Pin Li
Key-Embedded Privacy for Decentralized AI in Biomedical Omics
Mar 30, 2026Abstract:The rapid adoption of data-driven methods in biomedicine has intensified concerns over privacy, governance, and regulation, limiting raw data sharing and hindering the assembly of representative cohorts for clinically relevant AI. This landscape necessitates practical, efficient privacy solutions, as cryptographic defenses often impose heavy overhead and differential privacy can degrade performance, leading to sub-optimal outcomes in real-world settings. Here, we present a lightweight federated learning method, INFL, based on Implicit Neural Representations that addresses these challenges. Our approach integrates plug-and-play, coordinate-conditioned modules into client models, embeds a secret key directly into the architecture, and supports seamless aggregation across heterogeneous sites. Across diverse biomedical omics tasks, including cohort-scale classification in bulk proteomics, regression for perturbation prediction in single-cell transcriptomics, and clustering in spatial transcriptomics and multi-omics with both public and private data, we demonstrate that INFL achieves strong, controllable privacy while maintaining utility, preserving the performance necessary for downstream scientific and clinical applications.
Recognition of convolutional neural network based on CUDA Technology
Aug 27, 2018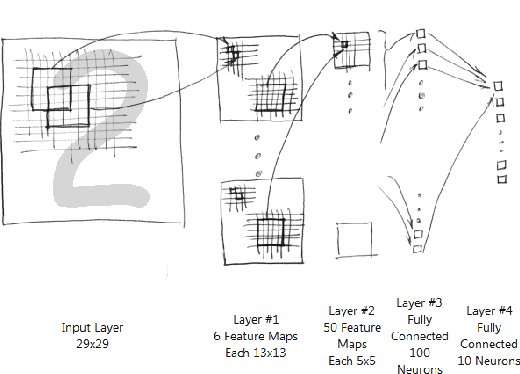
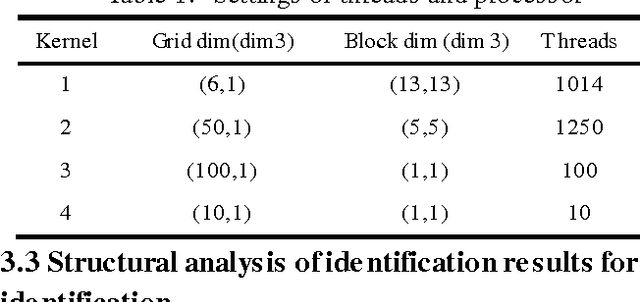

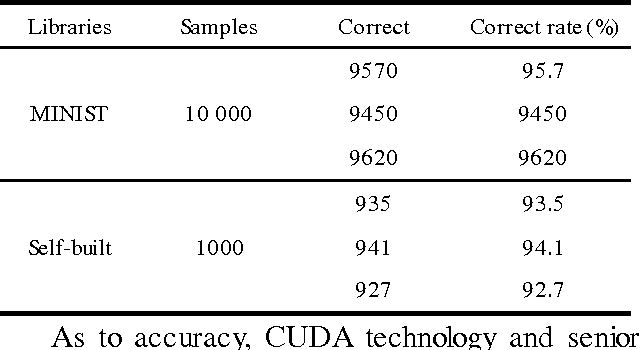
Abstract:For the problem whether Graphic Processing Unit(GPU),the stream processor with high performance of floating-point computing is applicable to neural networks, this paper proposes the parallel recognition algorithm of Convolutional Neural Networks(CNNs).It adopts Compute Unified Device Architecture(CUDA)technology, definite the parallel data structures, and describes the mapping mechanism for computing tasks on CUDA. It compares the parallel recognition algorithm achieved on GPU of GTX200 hardware architecture with the serial algorithm on CPU. It improves speed by nearly 60 times. Result shows that GPU based the stream processor architecture ate more applicable to some related applications about neural networks than CPU.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge