Hongyu Dong
Key-Embedded Privacy for Decentralized AI in Biomedical Omics
Mar 30, 2026Abstract:The rapid adoption of data-driven methods in biomedicine has intensified concerns over privacy, governance, and regulation, limiting raw data sharing and hindering the assembly of representative cohorts for clinically relevant AI. This landscape necessitates practical, efficient privacy solutions, as cryptographic defenses often impose heavy overhead and differential privacy can degrade performance, leading to sub-optimal outcomes in real-world settings. Here, we present a lightweight federated learning method, INFL, based on Implicit Neural Representations that addresses these challenges. Our approach integrates plug-and-play, coordinate-conditioned modules into client models, embeds a secret key directly into the architecture, and supports seamless aggregation across heterogeneous sites. Across diverse biomedical omics tasks, including cohort-scale classification in bulk proteomics, regression for perturbation prediction in single-cell transcriptomics, and clustering in spatial transcriptomics and multi-omics with both public and private data, we demonstrate that INFL achieves strong, controllable privacy while maintaining utility, preserving the performance necessary for downstream scientific and clinical applications.
Deeply Supervised Layer Selective Attention Network: Towards Label-Efficient Learning for Medical Image Classification
Sep 28, 2022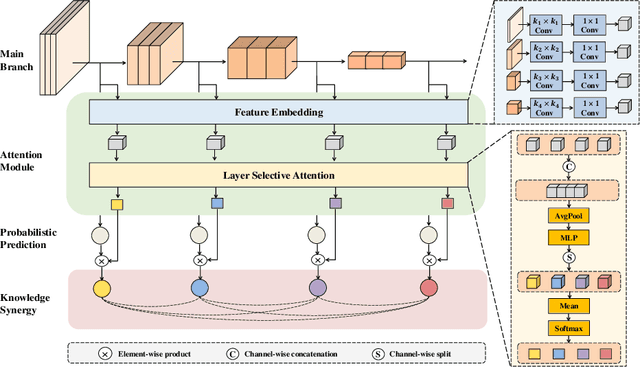

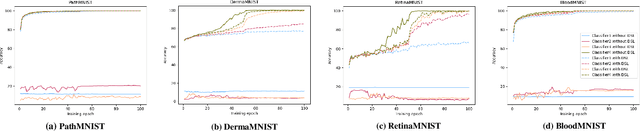
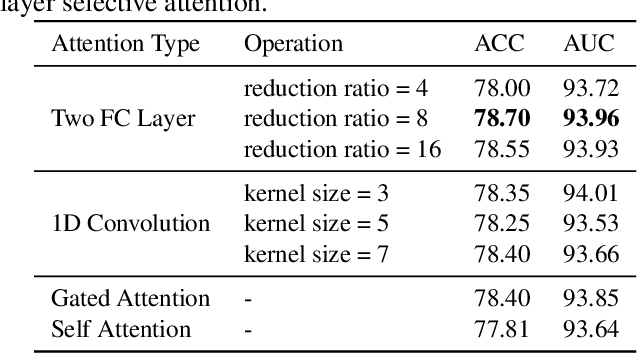
Abstract:Labeling medical images depends on professional knowledge, making it difficult to acquire large amount of annotated medical images with high quality in a short time. Thus, making good use of limited labeled samples in a small dataset to build a high-performance model is the key to medical image classification problem. In this paper, we propose a deeply supervised Layer Selective Attention Network (LSANet), which comprehensively uses label information in feature-level and prediction-level supervision. For feature-level supervision, in order to better fuse the low-level features and high-level features, we propose a novel visual attention module, Layer Selective Attention (LSA), to focus on the feature selection of different layers. LSA introduces a weight allocation scheme which can dynamically adjust the weighting factor of each auxiliary branch during the whole training process to further enhance deeply supervised learning and ensure its generalization. For prediction-level supervision, we adopt the knowledge synergy strategy to promote hierarchical information interactions among all supervision branches via pairwise knowledge matching. Using the public dataset, MedMNIST, which is a large-scale benchmark for biomedical image classification covering diverse medical specialties, we evaluate LSANet on multiple mainstream CNN architectures and various visual attention modules. The experimental results show the substantial improvements of our proposed method over its corresponding counterparts, demonstrating that LSANet can provide a promising solution for label-efficient learning in the field of medical image classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge