Haiming Tang
RelBench v2: A Large-Scale Benchmark and Repository for Relational Data
Feb 13, 2026Abstract:Relational deep learning (RDL) has emerged as a powerful paradigm for learning directly on relational databases by modeling entities and their relationships across multiple interconnected tables. As this paradigm evolves toward larger models and relational foundation models, scalable and realistic benchmarks are essential for enabling systematic evaluation and progress. In this paper, we introduce RelBench v2, a major expansion of the RelBench benchmark for RDL. RelBench v2 adds four large-scale relational datasets spanning scholarly publications, enterprise resource planning, consumer platforms, and clinical records, increasing the benchmark to 11 datasets comprising over 22 million rows across 29 tables. We further introduce autocomplete tasks, a new class of predictive objectives that require models to infer missing attribute values directly within relational tables while respecting temporal constraints, expanding beyond traditional forecasting tasks constructed via SQL queries. In addition, RelBench v2 expands beyond its native datasets by integrating external benchmarks and evaluation frameworks: we translate event streams from the Temporal Graph Benchmark into relational schemas for unified relational-temporal evaluation, interface with ReDeLEx to provide uniform access to 70+ real-world databases suitable for pretraining, and incorporate 4DBInfer datasets and tasks to broaden multi-table prediction coverage. Experimental results demonstrate that RDL models consistently outperform single-table baselines across autocomplete, forecasting, and recommendation tasks, highlighting the importance of modeling relational structure explicitly.
aiXiv: A Next-Generation Open Access Ecosystem for Scientific Discovery Generated by AI Scientists
Aug 20, 2025Abstract:Recent advances in large language models (LLMs) have enabled AI agents to autonomously generate scientific proposals, conduct experiments, author papers, and perform peer reviews. Yet this flood of AI-generated research content collides with a fragmented and largely closed publication ecosystem. Traditional journals and conferences rely on human peer review, making them difficult to scale and often reluctant to accept AI-generated research content; existing preprint servers (e.g. arXiv) lack rigorous quality-control mechanisms. Consequently, a significant amount of high-quality AI-generated research lacks appropriate venues for dissemination, hindering its potential to advance scientific progress. To address these challenges, we introduce aiXiv, a next-generation open-access platform for human and AI scientists. Its multi-agent architecture allows research proposals and papers to be submitted, reviewed, and iteratively refined by both human and AI scientists. It also provides API and MCP interfaces that enable seamless integration of heterogeneous human and AI scientists, creating a scalable and extensible ecosystem for autonomous scientific discovery. Through extensive experiments, we demonstrate that aiXiv is a reliable and robust platform that significantly enhances the quality of AI-generated research proposals and papers after iterative revising and reviewing on aiXiv. Our work lays the groundwork for a next-generation open-access ecosystem for AI scientists, accelerating the publication and dissemination of high-quality AI-generated research content. Code is available at https://github.com/aixiv-org. Website is available at https://forms.gle/DxQgCtXFsJ4paMtn8.
Region Proposal Rectification Towards Robust Instance Segmentation of Biological Images
Mar 06, 2022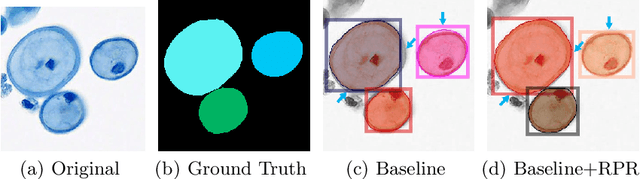
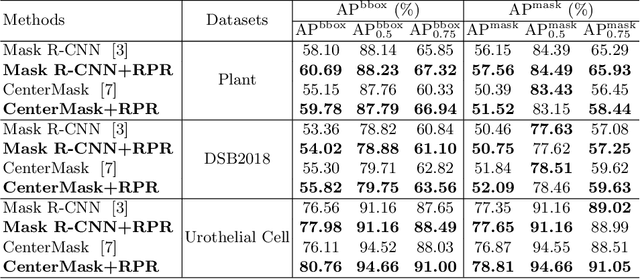
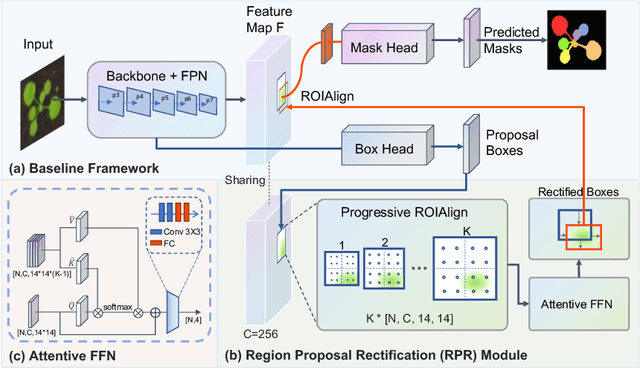
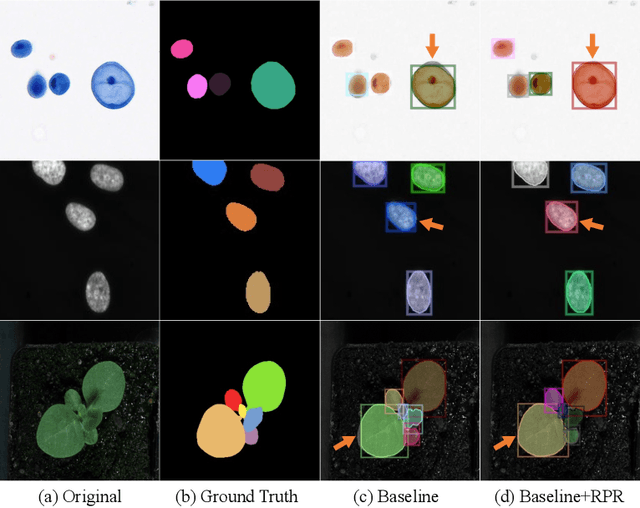
Abstract:Top-down instance segmentation framework has shown its superiority in object detection compared to the bottom-up framework. While it is efficient in addressing over-segmentation, top-down instance segmentation suffers from over-crop problem. However, a complete segmentation mask is crucial for biological image analysis as it delivers important morphological properties such as shapes and volumes. In this paper, we propose a region proposal rectification (RPR) module to address this challenging incomplete segmentation problem. In particular, we offer a progressive ROIAlign module to introduce neighbor information into a series of ROIs gradually. The ROI features are fed into an attentive feed-forward network (FFN) for proposal box regression. With additional neighbor information, the proposed RPR module shows significant improvement in correction of region proposal locations and thereby exhibits favorable instance segmentation performances on three biological image datasets compared to state-of-the-art baseline methods. Experimental results demonstrate that the proposed RPR module is effective in both anchor-based and anchor-free top-down instance segmentation approaches, suggesting the proposed method can be applied to general top-down instance segmentation of biological images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge