Daniel Ward
The Hyperfitting Phenomenon: Sharpening and Stabilizing LLMs for Open-Ended Text Generation
Dec 05, 2024



Abstract:This paper introduces the counter-intuitive generalization results of overfitting pre-trained large language models (LLMs) on very small datasets. In the setting of open-ended text generation, it is well-documented that LLMs tend to generate repetitive and dull sequences, a phenomenon that is especially apparent when generating using greedy decoding. This issue persists even with state-of-the-art LLMs containing billions of parameters, trained via next-token prediction on large datasets. We find that by further fine-tuning these models to achieve a near-zero training loss on a small set of samples -- a process we refer to as hyperfitting -- the long-sequence generative capabilities are greatly enhanced. Greedy decoding with these Hyperfitted models even outperform Top-P sampling over long-sequences, both in terms of diversity and human preferences. This phenomenon extends to LLMs of various sizes, different domains, and even autoregressive image generation. We further find this phenomena to be distinctly different from that of Grokking and double descent. Surprisingly, our experiments indicate that hyperfitted models rarely fall into repeating sequences they were trained on, and even explicitly blocking these sequences results in high-quality output. All hyperfitted models produce extremely low-entropy predictions, often allocating nearly all probability to a single token.
SoftCVI: contrastive variational inference with self-generated soft labels
Jul 22, 2024Abstract:Estimating a distribution given access to its unnormalized density is pivotal in Bayesian inference, where the posterior is generally known only up to an unknown normalizing constant. Variational inference and Markov chain Monte Carlo methods are the predominant tools for this task; however, both methods are often challenging to apply reliably, particularly when the posterior has complex geometry. Here, we introduce Soft Contrastive Variational Inference (SoftCVI), which allows a family of variational objectives to be derived through a contrastive estimation framework. These objectives have zero variance gradient when the variational approximation is exact, without the need for specialized gradient estimators. The approach involves parameterizing a classifier in terms of the variational distribution, which allows the inference task to be reframed as a contrastive estimation problem, aiming to identify a single true posterior sample among a set of samples. Despite this framing, we do not require positive or negative samples, but rather learn by sampling the variational distribution and computing ground truth soft classification labels from the unnormalized posterior itself. We empirically investigate the performance on a variety of Bayesian inference tasks, using both using both simple (e.g. normal) and expressive (normalizing flow) variational distributions. We find that SoftCVI objectives often outperform other commonly used variational objectives.
Robust Neural Posterior Estimation and Statistical Model Criticism
Oct 12, 2022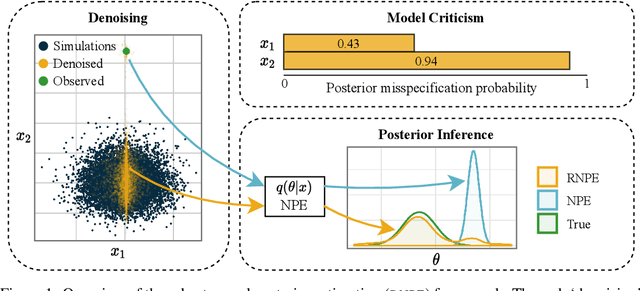
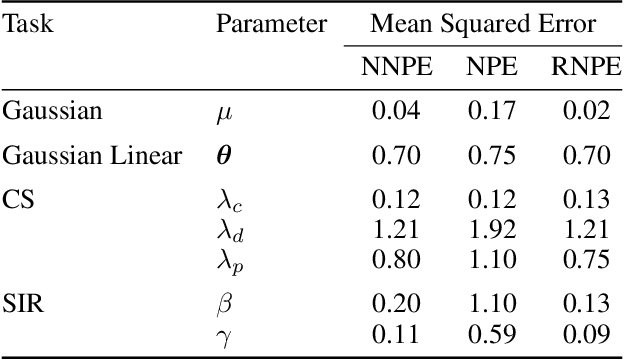
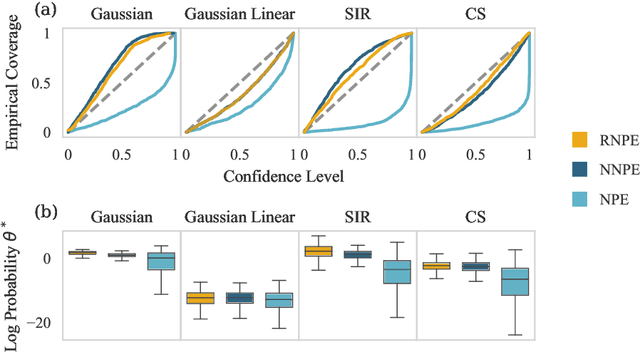
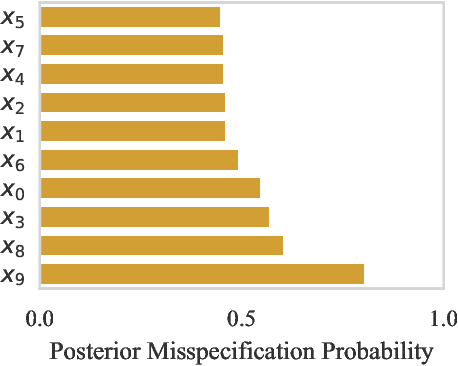
Abstract:Computer simulations have proven a valuable tool for understanding complex phenomena across the sciences. However, the utility of simulators for modelling and forecasting purposes is often restricted by low data quality, as well as practical limits to model fidelity. In order to circumvent these difficulties, we argue that modellers must treat simulators as idealistic representations of the true data generating process, and consequently should thoughtfully consider the risk of model misspecification. In this work we revisit neural posterior estimation (NPE), a class of algorithms that enable black-box parameter inference in simulation models, and consider the implication of a simulation-to-reality gap. While recent works have demonstrated reliable performance of these methods, the analyses have been performed using synthetic data generated by the simulator model itself, and have therefore only addressed the well-specified case. In this paper, we find that the presence of misspecification, in contrast, leads to unreliable inference when NPE is used naively. As a remedy we argue that principled scientific inquiry with simulators should incorporate a model criticism component, to facilitate interpretable identification of misspecification and a robust inference component, to fit 'wrong but useful' models. We propose robust neural posterior estimation (RNPE), an extension of NPE to simultaneously achieve both these aims, through explicitly modelling the discrepancies between simulations and the observed data. We assess the approach on a range of artificially misspecified examples, and find RNPE performs well across the tasks, whereas naively using NPE leads to misleading and erratic posteriors.
Investigating the Impact of Model Misspecification in Neural Simulation-based Inference
Sep 05, 2022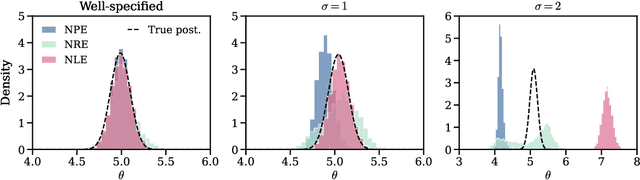
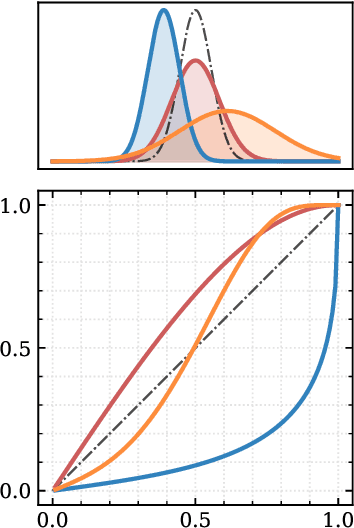
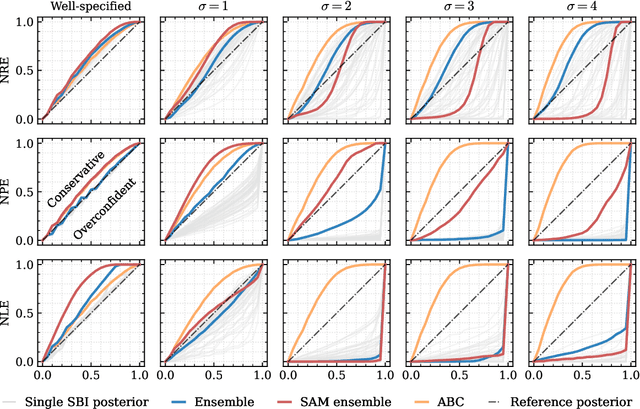
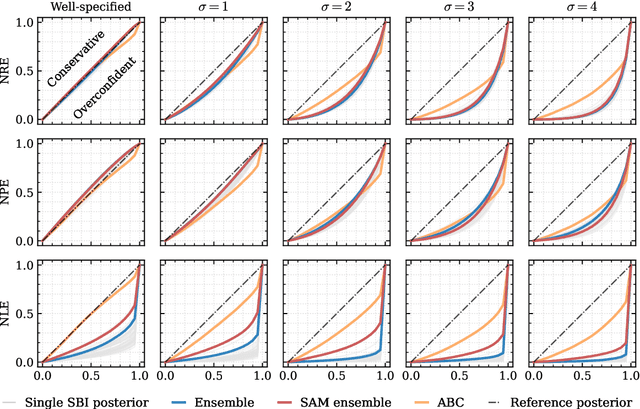
Abstract:Aided by advances in neural density estimation, considerable progress has been made in recent years towards a suite of simulation-based inference (SBI) methods capable of performing flexible, black-box, approximate Bayesian inference for stochastic simulation models. While it has been demonstrated that neural SBI methods can provide accurate posterior approximations, the simulation studies establishing these results have considered only well-specified problems -- that is, where the model and the data generating process coincide exactly. However, the behaviour of such algorithms in the case of model misspecification has received little attention. In this work, we provide the first comprehensive study of the behaviour of neural SBI algorithms in the presence of various forms of model misspecification. We find that misspecification can have a profoundly deleterious effect on performance. Some mitigation strategies are explored, but no approach tested prevents failure in all cases. We conclude that new approaches are required to address model misspecification if neural SBI algorithms are to be relied upon to derive accurate scientific conclusions.
Learning Arbitrary-Goal Fabric Folding with One Hour of Real Robot Experience
Oct 07, 2020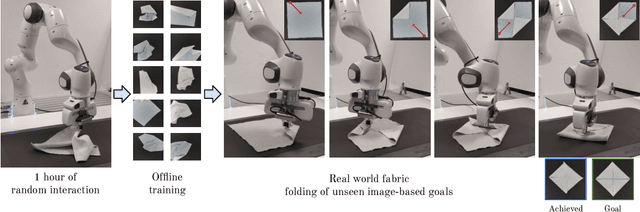

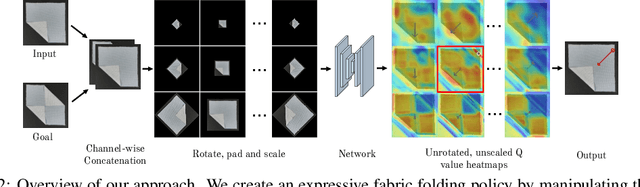
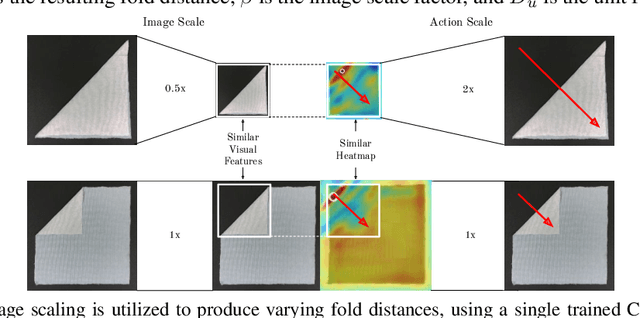
Abstract:Manipulating deformable objects, such as fabric, is a long standing problem in robotics, with state estimation and control posing a significant challenge for traditional methods. In this paper, we show that it is possible to learn fabric folding skills in only an hour of self-supervised real robot experience, without human supervision or simulation. Our approach relies on fully convolutional networks and the manipulation of visual inputs to exploit learned features, allowing us to create an expressive goal-conditioned pick and place policy that can be trained efficiently with real world robot data only. Folding skills are learned with only a sparse reward function and thus do not require reward function engineering, merely an image of the goal configuration. We demonstrate our method on a set of towel-folding tasks, and show that our approach is able to discover sequential folding strategies, purely from trial-and-error. We achieve state-of-the-art results without the need for demonstrations or simulation, used in prior approaches. Videos available at: https://sites.google.com/view/learningtofold
Temporally Coherent Embeddings for Self-Supervised Video Representation Learning
May 01, 2020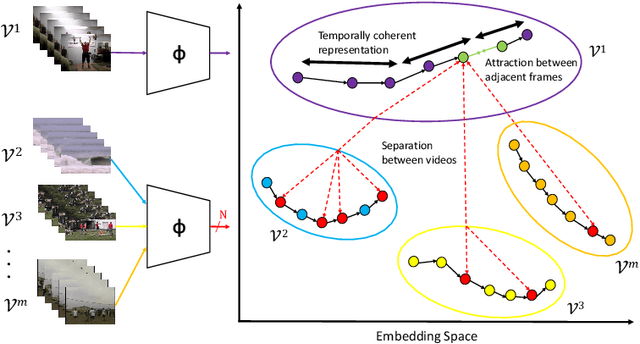
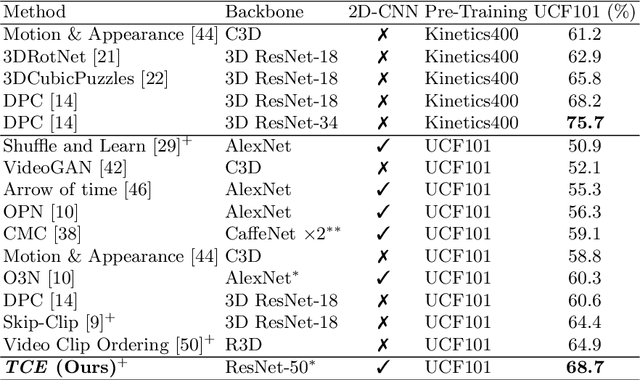
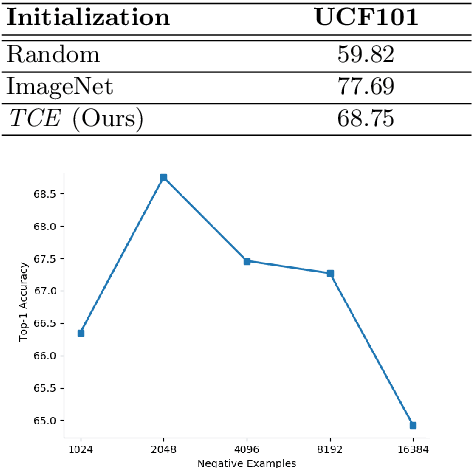
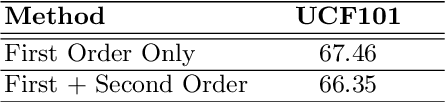
Abstract:This paper presents TCE: Temporally Coherent Embeddings for self-supervised video representation learning. The proposed method exploits inherent structure of unlabeled video data to explicitly enforce temporal coherency in the embedding space, rather than indirectly learning it through ranking or predictive pretext tasks. In the same way that high-level visual information in the world changes smoothly, we believe that nearby frames in learned representations should demonstrate similar properties. Using this assumption, we train the TCE model to encode videos such that adjacent frames exist close to each other and videos are separated from one another. Using TCE we learn robust representations from large quantities of unlabeled video data. We evaluate our self-supervised trained TCE model by adding a classification layer and finetuning the learned representation on the downstream task of video action recognition on the UCF101 dataset. We obtain 67.01% accuracy and outperform the state-of-the-art self-supervised methods trained on UCF101 despite using a significantly smaller dataset for pre-training. Notably, we demonstrate results competitive with more complex 3D-CNN based networks while training with a 2D-CNN network backbone on action recognition tasks. Our training code and pretrained models are available at https://github.com/csiro-robotics/TCE
Scalable learning for bridging the species gap in image-based plant phenotyping
Apr 24, 2020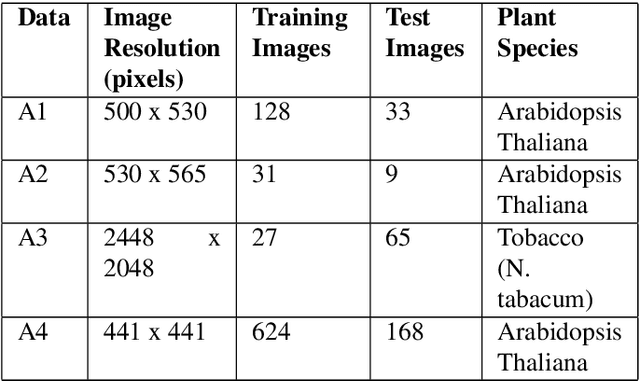
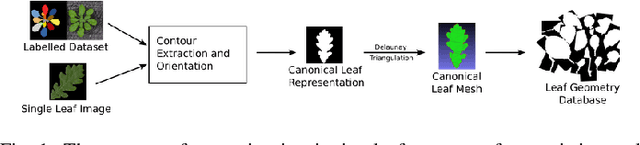
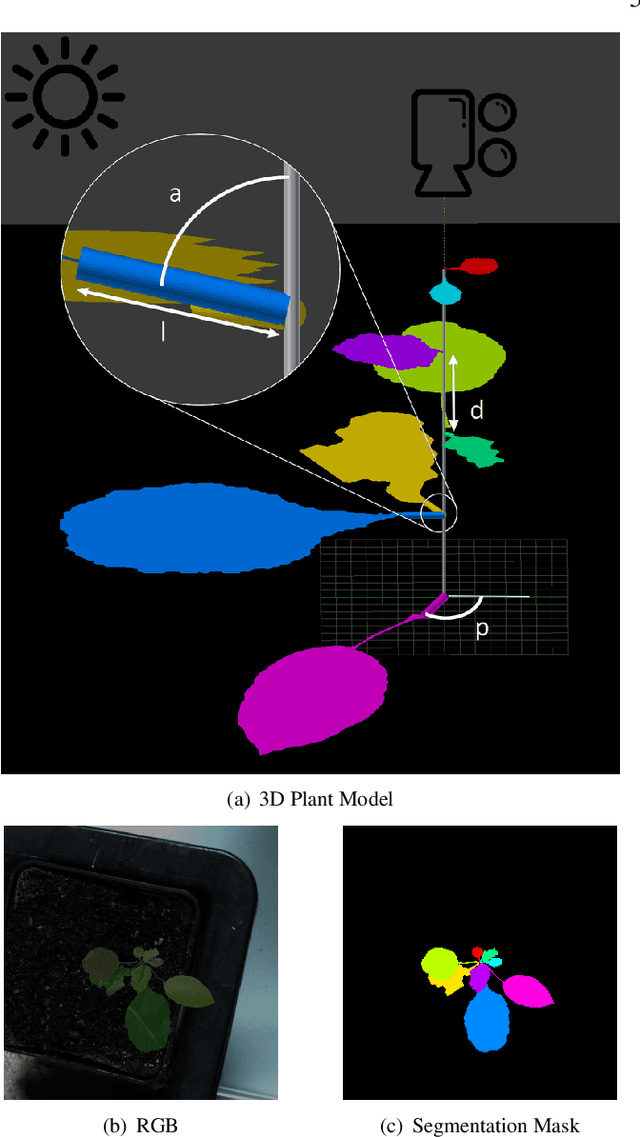
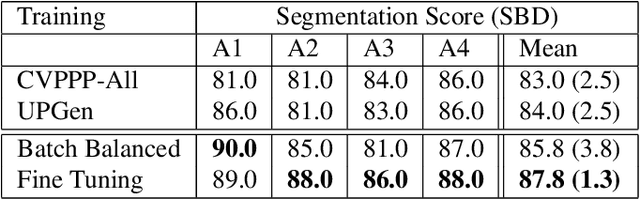
Abstract:The traditional paradigm of applying deep learning -- collect, annotate and train on data -- is not applicable to image-based plant phenotyping as almost 400,000 different plant species exists. Data costs include growing physical samples, imaging and labelling them. Model performance is impacted by the species gap between the domain of each plant species, it is not generalisable and may not transfer to unseen plant species. In this paper, we investigate the use of synthetic data for leaf instance segmentation. We study multiple synthetic data training regimes using Mask-RCNN when few or no annotated real data is available. We also present UPGen: a Universal Plant Generator for bridging the species gap. UPGen leverages domain randomisation to produce widely distributed data samples and models stochastic biological variation. Our methods outperform standard practices, such as transfer learning from publicly available plant data, by 26.6% and 51.46% on two unseen plant species respectively. We benchmark UPGen by competing in the CVPPP Leaf Segmentation Challenge and set a new state-of-the-art, a mean of 88% across A1-4 test datasets. This study is applicable to use of synthetic data for automating the measurement of phenotypic traits. Our synthetic dataset and pretrained model are available at https://csiro-robotics.github.io/UPGen_Webpage/.
Deep Leaf Segmentation Using Synthetic Data
Aug 17, 2018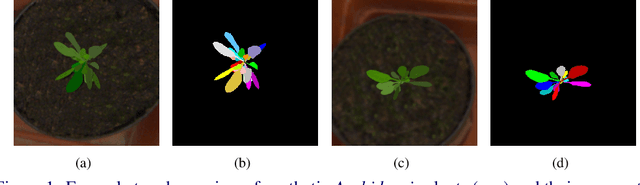
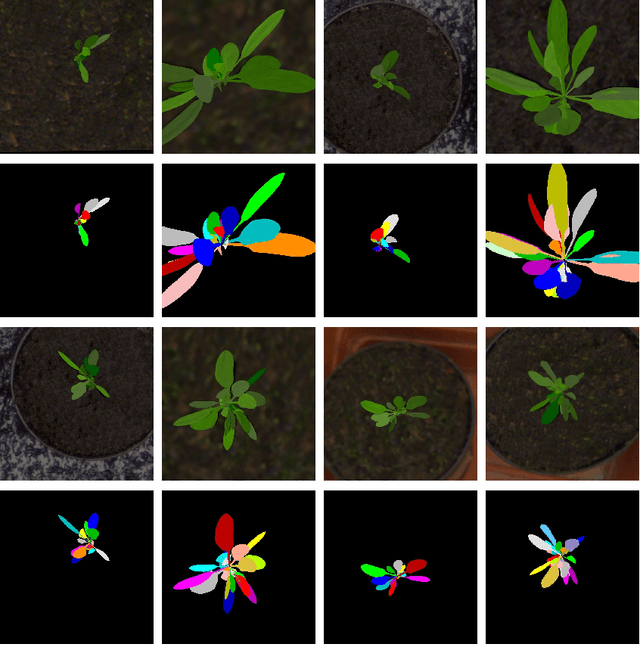
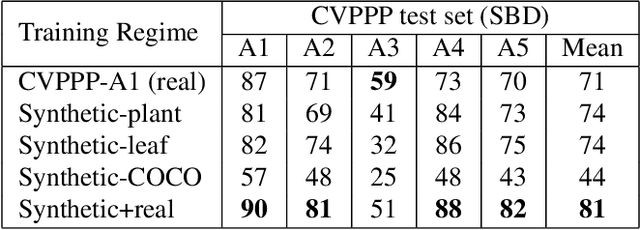

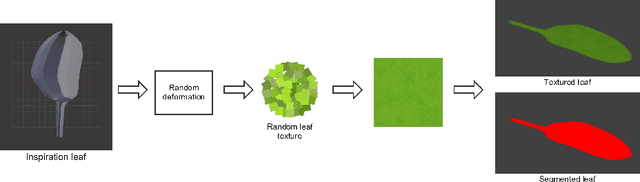
Abstract:Automated segmentation of individual leaves of a plant in an image is a prerequisite to measure more complex phenotypic traits in high-throughput phenotyping. Applying state-of-the-art machine learning approaches to tackle leaf instance segmentation requires a large amount of manually annotated training data. Currently, the benchmark datasets for leaf segmentation contain only a few hundred labeled training images. In this paper, we propose a framework for leaf instance segmentation by augmenting real plant datasets with generated synthetic images of plants inspired by domain randomisation. We train a state-of-the-art deep learning segmentation architecture (Mask-RCNN) with a combination of real and synthetic images of Arabidopsis plants. Our proposed approach achieves 90% leaf segmentation score on the A1 test set outperforming the-state-of-the-art approaches for the CVPPP Leaf Segmentation Challenge (LSC). Our approach also achieves 81% mean performance over all five test datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge