Peiyuan Li
SPAMoE: Spectrum-Aware Hybrid Operator Framework for Full-Waveform Inversion
Apr 08, 2026Abstract:Full-waveform inversion (FWI) is pivotal for reconstructing high-resolution subsurface velocity models but remains computationally intensive and ill-posed. While deep learning approaches promise efficiency, existing Convolutional Neural Networks (CNNs) and single-paradigm Neural Operators (NOs) struggle with one fundamental issue: frequency entanglement of multi-scale geological features. To address this challenge, we propose Spectral-Preserving Adaptive MoE (SPAMoE), a novel spectrum-aware framework for solving inverse problems with complex multi-scale structures. Our approach introduces a Spectral-Preserving DINO Encoder that enforces a lower bound on the high-to-low frequency energy ratio of the encoded representation, mitigating high-frequency collapse and stabilizing subsequent frequency-domain modeling. Furthermore, we design a novel Spectral Decomposition and Routing mechanism that dynamically assigns frequency bands to a Mixture-of-Experts (MoE) ensemble comprising FNO, MNO, and LNO. On the ten OpenFWI sub-datasets, experiments show that SPAMoE reduces the average MAE by 54.1% relative to the best officially reported OpenFWI baseline, thereby establishing a new architectural framework for learning-based full-waveform inversion.
VisualAD: Language-Free Zero-Shot Anomaly Detection via Vision Transformer
Mar 09, 2026Abstract:Zero-shot anomaly detection (ZSAD) requires detecting and localizing anomalies without access to target-class anomaly samples. Mainstream methods rely on vision-language models (VLMs) such as CLIP: they build hand-crafted or learned prompt sets for normal and abnormal semantics, then compute image-text similarities for open-set discrimination. While effective, this paradigm depends on a text encoder and cross-modal alignment, which can lead to training instability and parameter redundancy. This work revisits the necessity of the text branch in ZSAD and presents VisualAD, a purely visual framework built on Vision Transformers. We introduce two learnable tokens within a frozen backbone to directly encode normality and abnormality. Through multi-layer self-attention, these tokens interact with patch tokens, gradually acquiring high-level notions of normality and anomaly while guiding patches to highlight anomaly-related cues. Additionally, we incorporate a Spatial-Aware Cross-Attention (SCA) module and a lightweight Self-Alignment Function (SAF): SCA injects fine-grained spatial information into the tokens, and SAF recalibrates patch features before anomaly scoring. VisualAD achieves state-of-the-art performance on 13 zero-shot anomaly detection benchmarks spanning industrial and medical domains, and adapts seamlessly to pretrained vision backbones such as the CLIP image encoder and DINOv2. Code: https://github.com/7HHHHH/VisualAD
Unified Multi-Domain Graph Pre-training for Homogeneous and Heterogeneous Graphs via Domain-Specific Expert Encoding
Feb 13, 2026Abstract:Graph pre-training has achieved remarkable success in recent years, delivering transferable representations for downstream adaptation. However, most existing methods are designed for either homogeneous or heterogeneous graphs, thereby hindering unified graph modeling across diverse graph types. This separation contradicts real-world applications, where mixed homogeneous and heterogeneous graphs are ubiquitous, and distribution shifts between upstream pre-training and downstream deployment are common. In this paper, we empirically demonstrate that a balanced mixture of homogeneous and heterogeneous graph pre-training benefits downstream tasks and propose a unified multi-domain \textbf{G}raph \textbf{P}re-training method across \textbf{H}omogeneous and \textbf{H}eterogeneous graphs ($\mathbf{GPH^{2}}$). To address the lack of a unified encoder for homogeneous and heterogeneous graphs, we propose a Unified Multi-View Graph Construction that simultaneously encodes both without explicit graph-type-specific designs. To cope with the increased cross-domain distribution discrepancies arising from mixed graphs, we introduce domain-specific expert encoding. Each expert is independently pre-trained on a single graph to capture domain-specific knowledge, thereby shielding the pre-training encoder from the adverse effects of cross-domain discrepancies. For downstream tasks, we further design a Task-oriented Expert Fusion Strategy that adaptively integrates multiple experts based on their discriminative strengths. Extensive experiments on mixed graphs demonstrate that $\text{GPH}^{2}$ enables stable transfer across graph types and domains, significantly outperforming existing graph pre-training methods.
AugLiChem: Data Augmentation Library of Chemical Structures for Machine Learning
Dec 01, 2021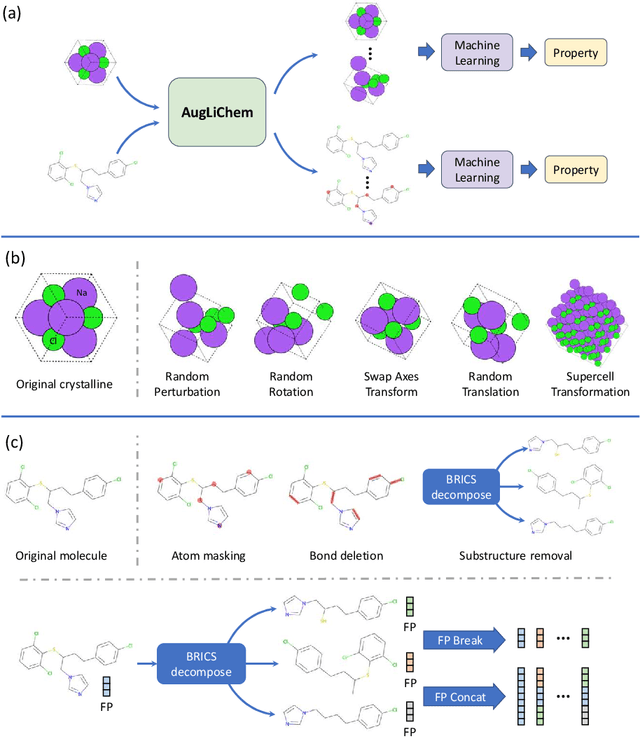
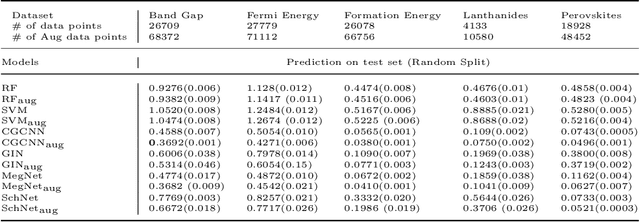
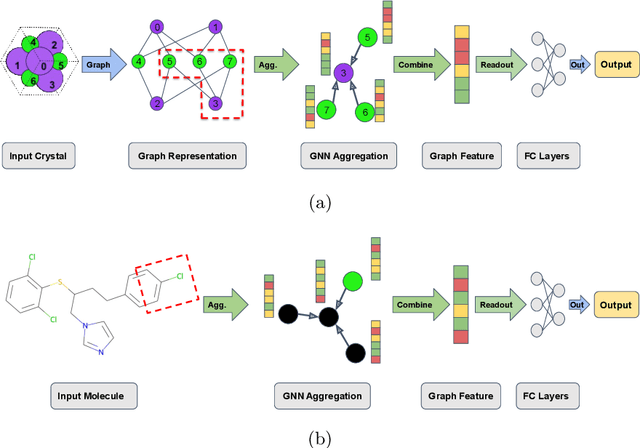
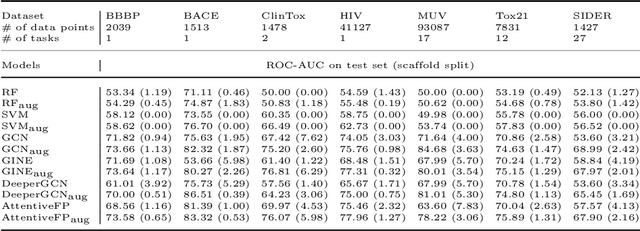
Abstract:Machine learning (ML) has demonstrated the promise for accurate and efficient property prediction of molecules and crystalline materials. To develop highly accurate ML models for chemical structure property prediction, datasets with sufficient samples are required. However, obtaining clean and sufficient data of chemical properties can be expensive and time-consuming, which greatly limits the performance of ML models. Inspired by the success of data augmentations in computer vision and natural language processing, we developed AugLiChem: the data augmentation library for chemical structures. Augmentation methods for both crystalline systems and molecules are introduced, which can be utilized for fingerprint-based ML models and Graph Neural Networks(GNNs). We show that using our augmentation strategies significantly improves the performance of ML models, especially when using GNNs. In addition, the augmentations that we developed can be used as a direct plug-in module during training and have demonstrated the effectiveness when implemented with different GNN models through the AugliChem library. The Python-based package for our implementation of Auglichem: Data augmentation library for chemical structures, is publicly available at: https://github.com/BaratiLab/AugLiChem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge