Giovanni Volpe
Inchworm-Inspired Soft Robot with Groove-Guided Locomotion
Dec 08, 2025



Abstract:Soft robots require directional control to navigate complex terrains. However, achieving such control often requires multiple actuators, which increases mechanical complexity, complicates control systems, and raises energy consumption. Here, we introduce an inchworm-inspired soft robot whose locomotion direction is controlled passively by patterned substrates. The robot employs a single rolled dielectric elastomer actuator, while groove patterns on a 3D-printed substrate guide its alignment and trajectory. Through systematic experiments, we demonstrate that varying groove angles enables precise control of locomotion direction without the need for complex actuation strategies. This groove-guided approach reduces energy consumption, simplifies robot design, and expands the applicability of bio-inspired soft robots in fields such as search and rescue, pipe inspection, and planetary exploration.
Global graph features unveiled by unsupervised geometric deep learning
Mar 07, 2025



Abstract:Graphs provide a powerful framework for modeling complex systems, but their structural variability makes analysis and classification challenging. To address this, we introduce GAUDI (Graph Autoencoder Uncovering Descriptive Information), a novel unsupervised geometric deep learning framework that captures both local details and global structure. GAUDI employs an innovative hourglass architecture with hierarchical pooling and upsampling layers, linked through skip connections to preserve essential connectivity information throughout the encoding-decoding process. By mapping different realizations of a system - generated from the same underlying parameters - into a continuous, structured latent space, GAUDI disentangles invariant process-level features from stochastic noise. We demonstrate its power across multiple applications, including modeling small-world networks, characterizing protein assemblies from super-resolution microscopy, analyzing collective motion in the Vicsek model, and capturing age-related changes in brain connectivity. This approach not only improves the analysis of complex graphs but also provides new insights into emergent phenomena across diverse scientific domains.
Spatial Clustering of Molecular Localizations with Graph Neural Networks
Nov 29, 2024Abstract:Single-molecule localization microscopy generates point clouds corresponding to fluorophore localizations. Spatial cluster identification and analysis of these point clouds are crucial for extracting insights about molecular organization. However, this task becomes challenging in the presence of localization noise, high point density, or complex biological structures. Here, we introduce MIRO (Multimodal Integration through Relational Optimization), an algorithm that uses recurrent graph neural networks to transform the point clouds in order to improve clustering efficiency when applying conventional clustering techniques. We show that MIRO supports simultaneous processing of clusters of different shapes and at multiple scales, demonstrating improved performance across varied datasets. Our comprehensive evaluation demonstrates MIRO's transformative potential for single-molecule localization applications, showcasing its capability to revolutionize cluster analysis and provide accurate, reliable details of molecular architecture. In addition, MIRO's robust clustering capabilities hold promise for applications in various fields such as neuroscience, for the analysis of neural connectivity patterns, and environmental science, for studying spatial distributions of ecological data.
Diffusion Models to Enhance the Resolution of Microscopy Images: A Tutorial
Sep 24, 2024Abstract:Diffusion models have emerged as a prominent technique in generative modeling with neural networks, making their mark in tasks like text-to-image translation and super-resolution. In this tutorial, we provide a comprehensive guide to build denoising diffusion probabilistic models (DDPMs) from scratch, with a specific focus on transforming low-resolution microscopy images into their corresponding high-resolution versions. We provide the theoretical background, mathematical derivations, and a detailed Python code implementation using PyTorch, along with techniques to enhance model performance.
Deep-learning-powered data analysis in plankton ecology
Sep 15, 2023Abstract:The implementation of deep learning algorithms has brought new perspectives to plankton ecology. Emerging as an alternative approach to established methods, deep learning offers objective schemes to investigate plankton organisms in diverse environments. We provide an overview of deep-learning-based methods including detection and classification of phyto- and zooplankton images, foraging and swimming behaviour analysis, and finally ecological modelling. Deep learning has the potential to speed up the analysis and reduce the human experimental bias, thus enabling data acquisition at relevant temporal and spatial scales with improved reproducibility. We also discuss shortcomings and show how deep learning architectures have evolved to mitigate imprecise readouts. Finally, we suggest opportunities where deep learning is particularly likely to catalyze plankton research. The examples are accompanied by detailed tutorials and code samples that allow readers to apply the methods described in this review to their own data.
Roadmap on Deep Learning for Microscopy
Mar 07, 2023



Abstract:Through digital imaging, microscopy has evolved from primarily being a means for visual observation of life at the micro- and nano-scale, to a quantitative tool with ever-increasing resolution and throughput. Artificial intelligence, deep neural networks, and machine learning are all niche terms describing computational methods that have gained a pivotal role in microscopy-based research over the past decade. This Roadmap is written collectively by prominent researchers and encompasses selected aspects of how machine learning is applied to microscopy image data, with the aim of gaining scientific knowledge by improved image quality, automated detection, segmentation, classification and tracking of objects, and efficient merging of information from multiple imaging modalities. We aim to give the reader an overview of the key developments and an understanding of possibilities and limitations of machine learning for microscopy. It will be of interest to a wide cross-disciplinary audience in the physical sciences and life sciences.
Preface: Characterisation of Physical Processes from Anomalous Diffusion Data
Jan 07, 2023Abstract:Preface to the special issue "Characterisation of Physical Processes from Anomalous Diffusion Data" associated with the Anomalous Diffusion Challenge ( https://andi-challenge.org ) and published in Journal of Physics A: Mathematical and Theoretical. The list of articles included in the special issue can be accessed at https://iopscience.iop.org/journal/1751-8121/page/Characterisation-of-Physical-Processes-from-Anomalous-Diffusion-Data .
Corneal endothelium assessment in specular microscopy images with Fuchs' dystrophy via deep regression of signed distance maps
Oct 13, 2022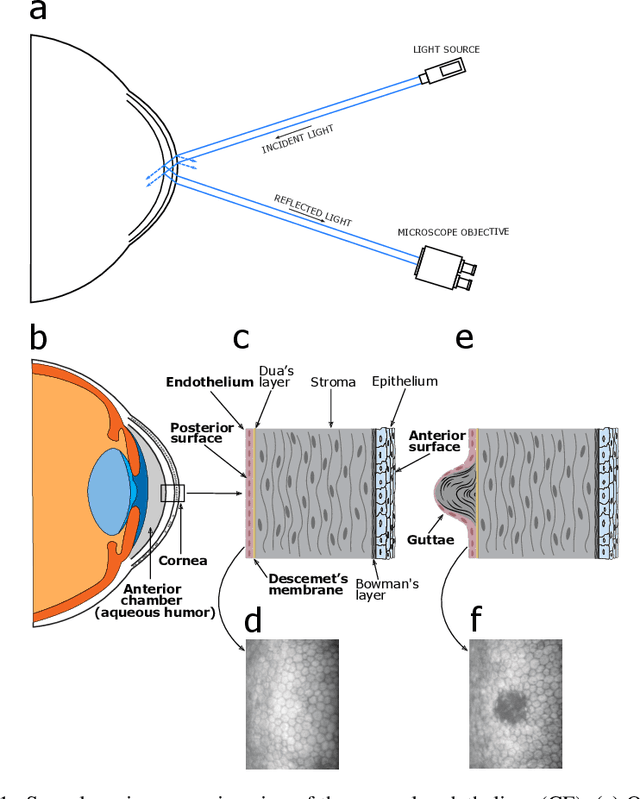
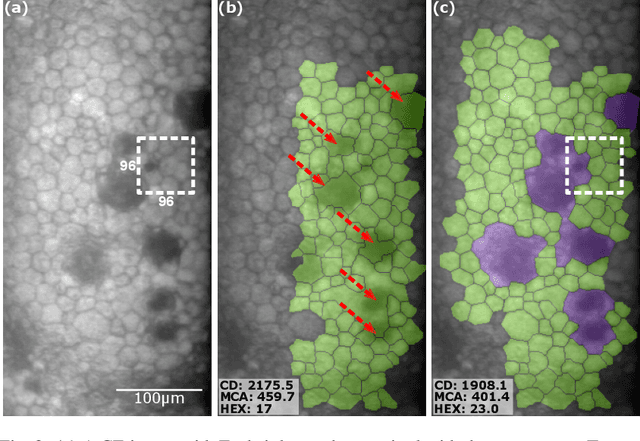
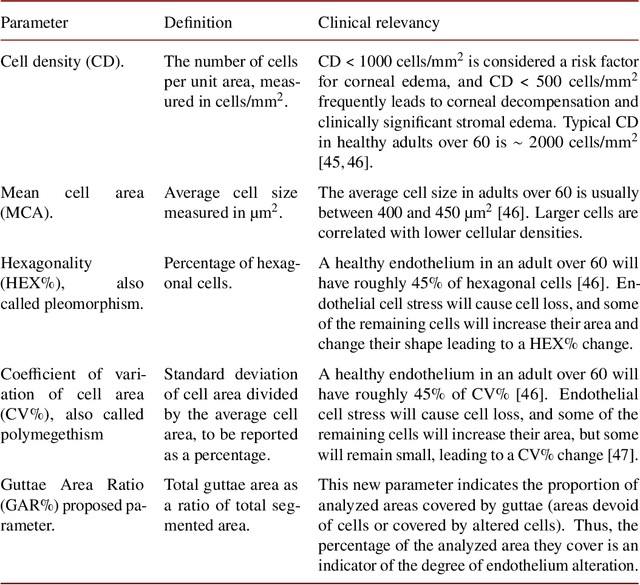

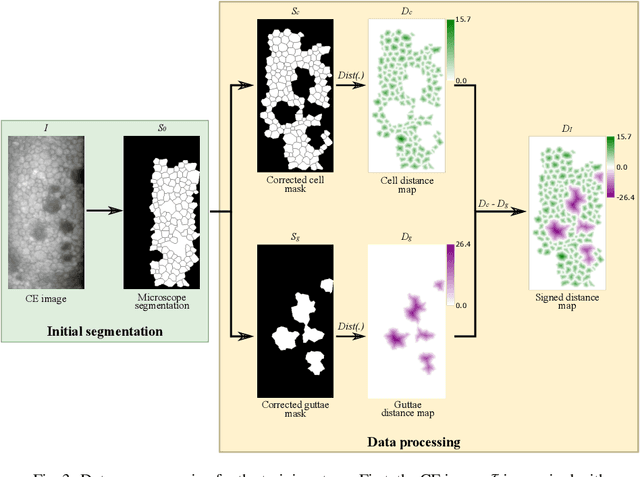
Abstract:Specular microscopy assessment of the human corneal endothelium (CE) in Fuchs' dystrophy is challenging due to the presence of dark image regions called guttae. This paper proposes a UNet-based segmentation approach that requires minimal post-processing and achieves reliable CE morphometric assessment and guttae identification across all degrees of Fuchs' dystrophy. We cast the segmentation problem as a regression task of the cell and gutta signed distance maps instead of a pixel-level classification task as typically done with UNets. Compared to the conventional UNet classification approach, the distance-map regression approach converges faster in clinically relevant parameters. It also produces morphometric parameters that agree with the manually-segmented ground-truth data, namely the average cell density difference of -41.9 cells/mm2 (95% confidence interval (CI) [-306.2, 222.5]) and the average difference of mean cell area of 14.8 um2 (95% CI [-41.9, 71.5]). These results suggest a promising alternative for CE assessment.
Single-shot self-supervised particle tracking
Feb 28, 2022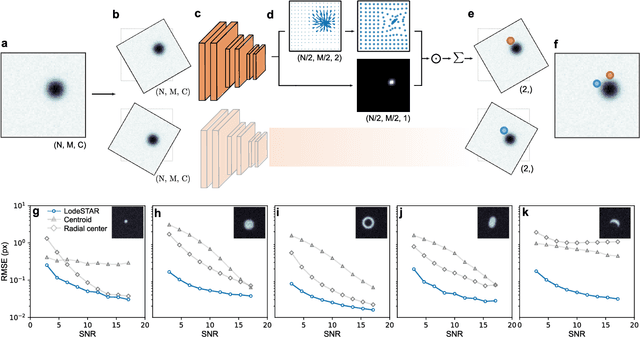
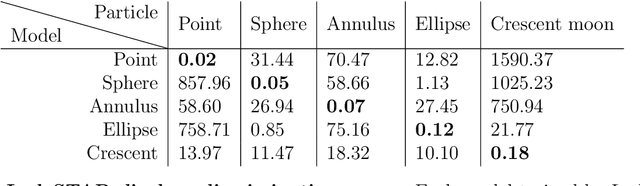
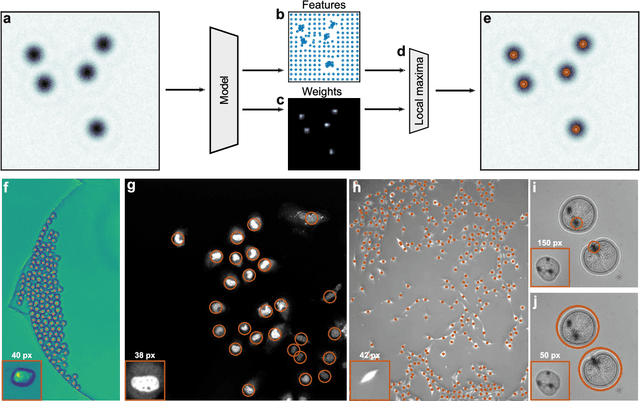
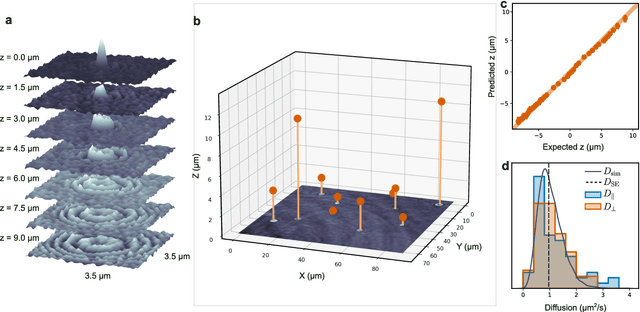
Abstract:Particle tracking is a fundamental task in digital microscopy. Recently, machine-learning approaches have made great strides in overcoming the limitations of more classical approaches. The training of state-of-the-art machine-learning methods almost universally relies on either vast amounts of labeled experimental data or the ability to numerically simulate realistic datasets. However, the data produced by experiments are often challenging to label and cannot be easily reproduced numerically. Here, we propose a novel deep-learning method, named LodeSTAR (Low-shot deep Symmetric Tracking And Regression), that learns to tracks objects with sub-pixel accuracy from a single unlabeled experimental image. This is made possible by exploiting the inherent roto-translational symmetries of the data. We demonstrate that LodeSTAR outperforms traditional methods in terms of accuracy. Furthermore, we analyze challenging experimental data containing densely packed cells or noisy backgrounds. We also exploit additional symmetries to extend the measurable particle properties to the particle's vertical position by propagating the signal in Fourier space and its polarizability by scaling the signal strength. Thanks to the ability to train deep-learning models with a single unlabeled image, LodeSTAR can accelerate the development of high-quality microscopic analysis pipelines for engineering, biology, and medicine.
Microplankton life histories revealed by holographic microscopy and deep learning
Feb 18, 2022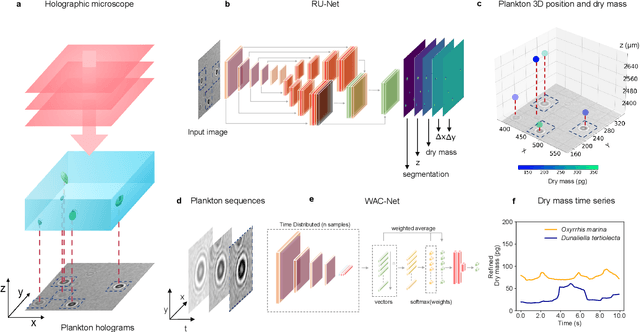
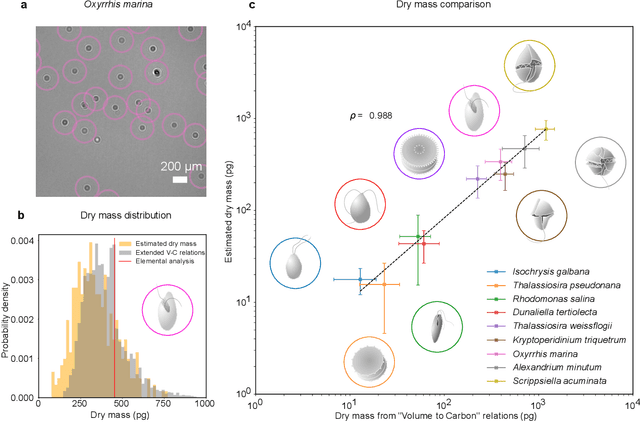
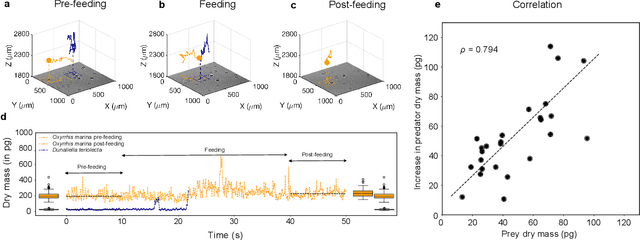
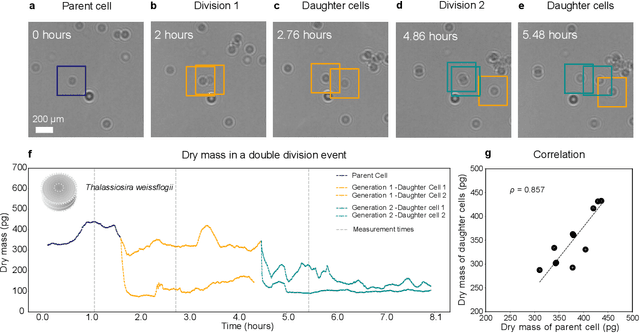
Abstract:The marine microbial food web plays a central role in the global carbon cycle. Our mechanistic understanding of the ocean, however, is biased towards its larger constituents, while rates and biomass fluxes in the microbial food web are mainly inferred from indirect measurements and ensemble averages. Yet, resolution at the level of the individual microplankton is required to advance our understanding of the oceanic food web. Here, we demonstrate that, by combining holographic microscopy with deep learning, we can follow microplanktons throughout their lifespan, continuously measuring their three dimensional position and dry mass. The deep learning algorithms circumvent the computationally intensive processing of holographic data and allow rapid measurements over extended time periods. This permits us to reliably estimate growth rates, both in terms of dry mass increase and cell divisions, as well as to measure trophic interactions between species such as predation events. The individual resolution provides information about selectivity, individual feeding rates and handling times for individual microplanktons. This method is particularly useful to explore the flux of carbon through micro-zooplankton, the most important and least known group of primary consumers in the global oceans. We exemplify this by detailed descriptions of micro-zooplankton feeding events, cell divisions, and long term monitoring of single cells from division to division.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge