Hangqi Zhou
ZScribbleSeg: A comprehensive segmentation framework with modeling of efficient annotation and maximization of scribble supervision
May 07, 2026Abstract:Curating fully annotated datasets for medical image segmentation is labour-intensive and expertise-demanding. To alleviate this problem, prior studies have explored scribble annotations for weakly supervised segmentation. Existing solutions mainly compute losses on annotated areas and generate pseudo labels by propagating annotations to adjacent regions. However, these methods often suffer from inaccurate and unrealistic segmentations due to insufficient supervision and incomplete shape information. In contrast, we first investigate the principle of good scribble annotations, which leads to efficient scribble forms via supervision maximization and randomness simulation. We further introduce regularization terms to encode the spatial relationship and the shape constraints, where the EM algorithm is utilized to estimate the mixture ratios of label classes. These ratios are critical in identifying the unlabeled pixels for each class and correcting erroneous predictions, thus the accurate estimation lays the foundation for the incorporation of spatial prior. Finally, we integrate the efficient scribble supervision with the prior into a framework, referred to as ZScribbleSeg, and apply it to multiple scenarios. Leveraging only scribble annotations, ZScribbleSeg achieves competitive performance on six segmentation tasks including ACDC, MSCMRseg, BTCV, MyoPS, Decathlon-BrainTumor and Decathlon-Prostate. Our code will be released via https://github.com/DLwbm123/ZScribbleSeg.
Beyond Forgetting in Continual Medical Image Segmentation: A Comprehensive Benchmark Study
May 07, 2026Abstract:Continual learning (CL) is essential for deploying medical image segmentation models in clinical environments where imaging domains, anatomical targets, and diagnostic tasks evolve over time. However, continual segmentation still faces three main challenges. First, the scenarios for this task remain insufficiently standardized for real-world clinical settings. Second, existing research has been primarily focused on mitigating forgetting, overlooking the other essential properties such as plasticity. Third, a benchmark work with comprehensive evaluation on existing methods is stll desirable. To address these gaps, we present such benchmark study of continual medical image segmentation. We first define three clinically motivated scenarios, namely Domain-CL, Class-CL, and Organ-CL, to respectively capture the cross-center domain shift, the incremental anatomical structure segmentation, and the cross-organ segmentation. We then introduce an evaluation framework that measures not only general performance and forgetting, but also plasticity, forward generalizability, parameter efficiency, and replay burden. The results, from extensive experiments with representative CL methods, showed that it was still challenging to develop a model that could satisfy all the requirements simultaneously. Nevertheless, these studies also suggested that the replay-based methods achieve the best overall balance between stability and plasticity, the parameter-isolation methods should be effective at reducing forgetting, though at the cost of increased model size, and the forward generalizability remain a significantly understudied aspect of this research field. Finally, we discuss related learning paradigms and outline future directions for continual medical image segmentation.
Learning Concept-Driven Logical Rules for Interpretable and Generalizable Medical Image Classification
May 20, 2025



Abstract:The pursuit of decision safety in clinical applications highlights the potential of concept-based methods in medical imaging. While these models offer active interpretability, they often suffer from concept leakages, where unintended information within soft concept representations undermines both interpretability and generalizability. Moreover, most concept-based models focus solely on local explanations (instance-level), neglecting the global decision logic (dataset-level). To address these limitations, we propose Concept Rule Learner (CRL), a novel framework to learn Boolean logical rules from binarized visual concepts. CRL employs logical layers to capture concept correlations and extract clinically meaningful rules, thereby providing both local and global interpretability. Experiments on two medical image classification tasks show that CRL achieves competitive performance with existing methods while significantly improving generalizability to out-of-distribution data. The code of our work is available at https://github.com/obiyoag/crl.
Multi-Center Fetal Brain Tissue Annotation (FeTA) Challenge 2022 Results
Feb 08, 2024



Abstract:Segmentation is a critical step in analyzing the developing human fetal brain. There have been vast improvements in automatic segmentation methods in the past several years, and the Fetal Brain Tissue Annotation (FeTA) Challenge 2021 helped to establish an excellent standard of fetal brain segmentation. However, FeTA 2021 was a single center study, and the generalizability of algorithms across different imaging centers remains unsolved, limiting real-world clinical applicability. The multi-center FeTA Challenge 2022 focuses on advancing the generalizability of fetal brain segmentation algorithms for magnetic resonance imaging (MRI). In FeTA 2022, the training dataset contained images and corresponding manually annotated multi-class labels from two imaging centers, and the testing data contained images from these two imaging centers as well as two additional unseen centers. The data from different centers varied in many aspects, including scanners used, imaging parameters, and fetal brain super-resolution algorithms applied. 16 teams participated in the challenge, and 17 algorithms were evaluated. Here, a detailed overview and analysis of the challenge results are provided, focusing on the generalizability of the submissions. Both in- and out of domain, the white matter and ventricles were segmented with the highest accuracy, while the most challenging structure remains the cerebral cortex due to anatomical complexity. The FeTA Challenge 2022 was able to successfully evaluate and advance generalizability of multi-class fetal brain tissue segmentation algorithms for MRI and it continues to benchmark new algorithms. The resulting new methods contribute to improving the analysis of brain development in utero.
BayeSeg: Bayesian Modeling for Medical Image Segmentation with Interpretable Generalizability
Mar 03, 2023Abstract:Due to the cross-domain distribution shift aroused from diverse medical imaging systems, many deep learning segmentation methods fail to perform well on unseen data, which limits their real-world applicability. Recent works have shown the benefits of extracting domain-invariant representations on domain generalization. However, the interpretability of domain-invariant features remains a great challenge. To address this problem, we propose an interpretable Bayesian framework (BayeSeg) through Bayesian modeling of image and label statistics to enhance model generalizability for medical image segmentation. Specifically, we first decompose an image into a spatial-correlated variable and a spatial-variant variable, assigning hierarchical Bayesian priors to explicitly force them to model the domain-stable shape and domain-specific appearance information respectively. Then, we model the segmentation as a locally smooth variable only related to the shape. Finally, we develop a variational Bayesian framework to infer the posterior distributions of these explainable variables. The framework is implemented with neural networks, and thus is referred to as deep Bayesian segmentation. Quantitative and qualitative experimental results on prostate segmentation and cardiac segmentation tasks have shown the effectiveness of our proposed method. Moreover, we investigated the interpretability of BayeSeg by explaining the posteriors and analyzed certain factors that affect the generalization ability through further ablation studies. Our code will be released via https://zmiclab.github.io/projects.html, once the manuscript is accepted for publication.
Joint Modeling of Image and Label Statistics for Enhancing Model Generalizability of Medical Image Segmentation
Jun 09, 2022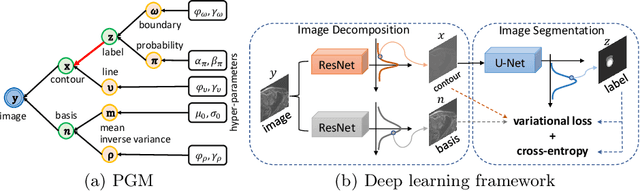
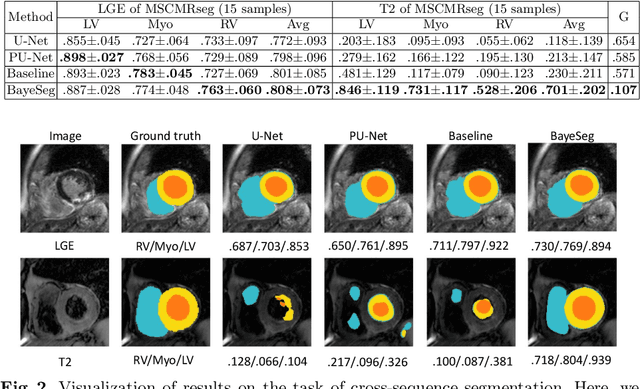
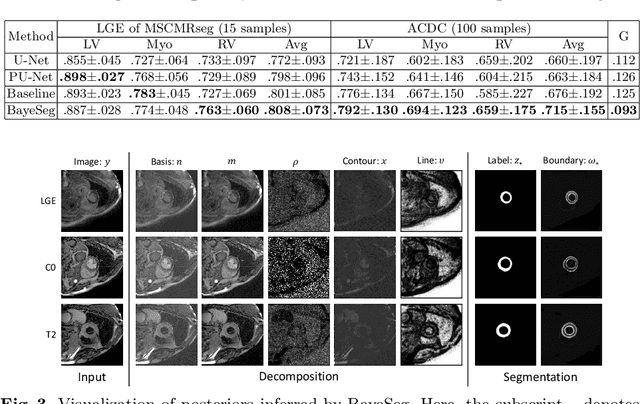
Abstract:Although supervised deep-learning has achieved promising performance in medical image segmentation, many methods cannot generalize well on unseen data, limiting their real-world applicability. To address this problem, we propose a deep learning-based Bayesian framework, which jointly models image and label statistics, utilizing the domain-irrelevant contour of a medical image for segmentation. Specifically, we first decompose an image into components of contour and basis. Then, we model the expected label as a variable only related to the contour. Finally, we develop a variational Bayesian framework to infer the posterior distributions of these variables, including the contour, the basis, and the label. The framework is implemented with neural networks, thus is referred to as deep Bayesian segmentation. Results on the task of cross-sequence cardiac MRI segmentation show that our method set a new state of the art for model generalizability. Particularly, the BayeSeg model trained with LGE MRI generalized well on T2 images and outperformed other models with great margins, i.e., over 0.47 in terms of average Dice. Our code is available at https://zmiclab.github.io/projects.html.
VSpSR: Explorable Super-Resolution via Variational Sparse Representation
Apr 17, 2021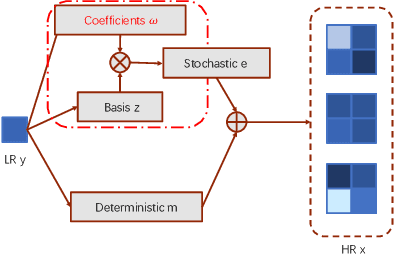
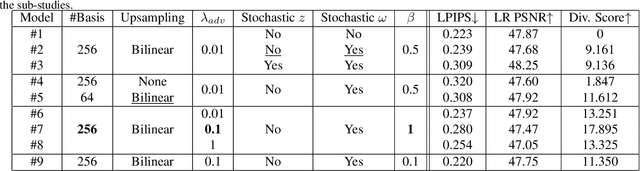
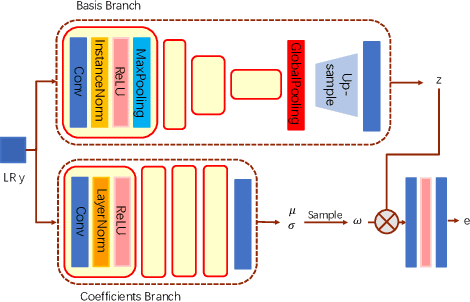
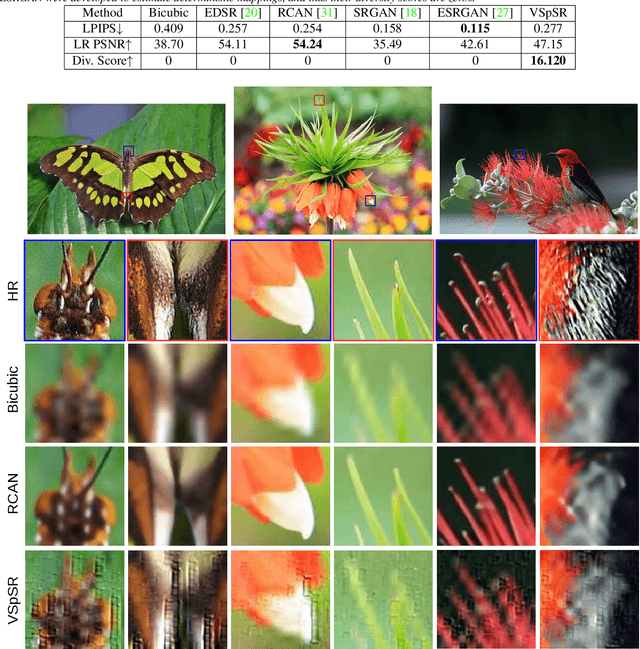
Abstract:Super-resolution (SR) is an ill-posed problem, which means that infinitely many high-resolution (HR) images can be degraded to the same low-resolution (LR) image. To study the one-to-many stochastic SR mapping, we implicitly represent the non-local self-similarity of natural images and develop a Variational Sparse framework for Super-Resolution (VSpSR) via neural networks. Since every small patch of a HR image can be well approximated by the sparse representation of atoms in an over-complete dictionary, we design a two-branch module, i.e., VSpM, to explore the SR space. Concretely, one branch of VSpM extracts patch-level basis from the LR input, and the other branch infers pixel-wise variational distributions with respect to the sparse coefficients. By repeatedly sampling coefficients, we could obtain infinite sparse representations, and thus generate diverse HR images. According to the preliminary results of NTIRE 2021 challenge on learning SR space, our team (FudanZmic21) ranks 7-th in terms of released scores. The implementation of VSpSR is released at https://zmiclab.github.io/.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge