Mario Krenn
Towards autonomous quantum physics research using LLM agents with access to intelligent tools
Nov 13, 2025Abstract:Artificial intelligence (AI) is used in numerous fields of science, yet the initial research questions and targets are still almost always provided by human researchers. AI-generated creative ideas in science are rare and often vague, so that it remains a human task to execute them. Automating idea generation and implementation in one coherent system would significantly shift the role of humans in the scientific process. Here we present AI-Mandel, an LLM agent that can generate and implement ideas in quantum physics. AI-Mandel formulates ideas from the literature and uses a domain-specific AI tool to turn them into concrete experiment designs that can readily be implemented in laboratories. The generated ideas by AI-Mandel are often scientifically interesting - for two of them we have already written independent scientific follow-up papers. The ideas include new variations of quantum teleportation, primitives of quantum networks in indefinite causal orders, and new concepts of geometric phases based on closed loops of quantum information transfer. AI-Mandel is a prototypical demonstration of an AI physicist that can generate and implement concrete, actionable ideas. Building such a system is not only useful to accelerate science, but it also reveals concrete open challenges on the path to human-level artificial scientists.
Quantum computing and artificial intelligence: status and perspectives
May 29, 2025


Abstract:This white paper discusses and explores the various points of intersection between quantum computing and artificial intelligence (AI). It describes how quantum computing could support the development of innovative AI solutions. It also examines use cases of classical AI that can empower research and development in quantum technologies, with a focus on quantum computing and quantum sensing. The purpose of this white paper is to provide a long-term research agenda aimed at addressing foundational questions about how AI and quantum computing interact and benefit one another. It concludes with a set of recommendations and challenges, including how to orchestrate the proposed theoretical work, align quantum AI developments with quantum hardware roadmaps, estimate both classical and quantum resources - especially with the goal of mitigating and optimizing energy consumption - advance this emerging hybrid software engineering discipline, and enhance European industrial competitiveness while considering societal implications.
Discovering emergent connections in quantum physics research via dynamic word embeddings
Nov 10, 2024Abstract:As the field of quantum physics evolves, researchers naturally form subgroups focusing on specialized problems. While this encourages in-depth exploration, it can limit the exchange of ideas across structurally similar problems in different subfields. To encourage cross-talk among these different specialized areas, data-driven approaches using machine learning have recently shown promise to uncover meaningful connections between research concepts, promoting cross-disciplinary innovation. Current state-of-the-art approaches represent concepts using knowledge graphs and frame the task as a link prediction problem, where connections between concepts are explicitly modeled. In this work, we introduce a novel approach based on dynamic word embeddings for concept combination prediction. Unlike knowledge graphs, our method captures implicit relationships between concepts, can be learned in a fully unsupervised manner, and encodes a broader spectrum of information. We demonstrate that this representation enables accurate predictions about the co-occurrence of concepts within research abstracts over time. To validate the effectiveness of our approach, we provide a comprehensive benchmark against existing methods and offer insights into the interpretability of these embeddings, particularly in the context of quantum physics research. Our findings suggest that this representation offers a more flexible and informative way of modeling conceptual relationships in scientific literature.
Meta-Designing Quantum Experiments with Language Models
Jun 04, 2024



Abstract:Artificial Intelligence (AI) has the potential to significantly advance scientific discovery by finding solutions beyond human capabilities. However, these super-human solutions are often unintuitive and require considerable effort to uncover underlying principles, if possible at all. Here, we show how a code-generating language model trained on synthetic data can not only find solutions to specific problems but can create meta-solutions, which solve an entire class of problems in one shot and simultaneously offer insight into the underlying design principles. Specifically, for the design of new quantum physics experiments, our sequence-to-sequence transformer architecture generates interpretable Python code that describes experimental blueprints for a whole class of quantum systems. We discover general and previously unknown design rules for infinitely large classes of quantum states. The ability to automatically generate generalized patterns in readable computer code is a crucial step toward machines that help discover new scientific understanding -- one of the central aims of physics.
Generation and human-expert evaluation of interesting research ideas using knowledge graphs and large language models
May 27, 2024



Abstract:Advanced artificial intelligence (AI) systems with access to millions of research papers could inspire new research ideas that may not be conceived by humans alone. However, how interesting are these AI-generated ideas, and how can we improve their quality? Here, we introduce SciMuse, a system that uses an evolving knowledge graph built from more than 58 million scientific papers to generate personalized research ideas via an interface to GPT-4. We conducted a large-scale human evaluation with over 100 research group leaders from the Max Planck Society, who ranked more than 4,000 personalized research ideas based on their level of interest. This evaluation allows us to understand the relationships between scientific interest and the core properties of the knowledge graph. We find that data-efficient machine learning can predict research interest with high precision, allowing us to optimize the interest-level of generated research ideas. This work represents a step towards an artificial scientific muse that could catalyze unforeseen collaborations and suggest interesting avenues for scientists.
Forecasting high-impact research topics via machine learning on evolving knowledge graphs
Feb 13, 2024Abstract:The exponential growth in scientific publications poses a severe challenge for human researchers. It forces attention to more narrow sub-fields, which makes it challenging to discover new impactful research ideas and collaborations outside one's own field. While there are ways to predict a scientific paper's future citation counts, they need the research to be finished and the paper written, usually assessing impact long after the idea was conceived. Here we show how to predict the impact of onsets of ideas that have never been published by researchers. For that, we developed a large evolving knowledge graph built from more than 21 million scientific papers. It combines a semantic network created from the content of the papers and an impact network created from the historic citations of papers. Using machine learning, we can predict the dynamic of the evolving network into the future with high accuracy, and thereby the impact of new research directions. We envision that the ability to predict the impact of new ideas will be a crucial component of future artificial muses that can inspire new impactful and interesting scientific ideas.
Deep Quantum Graph Dreaming: Deciphering Neural Network Insights into Quantum Experiments
Sep 13, 2023Abstract:Despite their promise to facilitate new scientific discoveries, the opaqueness of neural networks presents a challenge in interpreting the logic behind their findings. Here, we use a eXplainable-AI (XAI) technique called $inception$ or $deep$ $dreaming$, which has been invented in machine learning for computer vision. We use this techniques to explore what neural networks learn about quantum optics experiments. Our story begins by training a deep neural networks on the properties of quantum systems. Once trained, we "invert" the neural network -- effectively asking how it imagines a quantum system with a specific property, and how it would continuously modify the quantum system to change a property. We find that the network can shift the initial distribution of properties of the quantum system, and we can conceptualize the learned strategies of the neural network. Interestingly, we find that, in the first layers, the neural network identifies simple properties, while in the deeper ones, it can identify complex quantum structures and even quantum entanglement. This is in reminiscence of long-understood properties known in computer vision, which we now identify in a complex natural science task. Our approach could be useful in a more interpretable way to develop new advanced AI-based scientific discovery techniques in quantum physics.
Recent advances in the Self-Referencing Embedding Strings (SELFIES) library
Feb 07, 2023



Abstract:String-based molecular representations play a crucial role in cheminformatics applications, and with the growing success of deep learning in chemistry, have been readily adopted into machine learning pipelines. However, traditional string-based representations such as SMILES are often prone to syntactic and semantic errors when produced by generative models. To address these problems, a novel representation, SELF-referencIng Embedded Strings (SELFIES), was proposed that is inherently 100% robust, alongside an accompanying open-source implementation. Since then, we have generalized SELFIES to support a wider range of molecules and semantic constraints and streamlined its underlying grammar. We have implemented this updated representation in subsequent versions of \selfieslib, where we have also made major advances with respect to design, efficiency, and supported features. Hence, we present the current status of \selfieslib (version 2.1.1) in this manuscript.
Predicting the Future of AI with AI: High-quality link prediction in an exponentially growing knowledge network
Sep 23, 2022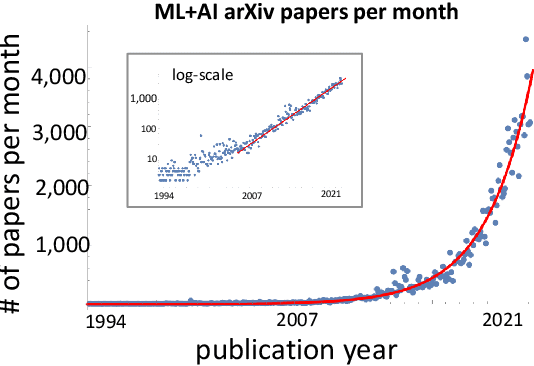
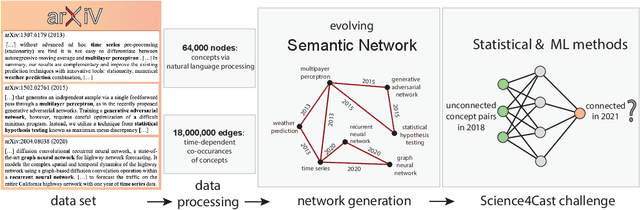
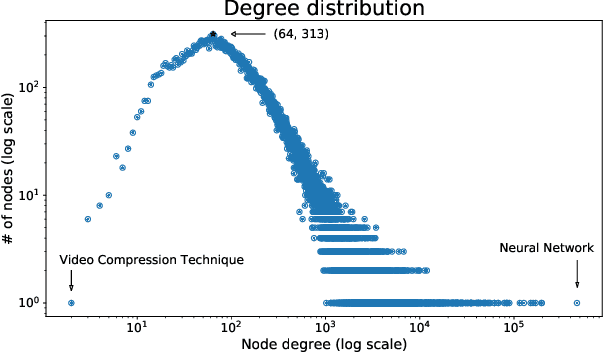
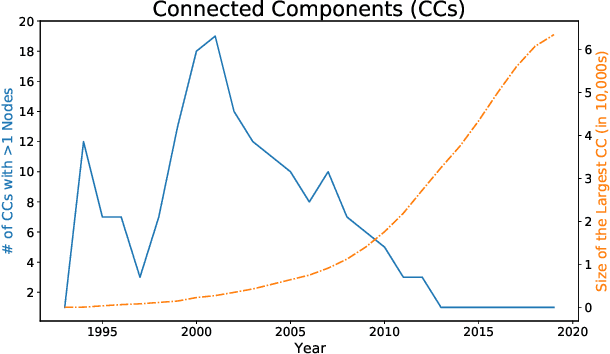
Abstract:A tool that could suggest new personalized research directions and ideas by taking insights from the scientific literature could significantly accelerate the progress of science. A field that might benefit from such an approach is artificial intelligence (AI) research, where the number of scientific publications has been growing exponentially over the last years, making it challenging for human researchers to keep track of the progress. Here, we use AI techniques to predict the future research directions of AI itself. We develop a new graph-based benchmark based on real-world data -- the Science4Cast benchmark, which aims to predict the future state of an evolving semantic network of AI. For that, we use more than 100,000 research papers and build up a knowledge network with more than 64,000 concept nodes. We then present ten diverse methods to tackle this task, ranging from pure statistical to pure learning methods. Surprisingly, the most powerful methods use a carefully curated set of network features, rather than an end-to-end AI approach. It indicates a great potential that can be unleashed for purely ML approaches without human knowledge. Ultimately, better predictions of new future research directions will be a crucial component of more advanced research suggestion tools.
Artificial Intelligence and Machine Learning for Quantum Technologies
Aug 07, 2022Abstract:In recent years, the dramatic progress in machine learning has begun to impact many areas of science and technology significantly. In the present perspective article, we explore how quantum technologies are benefiting from this revolution. We showcase in illustrative examples how scientists in the past few years have started to use machine learning and more broadly methods of artificial intelligence to analyze quantum measurements, estimate the parameters of quantum devices, discover new quantum experimental setups, protocols, and feedback strategies, and generally improve aspects of quantum computing, quantum communication, and quantum simulation. We highlight open challenges and future possibilities and conclude with some speculative visions for the next decade.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge