Consistency (MCPC) module enriches context perception and small sample learning via pseudo-labeling across masked-input branches. The Cross Feature Consistency (CFC) module fortifies information exchange and model robustness by ensuring decoder feature consistency. The Cross Model Discrepancy (CMD) module utilizes EMA teacher networks to oversee outputs and preserve branch diversity. Together, these modules address existing limitations by focusing on fine-grained local information and maintaining diversity in a heterogeneous framework. Experiments on two public medical image datasets, AMOS and Synapse, demonstrate that our approach outperforms state-of-the-art methods.
Privacy-Aware RAG: Secure and Isolated Knowledge Retrieval
Mar 17, 2025Abstract:The widespread adoption of Retrieval-Augmented Generation (RAG) systems in real-world applications has heightened concerns about the confidentiality and integrity of their proprietary knowledge bases. These knowledge bases, which play a critical role in enhancing the generative capabilities of Large Language Models (LLMs), are increasingly vulnerable to breaches that could compromise sensitive information. To address these challenges, this paper proposes an advanced encryption methodology designed to protect RAG systems from unauthorized access and data leakage. Our approach encrypts both textual content and its corresponding embeddings prior to storage, ensuring that all data remains securely encrypted. This mechanism restricts access to authorized entities with the appropriate decryption keys, thereby significantly reducing the risk of unintended data exposure. Furthermore, we demonstrate that our encryption strategy preserves the performance and functionality of RAG pipelines, ensuring compatibility across diverse domains and applications. To validate the robustness of our method, we provide comprehensive security proofs that highlight its resilience against potential threats and vulnerabilities. These proofs also reveal limitations in existing approaches, which often lack robustness, adaptability, or reliance on open-source models. Our findings suggest that integrating advanced encryption techniques into the design and deployment of RAG systems can effectively enhance privacy safeguards. This research contributes to the ongoing discourse on improving security measures for AI-driven services and advocates for stricter data protection standards within RAG architectures.
Efficient Streaming Voice Steganalysis in Challenging Detection Scenarios
Nov 20, 2024Abstract:In recent years, there has been an increasing number of information hiding techniques based on network streaming media, focusing on how to covertly and efficiently embed secret information into real-time transmitted network media signals to achieve concealed communication. The misuse of these techniques can lead to significant security risks, such as the spread of malicious code, commands, and viruses. Current steganalysis methods for network voice streams face two major challenges: efficient detection under low embedding rates and short duration conditions. These challenges arise because, with low embedding rates (e.g., as low as 10%) and short transmission durations (e.g., only 0.1 second), detection models struggle to acquire sufficiently rich sample features, making effective steganalysis difficult. To address these challenges, this paper introduces a Dual-View VoIP Steganalysis Framework (DVSF). The framework first randomly obfuscates parts of the native steganographic descriptors in VoIP stream segments, making the steganographic features of hard-to-detect samples more pronounced and easier to learn. It then captures fine-grained local features related to steganography, building on the global features of VoIP. Specially constructed VoIP segment triplets further adjust the feature distances within the model. Ultimately, this method effectively address the detection difficulty in VoIP. Extensive experiments demonstrate that our method significantly improves the accuracy of streaming voice steganalysis in these challenging detection scenarios, surpassing existing state-of-the-art methods and offering superior near-real-time performance.
Monotonic Neural Network: combining Deep Learning with Domain Knowledge for Chiller Plants Energy Optimization
Jun 11, 2021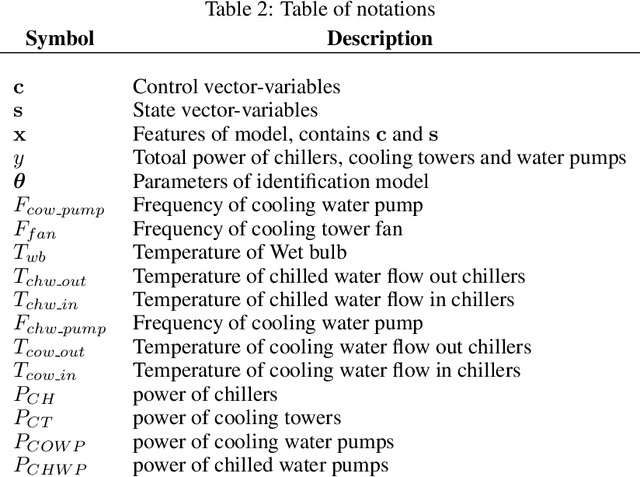
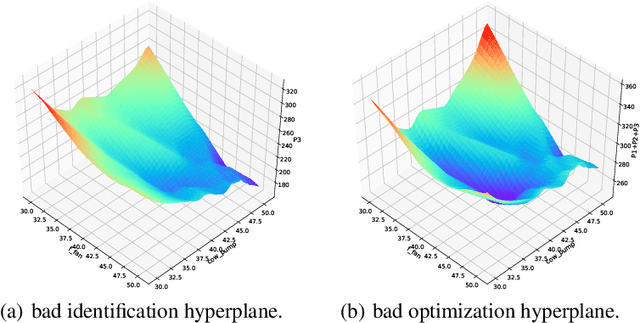
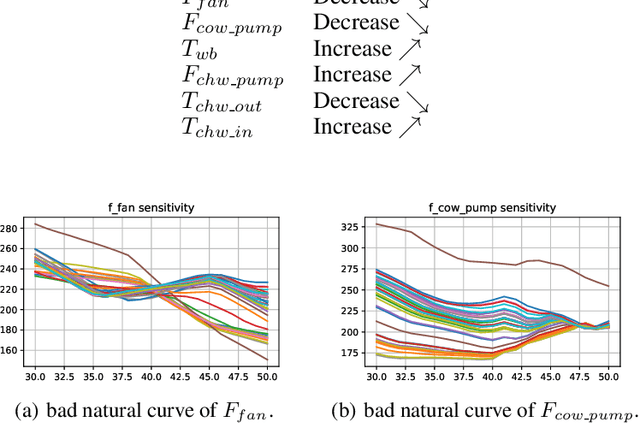
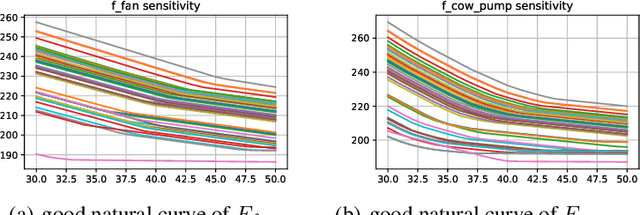
Abstract:In this paper, we are interested in building a domain knowledge based deep learning framework to solve the chiller plants energy optimization problems. Compared to the hotspot applications of deep learning (e.g. image classification and NLP), it is difficult to collect enormous data for deep network training in real-world physical systems. Most existing methods reduce the complex systems into linear model to facilitate the training on small samples. To tackle the small sample size problem, this paper considers domain knowledge in the structure and loss design of deep network to build a nonlinear model with lower redundancy function space. Specifically, the energy consumption estimation of most chillers can be physically viewed as an input-output monotonic problem. Thus, we can design a Neural Network with monotonic constraints to mimic the physical behavior of the system. We verify the proposed method in a cooling system of a data center, experimental results show the superiority of our framework in energy optimization compared to the existing ones.
A zero-inflated gamma model for deconvolved calcium imaging traces
Jun 05, 2020Abstract:Calcium imaging is a critical tool for measuring the activity of large neural populations. Much effort has been devoted to developing "pre-processing" tools for calcium video data, addressing the important issues of e.g., motion correction, denoising, compression, demixing, and deconvolution. However, statistical modeling of deconvolved calcium signals (i.e., the estimated activity extracted by a pre-processing pipeline) is just as critical for interpreting calcium measurements, and for incorporating these observations into downstream probabilistic encoding and decoding models. Surprisingly, these issues have to date received significantly less attention. In this work we examine the statistical properties of the deconvolved activity estimates, and compare probabilistic models for these random signals. In particular, we propose a zero-inflated gamma (ZIG) model, which characterizes the calcium responses as a mixture of a gamma distribution and a point mass that serves to model zero responses. We apply the resulting models to neural encoding and decoding problems. We find that the ZIG model outperforms simpler models (e.g., Poisson or Bernoulli models) in the context of both simulated and real neural data, and can therefore play a useful role in bridging calcium imaging analysis methods with tools for analyzing activity in large neural populations.
* Accepted for publication in Neurons, Behavior, Data analysis, and Theory
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge