Elsa Angelini
TSI
Co-Seg: An Image Segmentation Framework Against Label Corruption
Jan 31, 2021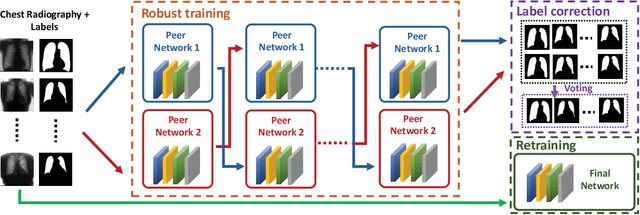
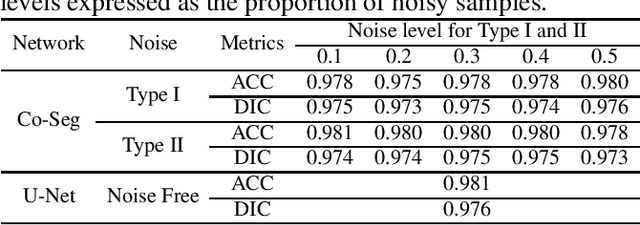
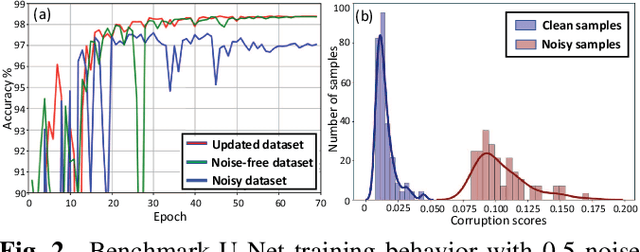
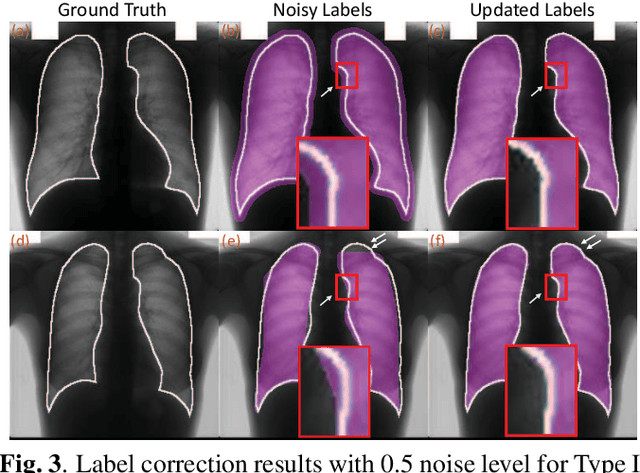
Abstract:Supervised deep learning performance is heavily tied to the availability of high-quality labels for training. Neural networks can gradually overfit corrupted labels if directly trained on noisy datasets, leading to severe performance degradation at test time. In this paper, we propose a novel deep learning framework, namely Co-Seg, to collaboratively train segmentation networks on datasets which include low-quality noisy labels. Our approach first trains two networks simultaneously to sift through all samples and obtain a subset with reliable labels. Then, an efficient yet easily-implemented label correction strategy is applied to enrich the reliable subset. Finally, using the updated dataset, we retrain the segmentation network to finalize its parameters. Experiments in two noisy labels scenarios demonstrate that our proposed model can achieve results comparable to those obtained from supervised learning trained on the noise-free labels. In addition, our framework can be easily implemented in any segmentation algorithm to increase its robustness to noisy labels.
Suggestive Annotation of Brain Tumour Images with Gradient-guided Sampling
Jul 03, 2020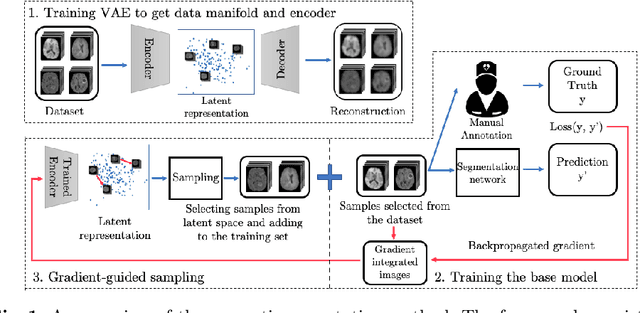
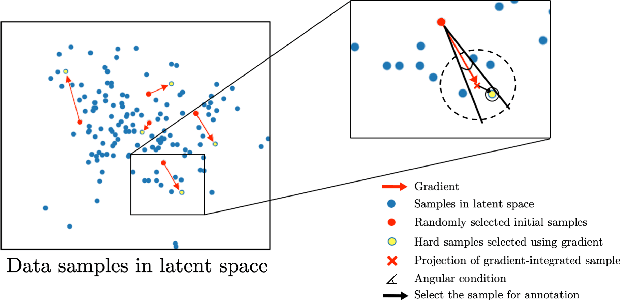
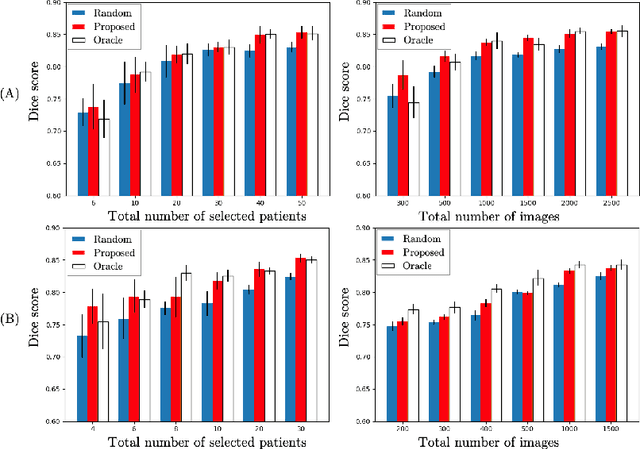
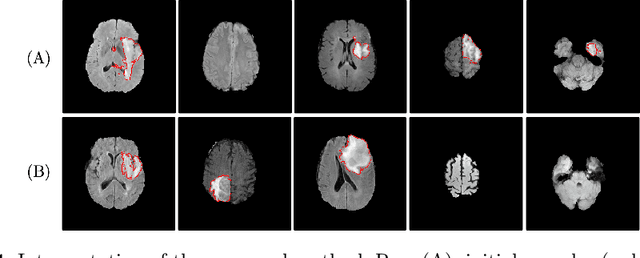
Abstract:Machine learning has been widely adopted for medical image analysis in recent years given its promising performance in image segmentation and classification tasks. As a data-driven science, the success of machine learning, in particular supervised learning, largely depends on the availability of manually annotated datasets. For medical imaging applications, such annotated datasets are not easy to acquire. It takes a substantial amount of time and resource to curate an annotated medical image set. In this paper, we propose an efficient annotation framework for brain tumour images that is able to suggest informative sample images for human experts to annotate. Our experiments show that training a segmentation model with only 19% suggestively annotated patient scans from BraTS 2019 dataset can achieve a comparable performance to training a model on the full dataset for whole tumour segmentation task. It demonstrates a promising way to save manual annotation cost and improve data efficiency in medical imaging applications.
Simultaneous Left Atrium Anatomy and Scar Segmentations via Deep Learning in Multiview Information with Attention
Feb 02, 2020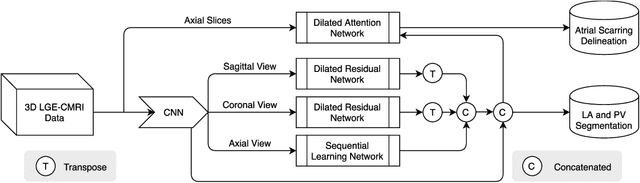
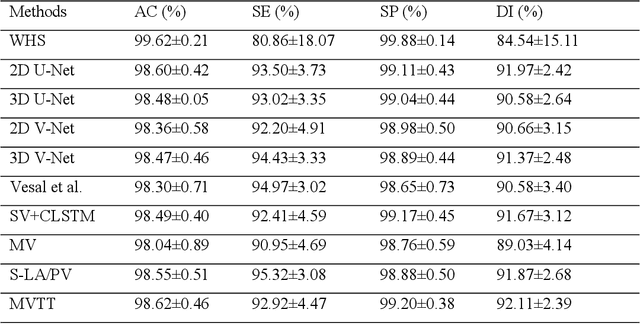
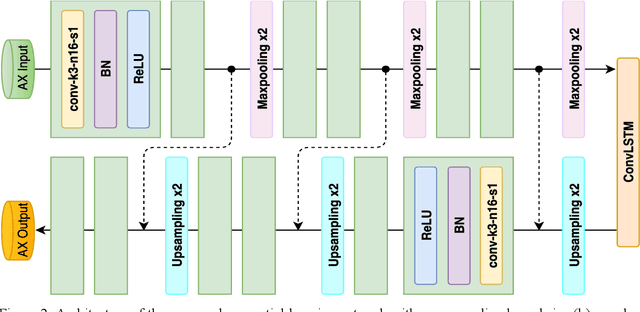
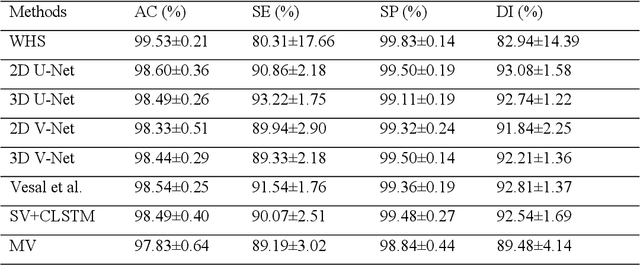
Abstract:Three-dimensional late gadolinium enhanced (LGE) cardiac MR (CMR) of left atrial scar in patients with atrial fibrillation (AF) has recently emerged as a promising technique to stratify patients, to guide ablation therapy and to predict treatment success. This requires a segmentation of the high intensity scar tissue and also a segmentation of the left atrium (LA) anatomy, the latter usually being derived from a separate bright-blood acquisition. Performing both segmentations automatically from a single 3D LGE CMR acquisition would eliminate the need for an additional acquisition and avoid subsequent registration issues. In this paper, we propose a joint segmentation method based on multiview two-task (MVTT) recursive attention model working directly on 3D LGE CMR images to segment the LA (and proximal pulmonary veins) and to delineate the scar on the same dataset. Using our MVTT recursive attention model, both the LA anatomy and scar can be segmented accurately (mean Dice score of 93% for the LA anatomy and 87% for the scar segmentations) and efficiently (~0.27 seconds to simultaneously segment the LA anatomy and scars directly from the 3D LGE CMR dataset with 60-68 2D slices). Compared to conventional unsupervised learning and other state-of-the-art deep learning based methods, the proposed MVTT model achieved excellent results, leading to an automatic generation of a patient-specific anatomical model combined with scar segmentation for patients in AF.
Automatic Brain Tumour Segmentation and Biophysics-Guided Survival Prediction
Nov 19, 2019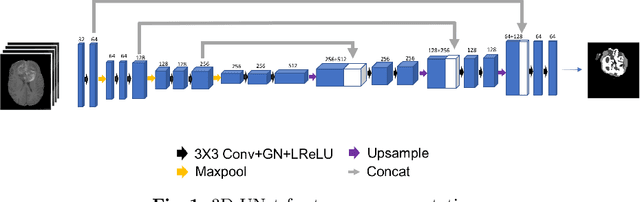
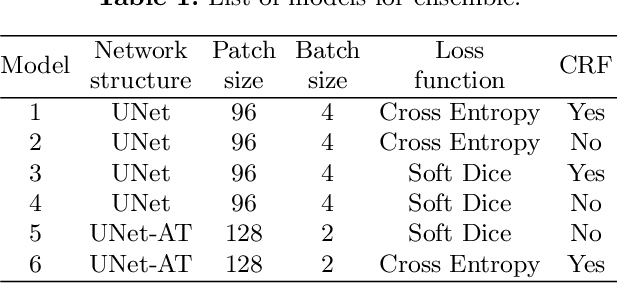
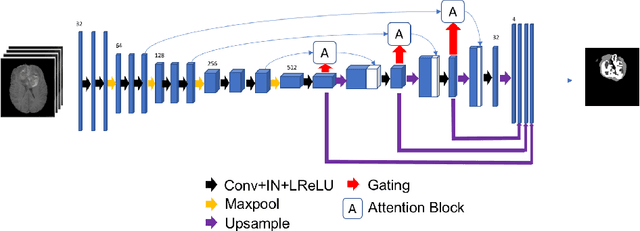
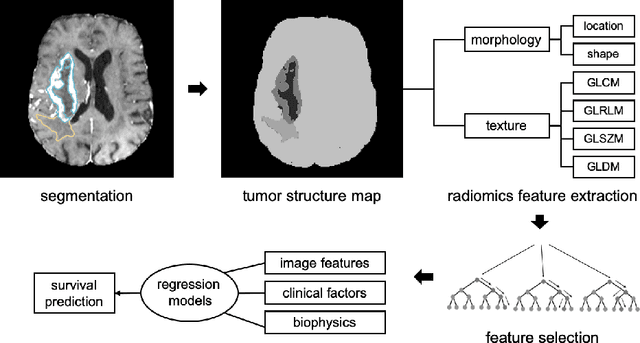
Abstract:Gliomas are the most common malignant brain tumourswith intrinsic heterogeneity. Accurate segmentation of gliomas and theirsub-regions on multi-parametric magnetic resonance images (mpMRI)is of great clinical importance, which defines tumour size, shape andappearance and provides abundant information for preoperative diag-nosis, treatment planning and survival prediction. Recent developmentson deep learning have significantly improved the performance of auto-mated medical image segmentation. In this paper, we compare severalstate-of-the-art convolutional neural network models for brain tumourimage segmentation. Based on the ensembled segmentation, we presenta biophysics-guided prognostic model for patient overall survival predic-tion which outperforms a data-driven radiomics approach. Our methodwon the second place of the MICCAI 2019 BraTS Challenge for theoverall survival prediction.
Transfer Learning from Partial Annotations for Whole Brain Segmentation
Aug 28, 2019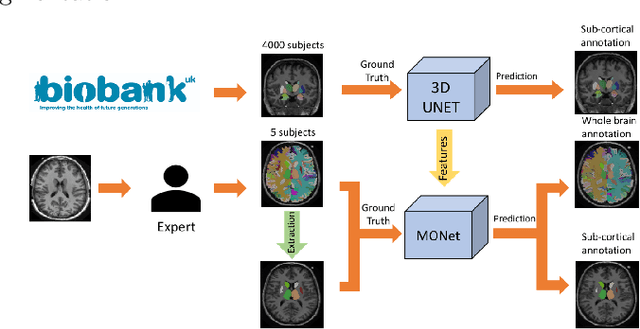

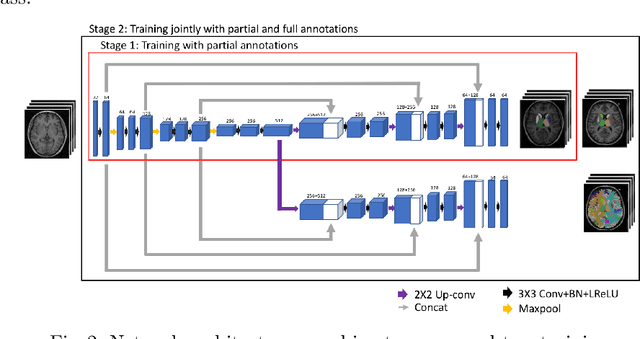
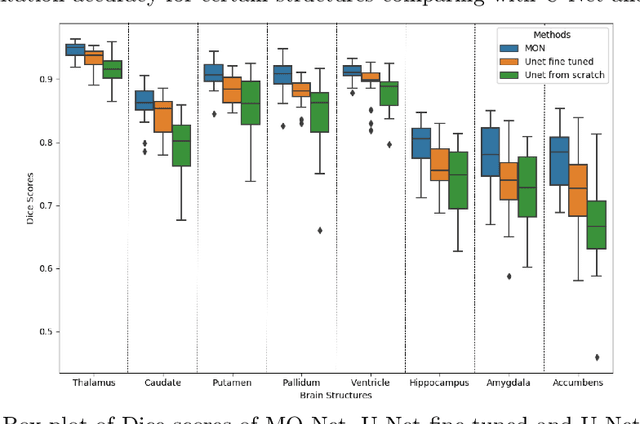
Abstract:Brain MR image segmentation is a key task in neuroimaging studies. It is commonly conducted using standard computational tools, such as FSL, SPM, multi-atlas segmentation etc, which are often registration-based and suffer from expensive computation cost. Recently, there is an increased interest using deep neural networks for brain image segmentation, which have demonstrated advantages in both speed and performance. However, neural networks-based approaches normally require a large amount of manual annotations for optimising the massive amount of network parameters. For 3D networks used in volumetric image segmentation, this has become a particular challenge, as a 3D network consists of many more parameters compared to its 2D counterpart. Manual annotation of 3D brain images is extremely time-consuming and requires extensive involvement of trained experts. To address the challenge with limited manual annotations, here we propose a novel multi-task learning framework for brain image segmentation, which utilises a large amount of automatically generated partial annotations together with a small set of manually created full annotations for network training. Our method yields a high performance comparable to state-of-the-art methods for whole brain segmentation.
SAPSAM - Sparsely Annotated Pathological Sign Activation Maps - A novel approach to train Convolutional Neural Networks on lung CT scans using binary labels only
Feb 06, 2019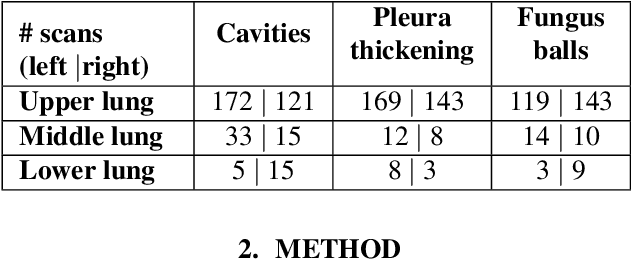
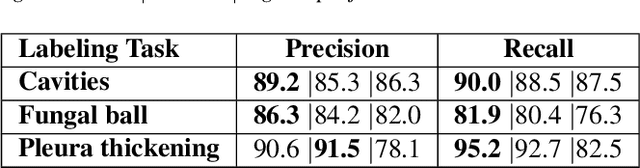
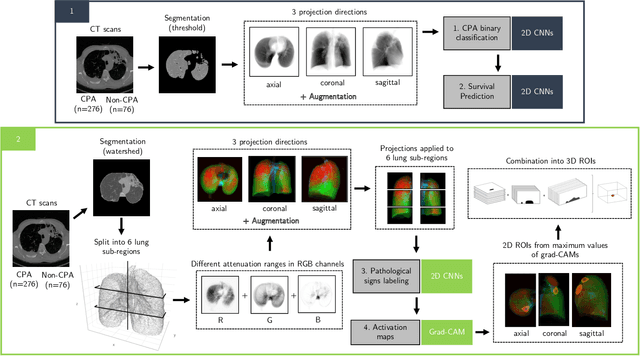

Abstract:Chronic Pulmonary Aspergillosis (CPA) is a complex lung disease caused by infection with Aspergillus. Computed tomography (CT) images are frequently requested in patients with suspected and established disease, but the radiological signs on CT are difficult to quantify making accurate follow-up challenging. We propose a novel method to train Convolutional Neural Networks using only regional labels on the presence of pathological signs, to not only detect CPA, but also spatially localize pathological signs. We use average intensity projections within different ranges of Hounsfield-unit (HU) values, transforming input 3D CT scans into 2D RGB-like images. CNN architectures are trained for hierarchical tasks, leading to precise activation maps of pathological patterns. Results on a cohort of 352 subjects demonstrate high classification accuracy, localization precision and predictive power of 2 year survival. Such tool opens the way to CPA patient stratification and quantitative follow-up of CPA pathological signs, for patients under drug therapy.
* Accepted paper for ISBI2019
Multiview Two-Task Recursive Attention Model for Left Atrium and Atrial Scars Segmentation
Jun 12, 2018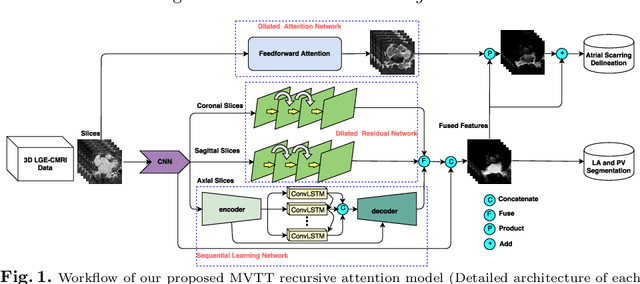
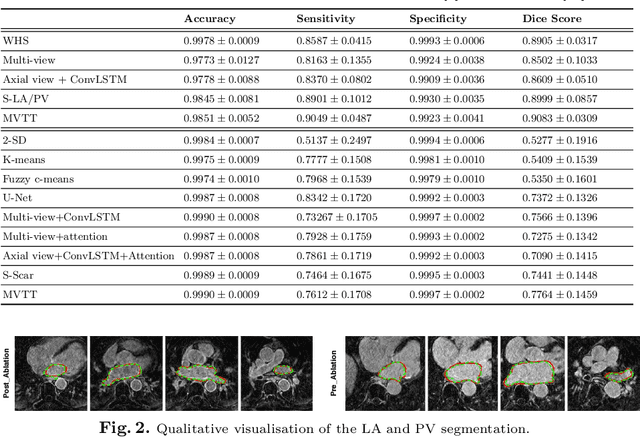
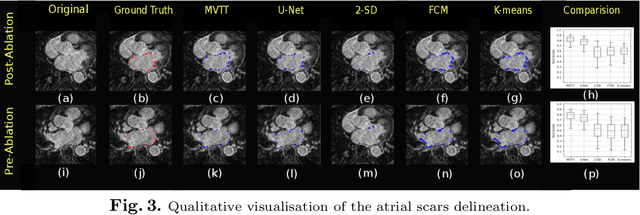
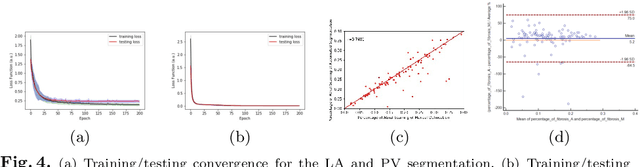
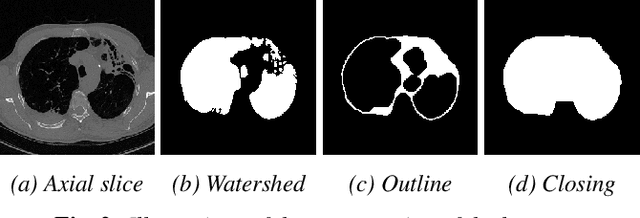
Abstract:Late Gadolinium Enhanced Cardiac MRI (LGE-CMRI) for detecting atrial scars in atrial fibrillation (AF) patients has recently emerged as a promising technique to stratify patients, guide ablation therapy and predict treatment success. Visualisation and quantification of scar tissues require a segmentation of both the left atrium (LA) and the high intensity scar regions from LGE-CMRI images. These two segmentation tasks are challenging due to the cancelling of healthy tissue signal, low signal-to-noise ratio and often limited image quality in these patients. Most approaches require manual supervision and/or a second bright-blood MRI acquisition for anatomical segmentation. Segmenting both the LA anatomy and the scar tissues automatically from a single LGE-CMRI acquisition is highly in demand. In this study, we proposed a novel fully automated multiview two-task (MVTT) recursive attention model working directly on LGE-CMRI images that combines a sequential learning and a dilated residual learning to segment the LA (including attached pulmonary veins) and delineate the atrial scars simultaneously via an innovative attention model. Compared to other state-of-the-art methods, the proposed MVTT achieves compelling improvement, enabling to generate a patient-specific anatomical and atrial scar assessment model.
Compressed Sensing with off-axis frequency-shifting holography
Apr 29, 2010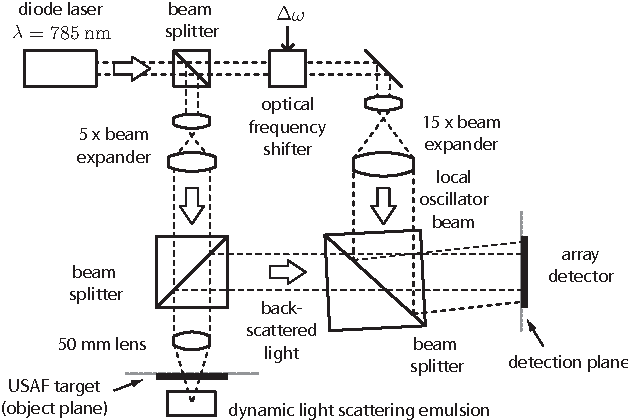
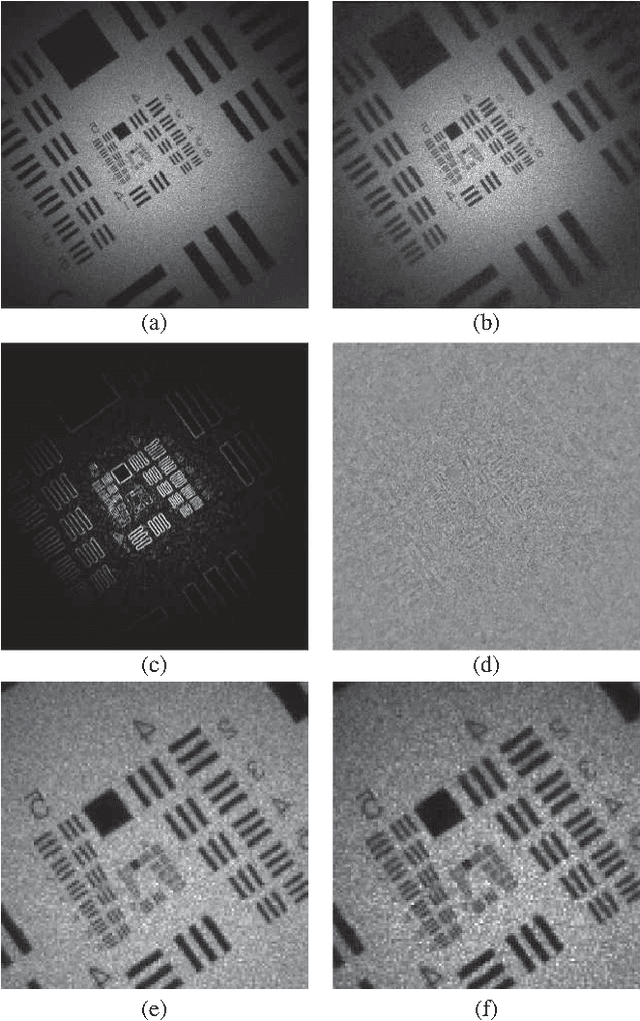
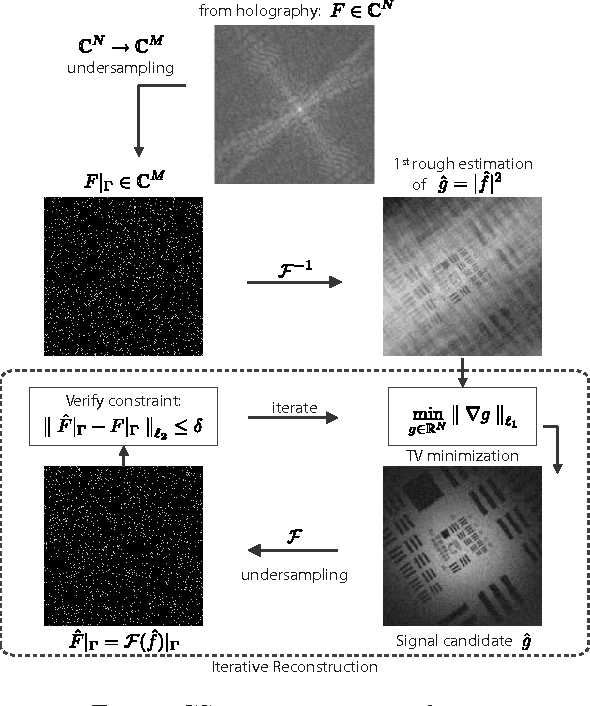

Abstract:This work reveals an experimental microscopy acquisition scheme successfully combining Compressed Sensing (CS) and digital holography in off-axis and frequency-shifting conditions. CS is a recent data acquisition theory involving signal reconstruction from randomly undersampled measurements, exploiting the fact that most images present some compact structure and redundancy. We propose a genuine CS-based imaging scheme for sparse gradient images, acquiring a diffraction map of the optical field with holographic microscopy and recovering the signal from as little as 7% of random measurements. We report experimental results demonstrating how CS can lead to an elegant and effective way to reconstruct images, opening the door for new microscopy applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge