Brian Dromey
A multi-centre, multi-device benchmark dataset for landmark-based comprehensive fetal biometry
Dec 19, 2025Abstract:Accurate fetal growth assessment from ultrasound (US) relies on precise biometry measured by manually identifying anatomical landmarks in standard planes. Manual landmarking is time-consuming, operator-dependent, and sensitive to variability across scanners and sites, limiting the reproducibility of automated approaches. There is a need for multi-source annotated datasets to develop artificial intelligence-assisted fetal growth assessment methods. To address this bottleneck, we present an open, multi-centre, multi-device benchmark dataset of fetal US images with expert anatomical landmark annotations for clinically used fetal biometric measurements. These measurements include head bi-parietal and occipito-frontal diameters, abdominal transverse and antero-posterior diameters, and femoral length. The dataset comprises 4,513 de-identified US images from 1,904 subjects acquired at three clinical sites using seven different US devices. We provide standardised, subject-disjoint train/test splits, evaluation code, and baseline results to enable fair and reproducible comparison of methods. Using an automatic biometry model, we quantify domain shift and demonstrate that training and evaluation confined to a single centre substantially overestimate performance relative to multi-centre testing. To the best of our knowledge, this is the first publicly available multi-centre, multi-device, landmark-annotated dataset that covers all primary fetal biometry measures, providing a robust benchmark for domain adaptation and multi-centre generalisation in fetal biometry and enabling more reliable AI-assisted fetal growth assessment across centres. All data, annotations, training code, and evaluation pipelines are made publicly available.
HUP-3D: A 3D multi-view synthetic dataset for assisted-egocentric hand-ultrasound pose estimation
Jul 12, 2024



Abstract:We present HUP-3D, a 3D multi-view multi-modal synthetic dataset for hand-ultrasound (US) probe pose estimation in the context of obstetric ultrasound. Egocentric markerless 3D joint pose estimation has potential applications in mixed reality based medical education. The ability to understand hand and probe movements programmatically opens the door to tailored guidance and mentoring applications. Our dataset consists of over 31k sets of RGB, depth and segmentation mask frames, including pose related ground truth data, with a strong emphasis on image diversity and complexity. Adopting a camera viewpoint-based sphere concept allows us to capture a variety of views and generate multiple hand grasp poses using a pre-trained network. Additionally, our approach includes a software-based image rendering concept, enhancing diversity with various hand and arm textures, lighting conditions, and background images. Furthermore, we validated our proposed dataset with state-of-the-art learning models and we obtained the lowest hand-object keypoint errors. The dataset and other details are provided with the supplementary material. The source code of our grasp generation and rendering pipeline will be made publicly available.
Learning ultrasound plane pose regression: assessing generalized pose coordinates in the fetal brain
Jan 19, 2023



Abstract:In obstetric ultrasound (US) scanning, the learner's ability to mentally build a three-dimensional (3D) map of the fetus from a two-dimensional (2D) US image represents a significant challenge in skill acquisition. We aim to build a US plane localization system for 3D visualization, training, and guidance without integrating additional sensors. This work builds on top of our previous work, which predicts the six-dimensional (6D) pose of arbitrarily-oriented US planes slicing the fetal brain with respect to a normalized reference frame using a convolutional neural network (CNN) regression network. Here, we analyze in detail the assumptions of the normalized fetal brain reference frame and quantify its accuracy with respect to the acquisition of transventricular (TV) standard plane (SP) for fetal biometry. We investigate the impact of registration quality in the training and testing data and its subsequent effect on trained models. Finally, we introduce data augmentations and larger training sets that improve the results of our previous work, achieving median errors of 3.53 mm and 6.42 degrees for translation and rotation, respectively.
BiometryNet: Landmark-based Fetal Biometry Estimation from Standard Ultrasound Planes
Jun 29, 2022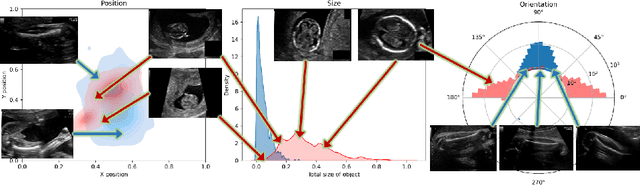
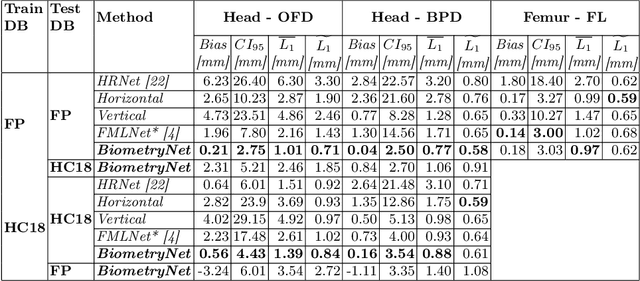
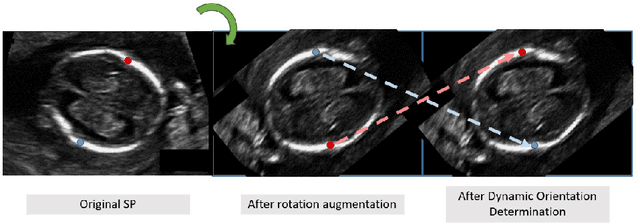
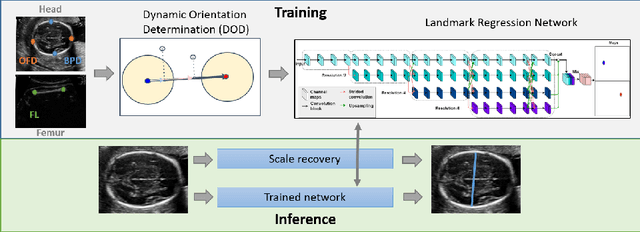
Abstract:Fetal growth assessment from ultrasound is based on a few biometric measurements that are performed manually and assessed relative to the expected gestational age. Reliable biometry estimation depends on the precise detection of landmarks in standard ultrasound planes. Manual annotation can be time-consuming and operator dependent task, and may results in high measurements variability. Existing methods for automatic fetal biometry rely on initial automatic fetal structure segmentation followed by geometric landmark detection. However, segmentation annotations are time-consuming and may be inaccurate, and landmark detection requires developing measurement-specific geometric methods. This paper describes BiometryNet, an end-to-end landmark regression framework for fetal biometry estimation that overcomes these limitations. It includes a novel Dynamic Orientation Determination (DOD) method for enforcing measurement-specific orientation consistency during network training. DOD reduces variabilities in network training, increases landmark localization accuracy, thus yields accurate and robust biometric measurements. To validate our method, we assembled a dataset of 3,398 ultrasound images from 1,829 subjects acquired in three clinical sites with seven different ultrasound devices. Comparison and cross-validation of three different biometric measurements on two independent datasets shows that BiometryNet is robust and yields accurate measurements whose errors are lower than the clinically permissible errors, outperforming other existing automated biometry estimation methods. Code is available at https://github.com/netanellavisdris/fetalbiometry.
AutoFB: Automating Fetal Biometry Estimation from Standard Ultrasound Planes
Jul 12, 2021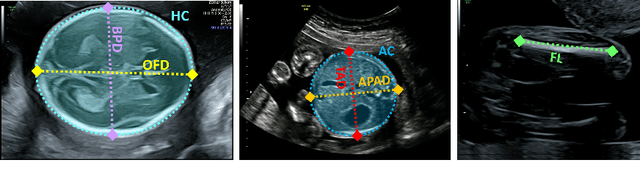
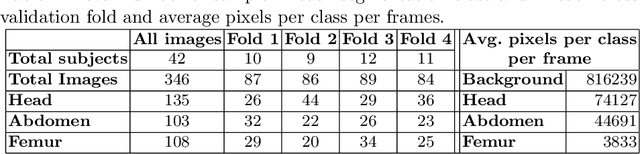
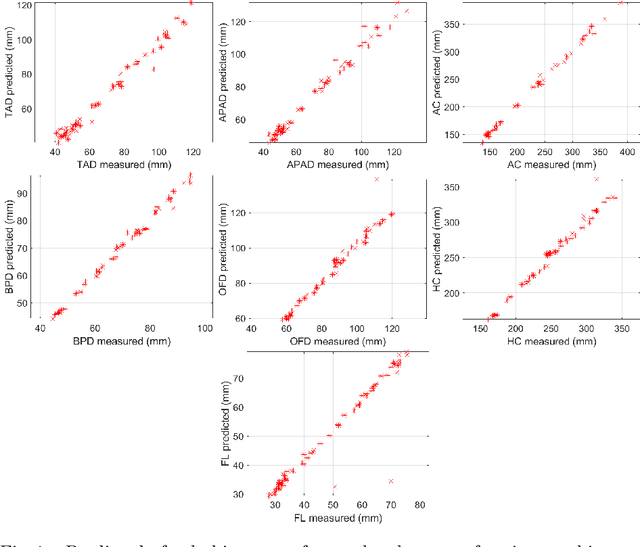
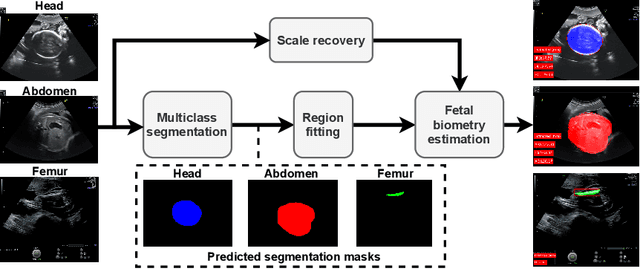
Abstract:During pregnancy, ultrasound examination in the second trimester can assess fetal size according to standardized charts. To achieve a reproducible and accurate measurement, a sonographer needs to identify three standard 2D planes of the fetal anatomy (head, abdomen, femur) and manually mark the key anatomical landmarks on the image for accurate biometry and fetal weight estimation. This can be a time-consuming operator-dependent task, especially for a trainee sonographer. Computer-assisted techniques can help in automating the fetal biometry computation process. In this paper, we present a unified automated framework for estimating all measurements needed for the fetal weight assessment. The proposed framework semantically segments the key fetal anatomies using state-of-the-art segmentation models, followed by region fitting and scale recovery for the biometry estimation. We present an ablation study of segmentation algorithms to show their robustness through 4-fold cross-validation on a dataset of 349 ultrasound standard plane images from 42 pregnancies. Moreover, we show that the network with the best segmentation performance tends to be more accurate for biometry estimation. Furthermore, we demonstrate that the error between clinically measured and predicted fetal biometry is lower than the permissible error during routine clinical measurements.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge