Haopeng Liu
Refinement Contrastive Learning of Cell-Gene Associations for Unsupervised Cell Type Identification
Dec 11, 2025Abstract:Unsupervised cell type identification is crucial for uncovering and characterizing heterogeneous populations in single cell omics studies. Although a range of clustering methods have been developed, most focus exclusively on intrinsic cellular structure and ignore the pivotal role of cell-gene associations, which limits their ability to distinguish closely related cell types. To this end, we propose a Refinement Contrastive Learning framework (scRCL) that explicitly incorporates cell-gene interactions to derive more informative representations. Specifically, we introduce two contrastive distribution alignment components that reveal reliable intrinsic cellular structures by effectively exploiting cell-cell structural relationships. Additionally, we develop a refinement module that integrates gene-correlation structure learning to enhance cell embeddings by capturing underlying cell-gene associations. This module strengthens connections between cells and their associated genes, refining the representation learning to exploiting biologically meaningful relationships. Extensive experiments on several single-cell RNA-seq and spatial transcriptomics benchmark datasets demonstrate that our method consistently outperforms state-of-the-art baselines in cell-type identification accuracy. Moreover, downstream biological analyses confirm that the recovered cell populations exhibit coherent gene-expression signatures, further validating the biological relevance of our approach. The code is available at https://github.com/THPengL/scRCL.
Detection and Attention: Diagnosing Pulmonary Lung Cancer from CT by Imitating Physicians
Dec 14, 2017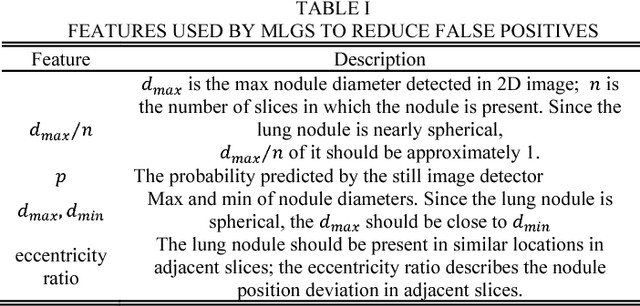

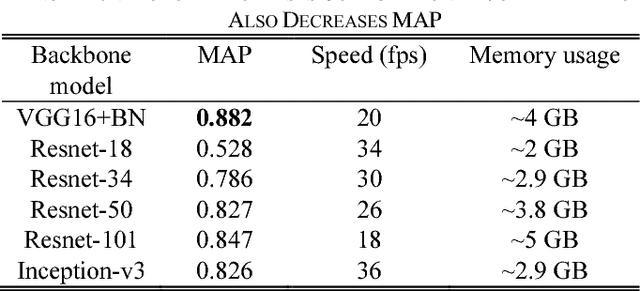
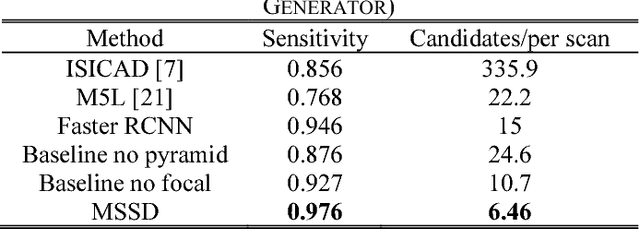
Abstract:This paper proposes a novel and efficient method to build a Computer-Aided Diagnoses (CAD) system for lung nodule detection based on Computed Tomography (CT). This task was treated as an Object Detection on Video (VID) problem by imitating how a radiologist reads CT scans. A lung nodule detector was trained to automatically learn nodule features from still images to detect lung nodule candidates with both high recall and accuracy. Unlike previous work which used 3-dimensional information around the nodule to reduce false positives, we propose two simple but efficient methods, Multi-slice propagation (MSP) and Motionless-guide suppression (MLGS), which analyze sequence information of CT scans to reduce false negatives and suppress false positives. We evaluated our method in open-source LUNA16 dataset which contains 888 CT scans, and obtained state-of-the-art result (Free-Response Receiver Operating Characteristic score of 0.892) with detection speed (end to end within 20 seconds per patient on a single NVidia GTX 1080) much higher than existing methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge