Lavender Yao Jiang
BPQA Dataset: Evaluating How Well Language Models Leverage Blood Pressures to Answer Biomedical Questions
Mar 06, 2025



Abstract:Clinical measurements such as blood pressures and respiration rates are critical in diagnosing and monitoring patient outcomes. It is an important component of biomedical data, which can be used to train transformer-based language models (LMs) for improving healthcare delivery. It is, however, unclear whether LMs can effectively interpret and use clinical measurements. We investigate two questions: First, can LMs effectively leverage clinical measurements to answer related medical questions? Second, how to enhance an LM's performance on medical question-answering (QA) tasks that involve measurements? We performed a case study on blood pressure readings (BPs), a vital sign routinely monitored by medical professionals. We evaluated the performance of four LMs: BERT, BioBERT, MedAlpaca, and GPT-3.5, on our newly developed dataset, BPQA (Blood Pressure Question Answering). BPQA contains $100$ medical QA pairs that were verified by medical students and designed to rely on BPs . We found that GPT-3.5 and MedAlpaca (larger and medium sized LMs) benefit more from the inclusion of BPs than BERT and BioBERT (small sized LMs). Further, augmenting measurements with labels improves the performance of BioBERT and Medalpaca (domain specific LMs), suggesting that retrieval may be useful for improving domain-specific LMs.
MedG-KRP: Medical Graph Knowledge Representation Probing
Dec 17, 2024Abstract:Large language models (LLMs) have recently emerged as powerful tools, finding many medical applications. LLMs' ability to coalesce vast amounts of information from many sources to generate a response-a process similar to that of a human expert-has led many to see potential in deploying LLMs for clinical use. However, medicine is a setting where accurate reasoning is paramount. Many researchers are questioning the effectiveness of multiple choice question answering (MCQA) benchmarks, frequently used to test LLMs. Researchers and clinicians alike must have complete confidence in LLMs' abilities for them to be deployed in a medical setting. To address this need for understanding, we introduce a knowledge graph (KG)-based method to evaluate the biomedical reasoning abilities of LLMs. Essentially, we map how LLMs link medical concepts in order to better understand how they reason. We test GPT-4, Llama3-70b, and PalmyraMed-70b, a specialized medical model. We enlist a panel of medical students to review a total of 60 LLM-generated graphs and compare these graphs to BIOS, a large biomedical KG. We observe GPT-4 to perform best in our human review but worst in our ground truth comparison; vice-versa with PalmyraMed, the medical model. Our work provides a means of visualizing the medical reasoning pathways of LLMs so they can be implemented in clinical settings safely and effectively.
Refining Packing and Shuffling Strategies for Enhanced Performance in Generative Language Models
Aug 19, 2024



Abstract:Packing and shuffling tokens is a common practice in training auto-regressive language models (LMs) to prevent overfitting and improve efficiency. Typically documents are concatenated to chunks of maximum sequence length (MSL) and then shuffled. However setting the atom size, the length for each data chunk accompanied by random shuffling, to MSL may lead to contextual incoherence due to tokens from different documents being packed into the same chunk. An alternative approach is to utilize padding, another common data packing strategy, to avoid contextual incoherence by only including one document in each shuffled chunk. To optimize both packing strategies (concatenation vs padding), we investigated the optimal atom size for shuffling and compared their performance and efficiency. We found that matching atom size to MSL optimizes performance for both packing methods (concatenation and padding), and padding yields lower final perplexity (higher performance) than concatenation at the cost of more training steps and lower compute efficiency. This trade-off informs the choice of packing methods in training language models.
Generalization in Healthcare AI: Evaluation of a Clinical Large Language Model
Feb 24, 2024Abstract:Advances in large language models (LLMs) provide new opportunities in healthcare for improved patient care, clinical decision-making, and enhancement of physician and administrator workflows. However, the potential of these models importantly depends on their ability to generalize effectively across clinical environments and populations, a challenge often underestimated in early development. To better understand reasons for these challenges and inform mitigation approaches, we evaluated ClinicLLM, an LLM trained on [HOSPITAL]'s clinical notes, analyzing its performance on 30-day all-cause readmission prediction focusing on variability across hospitals and patient characteristics. We found poorer generalization particularly in hospitals with fewer samples, among patients with government and unspecified insurance, the elderly, and those with high comorbidities. To understand reasons for lack of generalization, we investigated sample sizes for fine-tuning, note content (number of words per note), patient characteristics (comorbidity level, age, insurance type, borough), and health system aspects (hospital, all-cause 30-day readmission, and mortality rates). We used descriptive statistics and supervised classification to identify features. We found that, along with sample size, patient age, number of comorbidities, and the number of words in notes are all important factors related to generalization. Finally, we compared local fine-tuning (hospital specific), instance-based augmented fine-tuning and cluster-based fine-tuning for improving generalization. Among these, local fine-tuning proved most effective, increasing AUC by 0.25% to 11.74% (most helpful in settings with limited data). Overall, this study provides new insights for enhancing the deployment of large language models in the societally important domain of healthcare, and improving their performance for broader populations.
Making the Most Out of the Limited Context Length: Predictive Power Varies with Clinical Note Type and Note Section
Jul 13, 2023


Abstract:Recent advances in large language models have led to renewed interest in natural language processing in healthcare using the free text of clinical notes. One distinguishing characteristic of clinical notes is their long time span over multiple long documents. The unique structure of clinical notes creates a new design choice: when the context length for a language model predictor is limited, which part of clinical notes should we choose as the input? Existing studies either choose the inputs with domain knowledge or simply truncate them. We propose a framework to analyze the sections with high predictive power. Using MIMIC-III, we show that: 1) predictive power distribution is different between nursing notes and discharge notes and 2) combining different types of notes could improve performance when the context length is large. Our findings suggest that a carefully selected sampling function could enable more efficient information extraction from clinical notes.
* Our code is publicly available on GitHub (https://github.com/nyuolab/EfficientTransformer)
Edge Entropy as an Indicator of the Effectiveness of GNNs over CNNs for Node Classification
Dec 16, 2020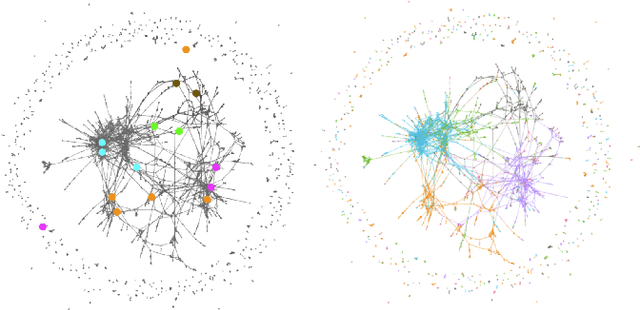
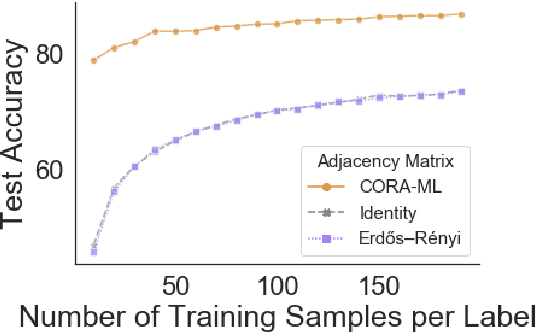
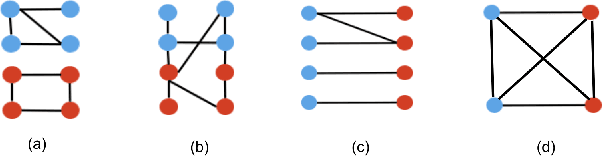
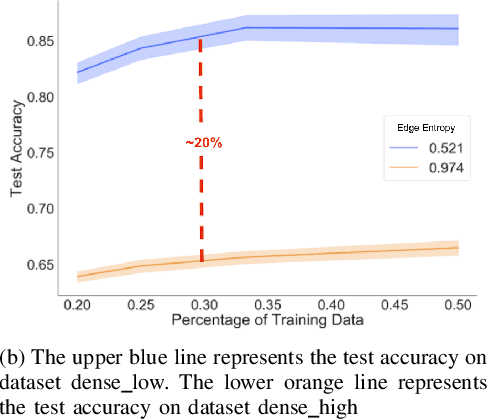
Abstract:Graph neural networks (GNNs) extend convolutional neural networks (CNNs) to graph-based data. A question that arises is how much performance improvement does the underlying graph structure in the GNN provide over the CNN (that ignores this graph structure). To address this question, we introduce edge entropy and evaluate how good an indicator it is for possible performance improvement of GNNs over CNNs. Our results on node classification with synthetic and real datasets show that lower values of edge entropy predict larger expected performance gains of GNNs over CNNs, and, conversely, higher edge entropy leads to expected smaller improvement gains.
Pooling in Graph Convolutional Neural Networks
Apr 07, 2020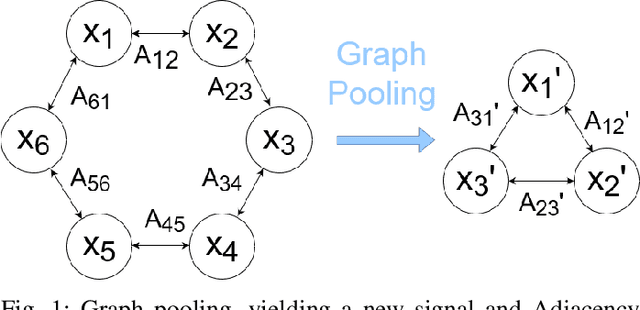
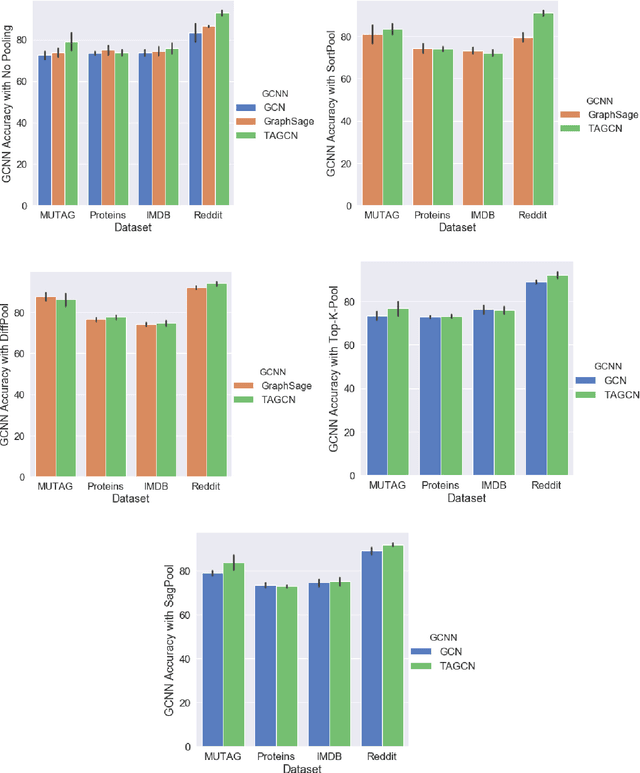
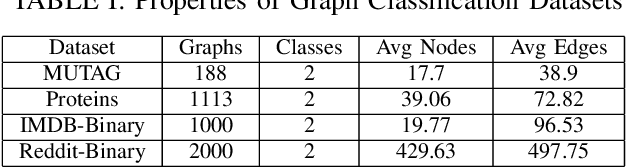
Abstract:Graph convolutional neural networks (GCNNs) are a powerful extension of deep learning techniques to graph-structured data problems. We empirically evaluate several pooling methods for GCNNs, and combinations of those graph pooling methods with three different architectures: GCN, TAGCN, and GraphSAGE. We confirm that graph pooling, especially DiffPool, improves classification accuracy on popular graph classification datasets and find that, on average, TAGCN achieves comparable or better accuracy than GCN and GraphSAGE, particularly for datasets with larger and sparser graph structures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge