Marleen de Bruijne
for the ALFA study
Semi-Supervised Medical Image Segmentation via Learning Consistency under Transformations
Nov 04, 2019
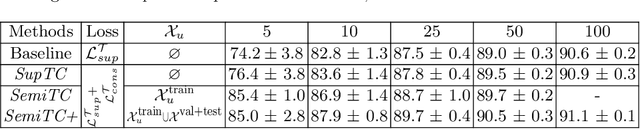
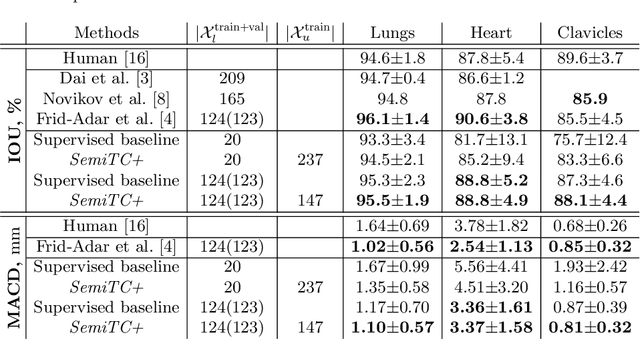
Abstract:The scarcity of labeled data often limits the application of supervised deep learning techniques for medical image segmentation. This has motivated the development of semi-supervised techniques that learn from a mixture of labeled and unlabeled images. In this paper, we propose a novel semi-supervised method that, in addition to supervised learning on labeled training images, learns to predict segmentations consistent under a given class of transformations on both labeled and unlabeled images. More specifically, in this work we explore learning equivariance to elastic deformations. We implement this through: 1) a Siamese architecture with two identical branches, each of which receives a differently transformed image, and 2) a composite loss function with a supervised segmentation loss term and an unsupervised term that encourages segmentation consistency between the predictions of the two branches. We evaluate the method on a public dataset of chest radiographs with segmentations of anatomical structures using 5-fold cross-validation. The proposed method reaches significantly higher segmentation accuracy compared to supervised learning. This is due to learning transformation consistency on both labeled and unlabeled images, with the latter contributing the most. We achieve the performance comparable to state-of-the-art chest X-ray segmentation methods while using substantially fewer labeled images.
APIR-Net: Autocalibrated Parallel Imaging Reconstruction using a Neural Network
Sep 19, 2019



Abstract:Deep learning has been successfully demonstrated in MRI reconstruction of accelerated acquisitions. However, its dependence on representative training data limits the application across different contrasts, anatomies, or image sizes. To address this limitation, we propose an unsupervised, auto-calibrated k-space completion method, based on a uniquely designed neural network that reconstructs the full k-space from an undersampled k-space, exploiting the redundancy among the multiple channels in the receive coil in a parallel imaging acquisition. To achieve this, contrary to common convolutional network approaches, the proposed network has a decreasing number of feature maps of constant size. In contrast to conventional parallel imaging methods such as GRAPPA that estimate the prediction kernel from the fully sampled autocalibration signals in a linear way, our method is able to learn nonlinear relations between sampled and unsampled positions in k-space. The proposed method was compared to the start-of-the-art ESPIRiT and RAKI methods in terms of noise amplification and visual image quality in both phantom and in-vivo experiments. The experiments indicate that APIR-Net provides a promising alternative to the conventional parallel imaging methods, and results in improved image quality especially for low SNR acquisitions.
A joint 3D UNet-Graph Neural Network-based method for Airway Segmentation from chest CTs
Aug 22, 2019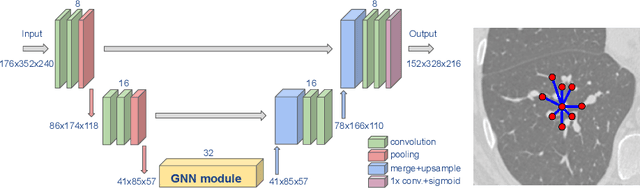
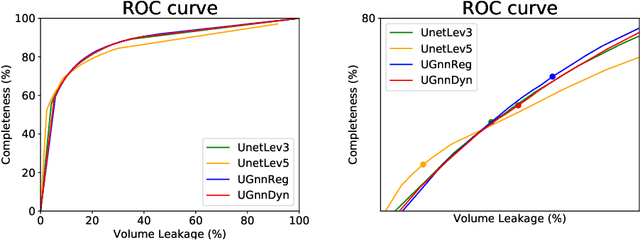
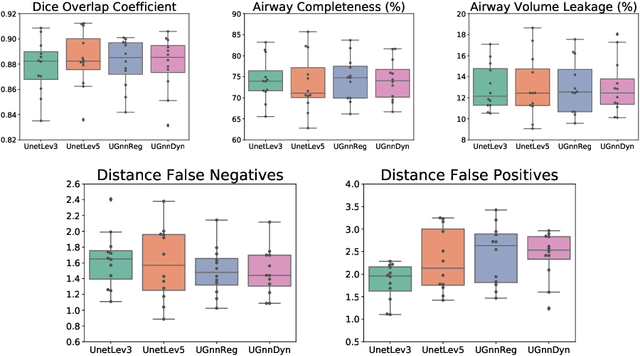
Abstract:We present an end-to-end deep learning segmentation method by combining a 3D UNet architecture with a graph neural network (GNN) model. In this approach, the convolutional layers at the deepest level of the UNet are replaced by a GNN-based module with a series of graph convolutions. The dense feature maps at this level are transformed into a graph input to the GNN module. The incorporation of graph convolutions in the UNet provides nodes in the graph with information that is based on node connectivity, in addition to the local features learnt through the downsampled paths. This information can help improve segmentation decisions. By stacking several graph convolution layers, the nodes can access higher order neighbourhood information without substantial increase in computational expense. We propose two types of node connectivity in the graph adjacency: i) one predefined and based on a regular node neighbourhood, and ii) one dynamically computed during training and using the nearest neighbour nodes in the feature space. We have applied this method to the task of segmenting the airway tree from chest CT scans. Experiments have been performed on 32 CTs from the Danish Lung Cancer Screening Trial dataset. We evaluate the performance of the UNet-GNN models with two types of graph adjacency and compare it with the baseline UNet.
Automated Lesion Detection by Regressing Intensity-Based Distance with a Neural Network
Jul 29, 2019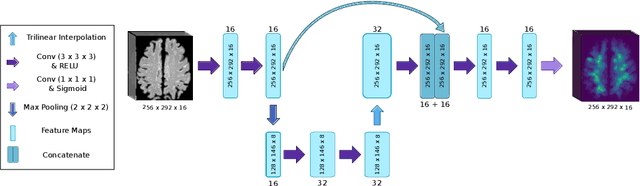
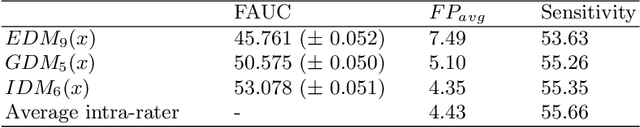

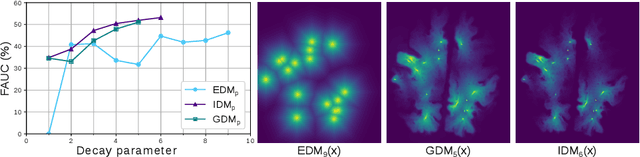
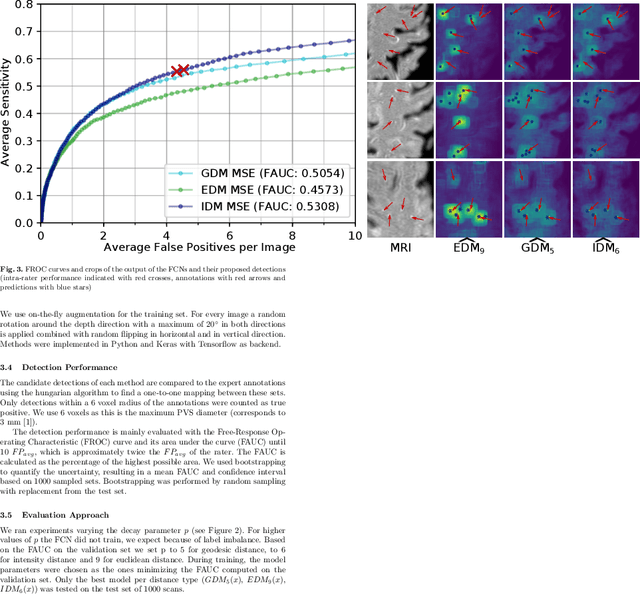
Abstract:Localization of focal vascular lesions on brain MRI is an important component of research on the etiology of neurological disorders. However, manual annotation of lesions can be challenging, time-consuming and subject to observer bias. Automated detection methods often need voxel-wise annotations for training. We propose a novel approach for automated lesion detection that can be trained on scans only annotated with a dot per lesion instead of a full segmentation. From the dot annotations and their corresponding intensity images we compute various distance maps (DMs), indicating the distance to a lesion based on spatial distance, intensity distance, or both. We train a fully convolutional neural network (FCN) to predict these DMs for unseen intensity images. The local optima in the predicted DMs are expected to correspond to lesion locations. We show the potential of this approach to detect enlarged perivascular spaces in white matter on a large brain MRI dataset with an independent test set of 1000 scans. Our method matches the intra-rater performance of the expert rater that was computed on an independent set. We compare the different types of distance maps, showing that incorporating intensity information in the distance maps used to train an FCN greatly improves performance.
Multi-Task Attention-Based Semi-Supervised Learning for Medical Image Segmentation
Jul 29, 2019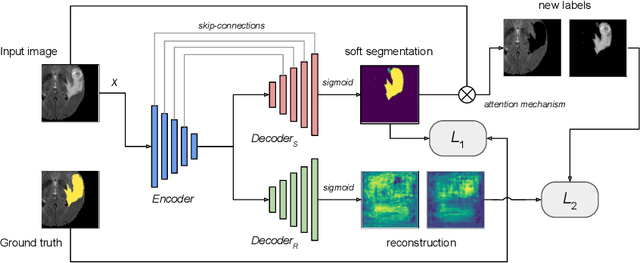
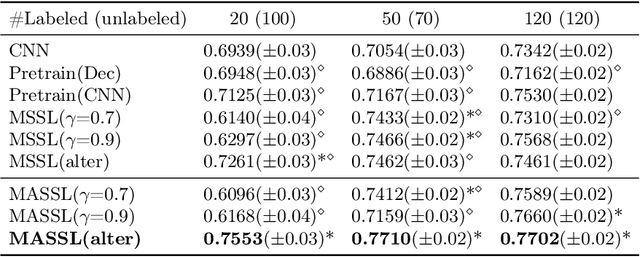
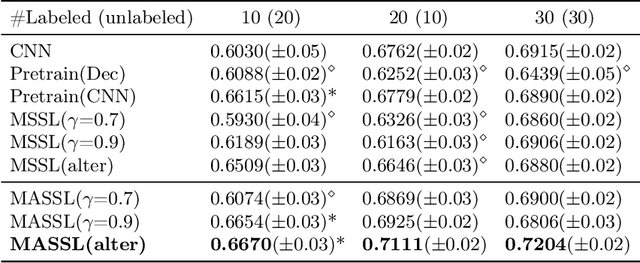
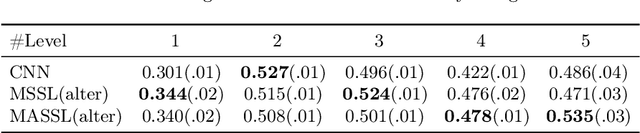
Abstract:We propose a novel semi-supervised image segmentation method that simultaneously optimizes a supervised segmentation and an unsupervised reconstruction objectives. The reconstruction objective uses an attention mechanism that separates the reconstruction of image areas corresponding to different classes. The proposed approach was evaluated on two applications: brain tumor and white matter hyperintensities segmentation. Our method, trained on unlabeled and a small number of labeled images, outperformed supervised CNNs trained with the same number of images and CNNs pre-trained on unlabeled data. In ablation experiments, we observed that the proposed attention mechanism substantially improves segmentation performance. We explore two multi-task training strategies: joint training and alternating training. Alternating training requires fewer hyperparameters and achieves a better, more stable performance than joint training. Finally, we analyze the features learned by different methods and find that the attention mechanism helps to learn more discriminative features in the deeper layers of encoders.
Automated Image Registration Quality Assessment Utilizing Deep-learning based Ventricle Extraction in Clinical Data
Jul 01, 2019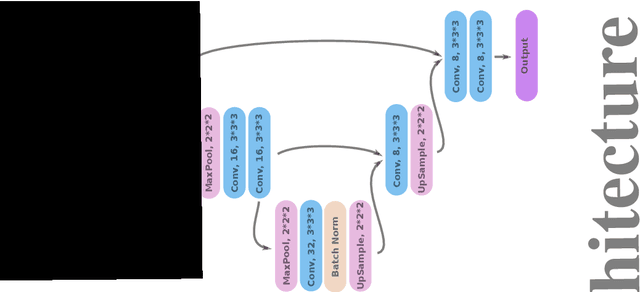
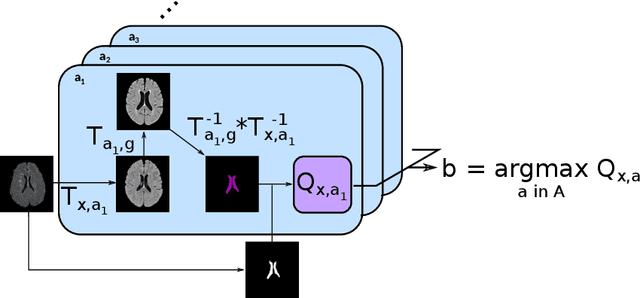
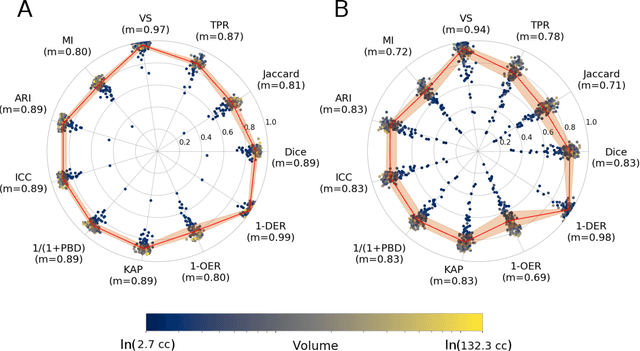
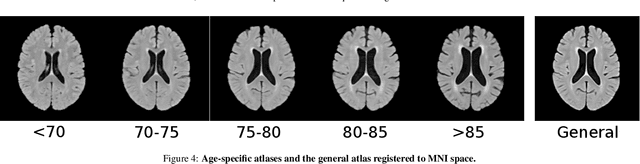
Abstract:Registration is a core component of many imaging pipelines. In case of clinical scans, with lower resolution and sometimes substantial motion artifacts, registration can produce poor results. Visual assessment of registration quality in large clinical datasets is inefficient. In this work, we propose to automatically assess the quality of registration to an atlas in clinical FLAIR MRI scans of the brain. The method consists of automatically segmenting the ventricles of a given scan using a neural network, and comparing the segmentation to the atlas' ventricles propagated to image space. We used the proposed method to improve clinical image registration to a general atlas by computing multiple registrations and then selecting the registration that yielded the highest ventricle overlap. Methods were evaluated in a single-site dataset of more than 1000 scans, as well as a multi-center dataset comprising 142 clinical scans from 12 sites. The automated ventricle segmentation reached a Dice coefficient with manual annotations of 0.89 in the single-site dataset, and 0.83 in the multi-center dataset. Registration via age-specific atlases could improve ventricle overlap compared to a direct registration to the general atlas (Dice similarity coefficient increase up to 0.15). Experiments also showed that selecting scans with the registration quality assessment method could improve the quality of average maps of white matter hyperintensity burden, instead of using all scans for the computation of the white matter hyperintensity map. In this work, we demonstrated the utility of an automated tool for assessing image registration quality in clinical scans. This image quality assessment step could ultimately assist in the translation of automated neuroimaging pipelines to the clinic.
Weakly Supervised Object Detection with 2D and 3D Regression Neural Networks
Jun 14, 2019
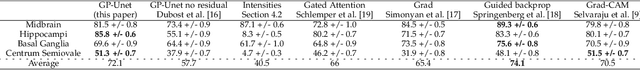
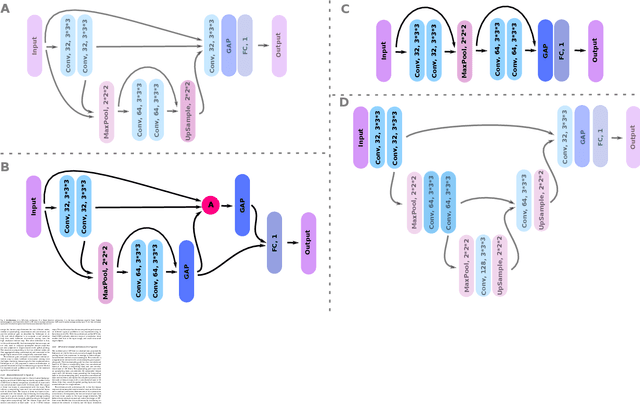
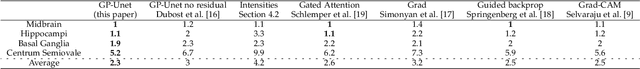
Abstract:Weakly supervised detection methods can infer the location of target objects in an image without requiring location or appearance information during training. We propose a weakly supervised deep learning method for the detection of objects that appear at multiple locations in an image. The method computes attention maps using the last feature maps of an encoder-decoder network optimized only with global labels: the number of occurrences of the target object in an image. In contrast with previous approaches, attention maps are generated at full input resolution thanks to the decoder part. The proposed approach is compared to multiple state-of-the-art methods in two tasks: the detection of digits in MNIST-based datasets, and the real life application of detection of enlarged perivascular spaces -- a type of brain lesion -- in four brain regions in a dataset of 2202 3D brain MRI scans. In MNIST-based datasets, the proposed method outperforms the other methods. In the brain dataset, several weakly supervised detection methods come close to the human intrarater agreement in each region. The proposed method reaches the lowest number of false positive detections in all brain regions at the operating point, while its average sensitivity is similar to that of the other best methods.
A cross-center smoothness prior for variational Bayesian brain tissue segmentation
Mar 11, 2019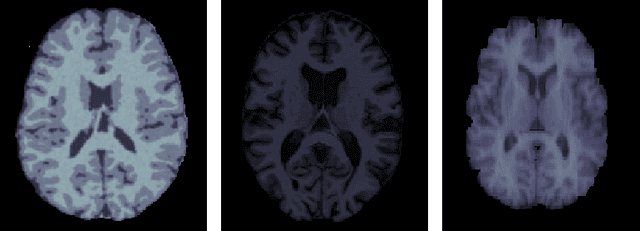
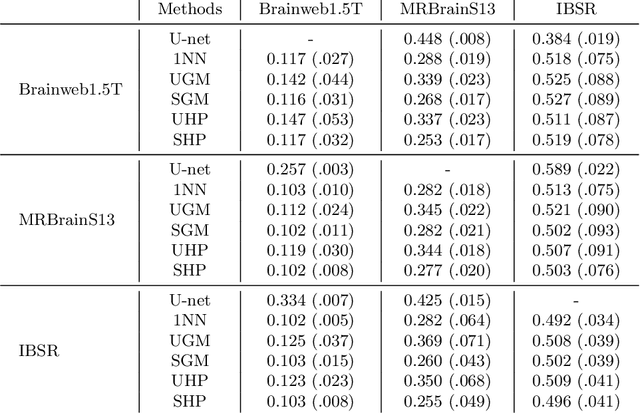
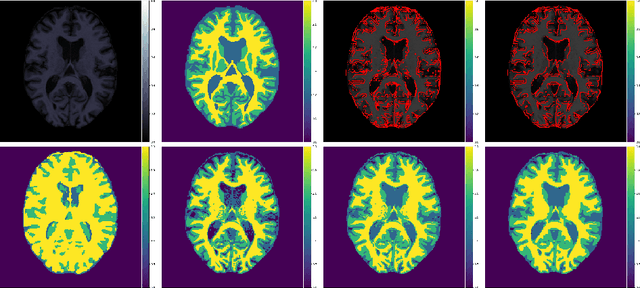
Abstract:Suppose one is faced with the challenge of tissue segmentation in MR images, without annotators at their center to provide labeled training data. One option is to go to another medical center for a trained classifier. Sadly, tissue classifiers do not generalize well across centers due to voxel intensity shifts caused by center-specific acquisition protocols. However, certain aspects of segmentations, such as spatial smoothness, remain relatively consistent and can be learned separately. Here we present a smoothness prior that is fit to segmentations produced at another medical center. This informative prior is presented to an unsupervised Bayesian model. The model clusters the voxel intensities, such that it produces segmentations that are similarly smooth to those of the other medical center. In addition, the unsupervised Bayesian model is extended to a semi-supervised variant, which needs no visual interpretation of clusters into tissues.
Event-Based Modeling with High-Dimensional Imaging Biomarkers for Estimating Spatial Progression of Dementia
Mar 08, 2019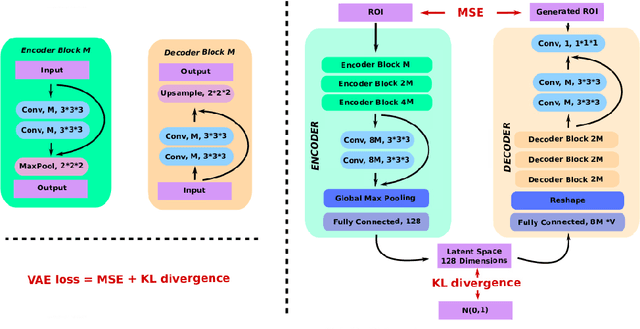
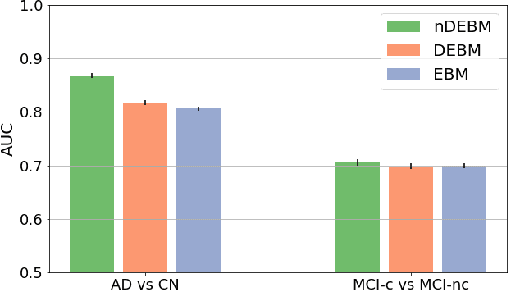
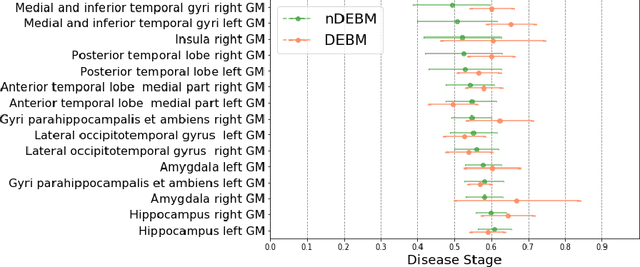
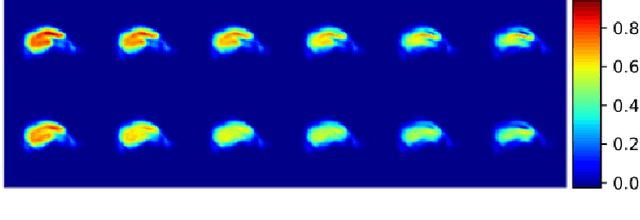
Abstract:Event-based models (EBM) are a class of disease progression models that can be used to estimate temporal ordering of neuropathological changes from cross-sectional data. Current EBMs only handle scalar biomarkers, such as regional volumes, as inputs. However, regional aggregates are a crude summary of the underlying high-resolution images, potentially limiting the accuracy of EBM. Therefore, we propose a novel method that exploits high-dimensional voxel-wise imaging biomarkers: n-dimensional discriminative EBM (nDEBM). nDEBM is based on an insight that mixture modeling, which is a key element of conventional EBMs, can be replaced by a more scalable semi-supervised support vector machine (SVM) approach. This SVM is used to estimate the degree of abnormality of each region which is then used to obtain subject-specific disease progression patterns. These patterns are in turn used for estimating the mean ordering by fitting a generalized Mallows model. In order to validate the biomarker ordering obtained using nDEBM, we also present a framework for Simulation of Imaging Biomarkers' Temporal Evolution (SImBioTE) that mimics neurodegeneration in brain regions. SImBioTE trains variational auto-encoders (VAE) in different brain regions independently to simulate images at varying stages of disease progression. We also validate nDEBM clinically using data from the Alzheimer's Disease Neuroimaging Initiative (ADNI). In both experiments, nDEBM using high-dimensional features gave better performance than state-of-the-art EBM methods using regional volume biomarkers. This suggests that nDEBM is a promising approach for disease progression modeling.
Graph Refinement based Tree Extraction using Mean-Field Networks and Graph Neural Networks
Nov 21, 2018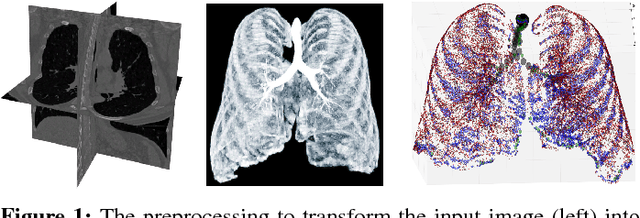
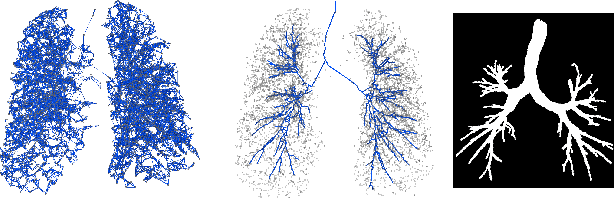
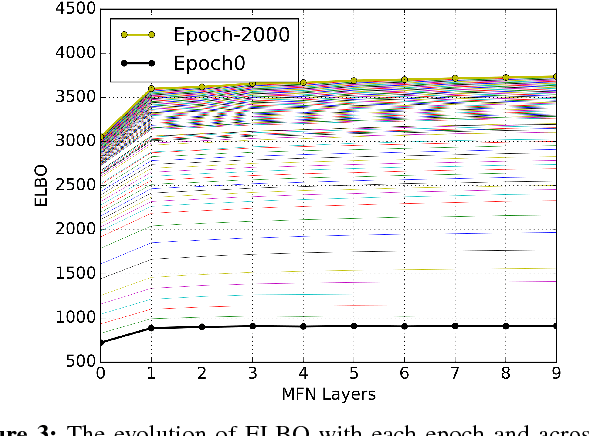
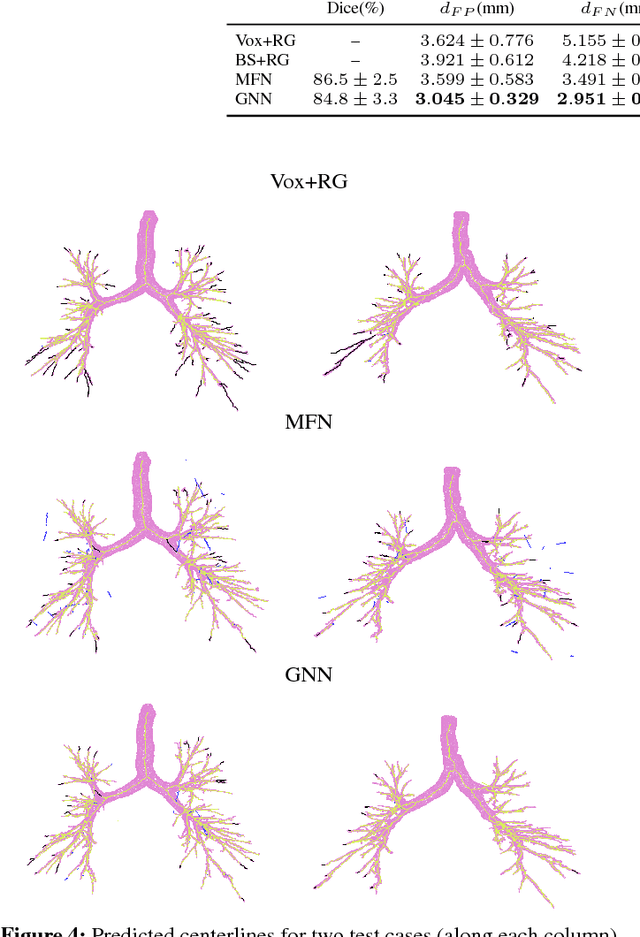
Abstract:Graph refinement, or the task of obtaining subgraphs of interest from over-complete graphs, can have many varied applications. In this work, we extract tree structures from image data by, first deriving a graph-based representation of the volumetric data and then, posing tree extraction as a graph refinement task. We present two methods to perform graph refinement. First, we use mean-field approximation (MFA) to approximate the posterior density over the subgraphs from which the optimal subgraph of interest can be estimated. Mean field networks (MFNs) are used for inference based on the interpretation that iterations of MFA can be seen as feed-forward operations in a neural network. This allows us to learn the model parameters using gradient descent. Second, we present a supervised learning approach using graph neural networks (GNNs) which can be seen as generalisations of MFNs. Subgraphs are obtained by jointly training a GNN based encoder-decoder pair, wherein the encoder learns useful edge embeddings from which the edge probabilities are predicted using a simple decoder. We discuss connections between the two classes of methods and compare them for the task of extracting airways from 3D, low-dose, chest CT data. We show that both the MFN and GNN models show significant improvement when compared to a baseline method, that is similar to a top performing method in the EXACT'09 Challenge, in detecting more branches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge