Bishesh Khanal
NAAMII, Nepal, Tangible Careers
Benchmarking Encoder-Decoder Architectures for Biplanar X-ray to 3D Shape Reconstruction
Sep 26, 2023



Abstract:Various deep learning models have been proposed for 3D bone shape reconstruction from two orthogonal (biplanar) X-ray images. However, it is unclear how these models compare against each other since they are evaluated on different anatomy, cohort and (often privately held) datasets. Moreover, the impact of the commonly optimized image-based segmentation metrics such as dice score on the estimation of clinical parameters relevant in 2D-3D bone shape reconstruction is not well known. To move closer toward clinical translation, we propose a benchmarking framework that evaluates tasks relevant to real-world clinical scenarios, including reconstruction of fractured bones, bones with implants, robustness to population shift, and error in estimating clinical parameters. Our open-source platform provides reference implementations of 8 models (many of whose implementations were not publicly available), APIs to easily collect and preprocess 6 public datasets, and the implementation of automatic clinical parameter and landmark extraction methods. We present an extensive evaluation of 8 2D-3D models on equal footing using 6 public datasets comprising images for four different anatomies. Our results show that attention-based methods that capture global spatial relationships tend to perform better across all anatomies and datasets; performance on clinically relevant subgroups may be overestimated without disaggregated reporting; ribs are substantially more difficult to reconstruct compared to femur, hip and spine; and the dice score improvement does not always bring a corresponding improvement in the automatic estimation of clinically relevant parameters.
Synthetic Boost: Leveraging Synthetic Data for Enhanced Vision-Language Segmentation in Echocardiography
Sep 22, 2023Abstract:Accurate segmentation is essential for echocardiography-based assessment of cardiovascular diseases (CVDs). However, the variability among sonographers and the inherent challenges of ultrasound images hinder precise segmentation. By leveraging the joint representation of image and text modalities, Vision-Language Segmentation Models (VLSMs) can incorporate rich contextual information, potentially aiding in accurate and explainable segmentation. However, the lack of readily available data in echocardiography hampers the training of VLSMs. In this study, we explore using synthetic datasets from Semantic Diffusion Models (SDMs) to enhance VLSMs for echocardiography segmentation. We evaluate results for two popular VLSMs (CLIPSeg and CRIS) using seven different kinds of language prompts derived from several attributes, automatically extracted from echocardiography images, segmentation masks, and their metadata. Our results show improved metrics and faster convergence when pretraining VLSMs on SDM-generated synthetic images before finetuning on real images. The code, configs, and prompts are available at https://github.com/naamiinepal/synthetic-boost.
Exploring Transfer Learning in Medical Image Segmentation using Vision-Language Models
Aug 15, 2023

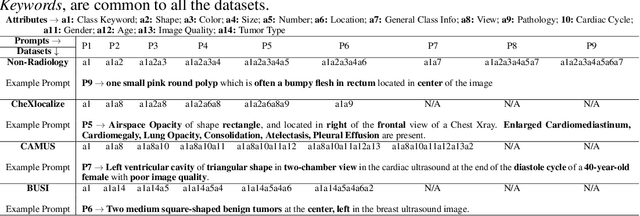

Abstract:Medical Image Segmentation is crucial in various clinical applications within the medical domain. While state-of-the-art segmentation models have proven effective, integrating textual guidance to enhance visual features for this task remains an area with limited progress. Existing segmentation models that utilize textual guidance are primarily trained on open-domain images, raising concerns about their direct applicability in the medical domain without manual intervention or fine-tuning. To address these challenges, we propose using multimodal vision-language models for capturing semantic information from image descriptions and images, enabling the segmentation of diverse medical images. This study comprehensively evaluates existing vision language models across multiple datasets to assess their transferability from the open domain to the medical field. Furthermore, we introduce variations of image descriptions for previously unseen images in the dataset, revealing notable variations in model performance based on the generated prompts. Our findings highlight the distribution shift between the open-domain images and the medical domain and show that the segmentation models trained on open-domain images are not directly transferrable to the medical field. But their performance can be increased by finetuning them in the medical datasets. We report the zero-shot and finetuned segmentation performance of 4 Vision Language Models (VLMs) on 11 medical datasets using 9 types of prompts derived from 14 attributes.
Improving Medical Image Classification in Noisy Labels Using Only Self-supervised Pretraining
Aug 08, 2023



Abstract:Noisy labels hurt deep learning-based supervised image classification performance as the models may overfit the noise and learn corrupted feature extractors. For natural image classification training with noisy labeled data, model initialization with contrastive self-supervised pretrained weights has shown to reduce feature corruption and improve classification performance. However, no works have explored: i) how other self-supervised approaches, such as pretext task-based pretraining, impact the learning with noisy label, and ii) any self-supervised pretraining methods alone for medical images in noisy label settings. Medical images often feature smaller datasets and subtle inter class variations, requiring human expertise to ensure correct classification. Thus, it is not clear if the methods improving learning with noisy labels in natural image datasets such as CIFAR would also help with medical images. In this work, we explore contrastive and pretext task-based self-supervised pretraining to initialize the weights of a deep learning classification model for two medical datasets with self-induced noisy labels -- NCT-CRC-HE-100K tissue histological images and COVID-QU-Ex chest X-ray images. Our results show that models initialized with pretrained weights obtained from self-supervised learning can effectively learn better features and improve robustness against noisy labels.
M-VAAL: Multimodal Variational Adversarial Active Learning for Downstream Medical Image Analysis Tasks
Jun 21, 2023Abstract:Acquiring properly annotated data is expensive in the medical field as it requires experts, time-consuming protocols, and rigorous validation. Active learning attempts to minimize the need for large annotated samples by actively sampling the most informative examples for annotation. These examples contribute significantly to improving the performance of supervised machine learning models, and thus, active learning can play an essential role in selecting the most appropriate information in deep learning-based diagnosis, clinical assessments, and treatment planning. Although some existing works have proposed methods for sampling the best examples for annotation in medical image analysis, they are not task-agnostic and do not use multimodal auxiliary information in the sampler, which has the potential to increase robustness. Therefore, in this work, we propose a Multimodal Variational Adversarial Active Learning (M-VAAL) method that uses auxiliary information from additional modalities to enhance the active sampling. We applied our method to two datasets: i) brain tumor segmentation and multi-label classification using the BraTS2018 dataset, and ii) chest X-ray image classification using the COVID-QU-Ex dataset. Our results show a promising direction toward data-efficient learning under limited annotations.
Deep-learning assisted detection and quantification of (oo)cysts of Giardia and Cryptosporidium on smartphone microscopy images
Apr 11, 2023



Abstract:The consumption of microbial-contaminated food and water is responsible for the deaths of millions of people annually. Smartphone-based microscopy systems are portable, low-cost, and more accessible alternatives for the detection of Giardia and Cryptosporidium than traditional brightfield microscopes. However, the images from smartphone microscopes are noisier and require manual cyst identification by trained technicians, usually unavailable in resource-limited settings. Automatic detection of (oo)cysts using deep-learning-based object detection could offer a solution for this limitation. We evaluate the performance of three state-of-the-art object detectors to detect (oo)cysts of Giardia and Cryptosporidium on a custom dataset that includes both smartphone and brightfield microscopic images from vegetable samples. Faster RCNN, RetinaNet, and you only look once (YOLOv8s) deep-learning models were employed to explore their efficacy and limitations. Our results show that while the deep-learning models perform better with the brightfield microscopy image dataset than the smartphone microscopy image dataset, the smartphone microscopy predictions are still comparable to the prediction performance of non-experts.
COVID-19-related Nepali Tweets Classification in a Low Resource Setting
Oct 11, 2022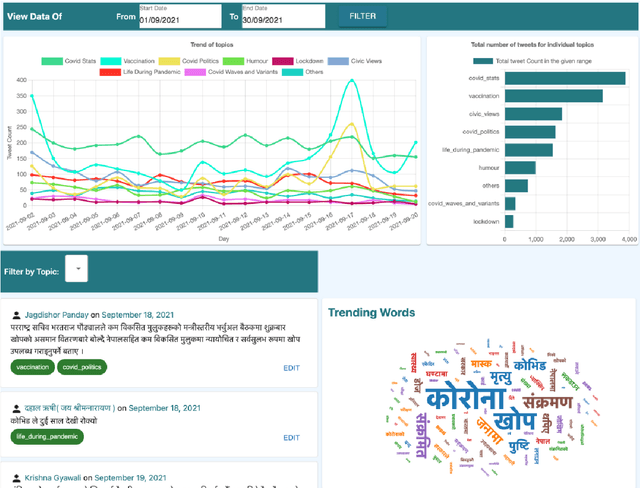
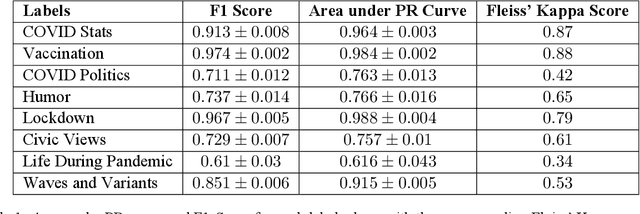
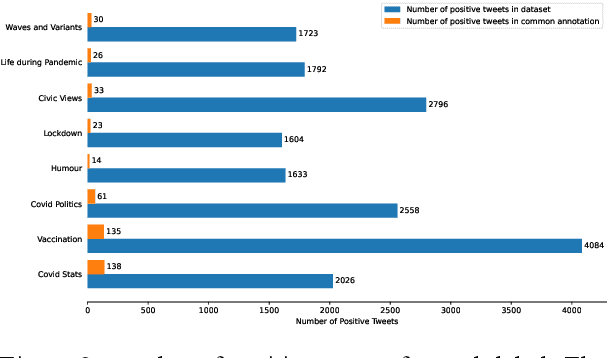
Abstract:Billions of people across the globe have been using social media platforms in their local languages to voice their opinions about the various topics related to the COVID-19 pandemic. Several organizations, including the World Health Organization, have developed automated social media analysis tools that classify COVID-19-related tweets into various topics. However, these tools that help combat the pandemic are limited to very few languages, making several countries unable to take their benefit. While multi-lingual or low-resource language-specific tools are being developed, they still need to expand their coverage, such as for the Nepali language. In this paper, we identify the eight most common COVID-19 discussion topics among the Twitter community using the Nepali language, set up an online platform to automatically gather Nepali tweets containing the COVID-19-related keywords, classify the tweets into the eight topics, and visualize the results across the period in a web-based dashboard. We compare the performance of two state-of-the-art multi-lingual language models for Nepali tweet classification, one generic (mBERT) and the other Nepali language family-specific model (MuRIL). Our results show that the models' relative performance depends on the data size, with MuRIL doing better for a larger dataset. The annotated data, models, and the web-based dashboard are open-sourced at https://github.com/naamiinepal/covid-tweet-classification.
FixMatchSeg: Fixing FixMatch for Semi-Supervised Semantic Segmentation
Aug 02, 2022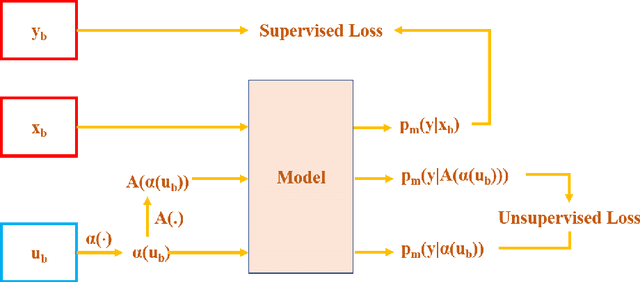
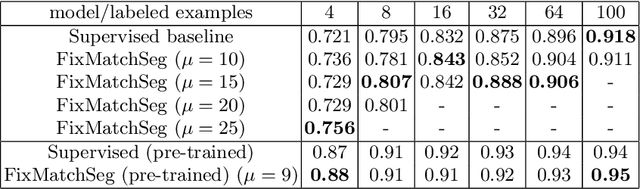
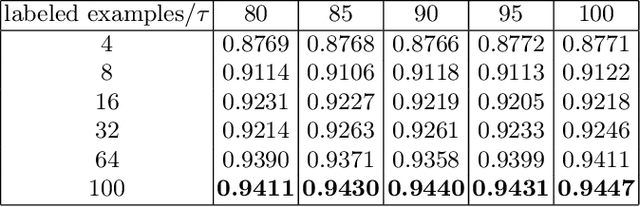
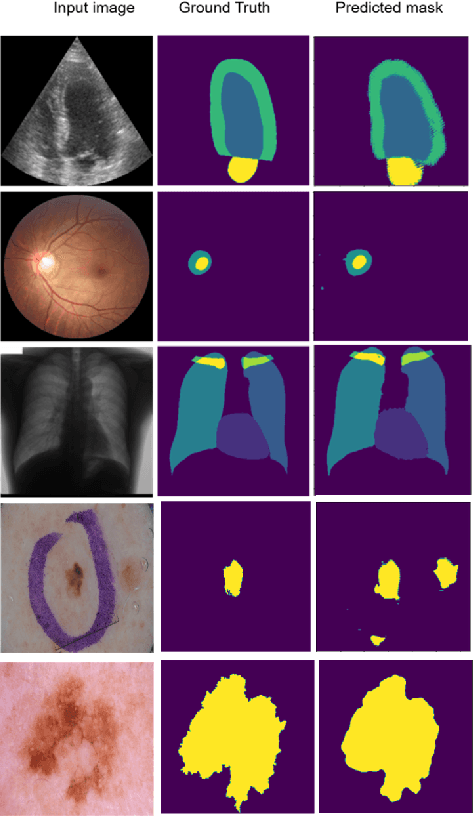
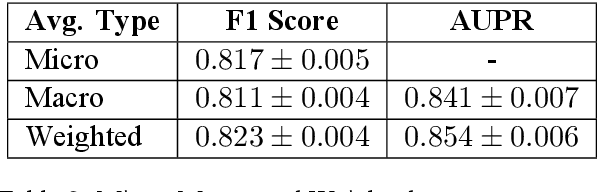
Abstract:Supervised deep learning methods for semantic medical image segmentation are getting increasingly popular in the past few years.However, in resource constrained settings, getting large number of annotated images is very difficult as it mostly requires experts, is expensive and time-consuming.Semi-supervised segmentation can be an attractive solution where a very few labeled images are used along with a large number of unlabeled ones. While the gap between supervised and semi-supervised methods have been dramatically reduced for classification problems in the past couple of years, there still remains a larger gap in segmentation methods. In this work, we adapt a state-of-the-art semi-supervised classification method FixMatch to semantic segmentation task, introducing FixMatchSeg. FixMatchSeg is evaluated in four different publicly available datasets of different anatomy and different modality: cardiac ultrasound, chest X-ray, retinal fundus image, and skin images. When there are few labels, we show that FixMatchSeg performs on par with strong supervised baselines.
Label Geometry Aware Discriminator for Conditional Generative Networks
May 12, 2021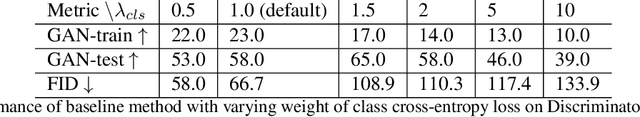
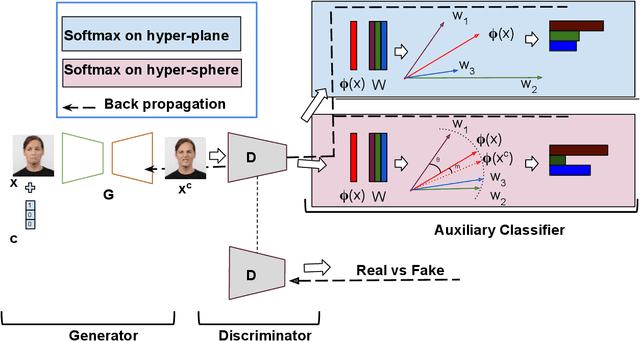
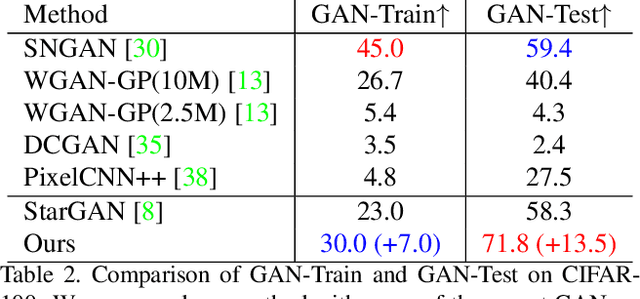
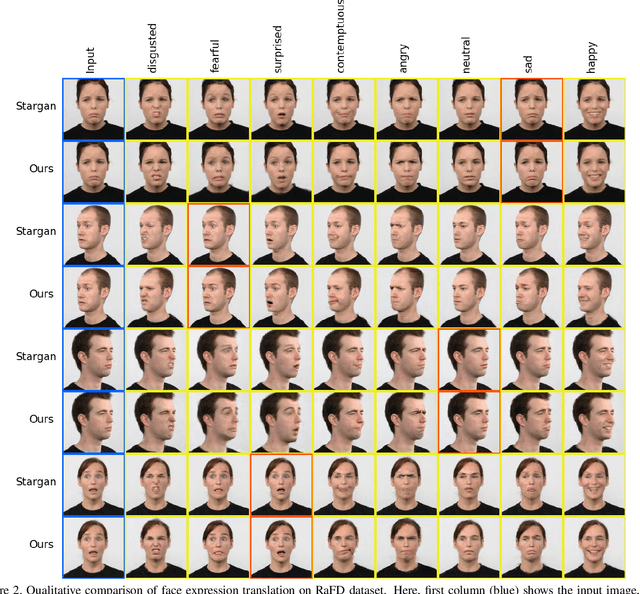
Abstract:Multi-domain image-to-image translation with conditional Generative Adversarial Networks (GANs) can generate highly photo realistic images with desired target classes, yet these synthetic images have not always been helpful to improve downstream supervised tasks such as image classification. Improving downstream tasks with synthetic examples requires generating images with high fidelity to the unknown conditional distribution of the target class, which many labeled conditional GANs attempt to achieve by adding soft-max cross-entropy loss based auxiliary classifier in the discriminator. As recent studies suggest that the soft-max loss in Euclidean space of deep feature does not leverage their intrinsic angular distribution, we propose to replace this loss in auxiliary classifier with an additive angular margin (AAM) loss that takes benefit of the intrinsic angular distribution, and promotes intra-class compactness and inter-class separation to help generator synthesize high fidelity images. We validate our method on RaFD and CIFAR-100, two challenging face expression and natural image classification data set. Our method outperforms state-of-the-art methods in several different evaluation criteria including recently proposed GAN-train and GAN-test metrics designed to assess the impact of synthetic data on downstream classification task, assessing the usefulness in data augmentation for supervised tasks with prediction accuracy score and average confidence score, and the well known FID metric.
Uncertainty Estimation in Deep 2D Echocardiography Segmentation
May 19, 2020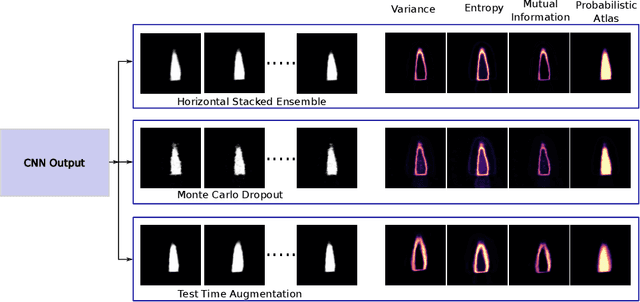
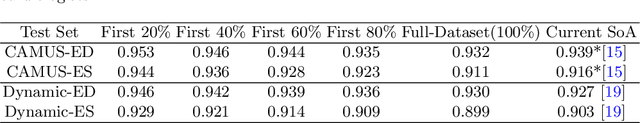
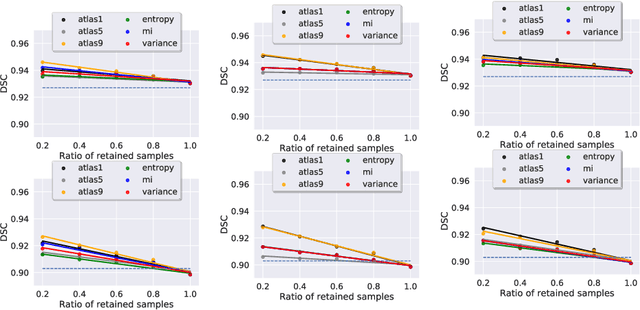
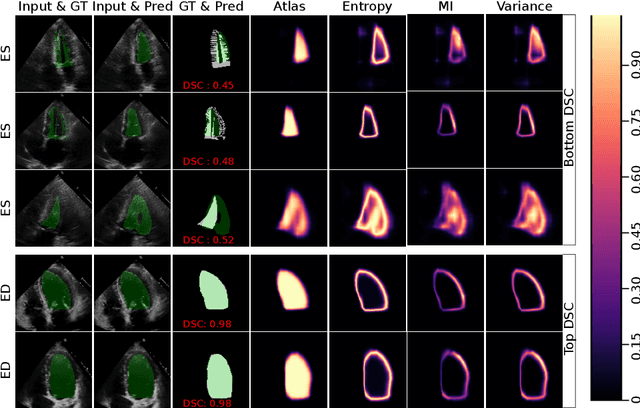
Abstract:2D echocardiography is the most common imaging modality for cardiovascular diseases. The portability and relatively low-cost nature of Ultrasound (US) enable the US devices needed for performing echocardiography to be made widely available. However, acquiring and interpreting cardiac US images is operator dependent, limiting its use to only places where experts are present. Recently, Deep Learning (DL) has been used in 2D echocardiography for automated view classification, and structure and function assessment. Although these recent works show promise in developing computer-guided acquisition and automated interpretation of echocardiograms, most of these methods do not model and estimate uncertainty which can be important when testing on data coming from a distribution further away from that of the training data. Uncertainty estimates can be beneficial both during the image acquisition phase (by providing real-time feedback to the operator on acquired image's quality), and during automated measurement and interpretation. The performance of uncertainty models and quantification metric may depend on the prediction task and the models being compared. Hence, to gain insight of uncertainty modelling for left ventricular segmentation from US images, we compare three ensembling based uncertainty models quantified using four different metrics (one newly proposed) on state-of-the-art baseline networks using two publicly available echocardiogram datasets. We further demonstrate how uncertainty estimation can be used to automatically reject poor quality images and improve state-of-the-art segmentation results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge