Berkin Bilgic
Harvard-MIT Health Sciences and Technology, Athinoula A. Martinos Center for Biomedical Imaging, Harvard Medical School
Chaos and COSMOS -- Considerations on QSM methods with multiple and single orientations and effects from local anisotropy
Sep 30, 2023Abstract:Purpose: Field-to-susceptibility inversion in quantitative susceptibility mapping (QSM) is ill-posed and needs numerical stabilization through either regularization or oversampling by acquiring data at three or more object orientations. Calculation Of Susceptibility through Multiple Orientations Sampling (COSMOS) is an established oversampling approach and regarded as QSM gold standard. It achieves a well-conditioned inverse problem, requiring rotations by 0{\deg}, 60{\deg} and 120{\deg} in the yz-plane. However, this is impractical in vivo, where head rotations are typically restricted to a range of +-25{\deg}. Non-ideal sampling degrades the conditioning with residual streaking artifacts whose mitigation needs further regularization. Moreover, susceptibility anisotropy in white matter is not considered in the COSMOS model, which may introduce additional bias. The current work presents a thorough investigation of these effects in primate brain. Methods: Gradient-recalled echo (GRE) data of an entire fixed chimpanzee brain were acquired at 7 T (350 microns resolution, 10 orientations) including ideal COSMOS sampling and realistic rotations in vivo. Comparisons of the results included ideal COSMOS, in-vivo feasible acquisitions with 3-8 orientations and single-orientation iLSQR QSM. Results: In-vivo feasible and optimal COSMOS yielded high-quality susceptibility maps with increased SNR resulting from averaging multiple acquisitions. COSMOS reconstructions from non-ideal rotations about a single axis required additional L2-regularization to mitigate residual streaking artifacts. Conclusion: In view of unconsidered anisotropy effects, added complexity of the reconstruction, and the general challenge of multi-orientation acquisitions, advantages of sub-optimal COSMOS schemes over regularized single-orientation QSM appear limited in in-vivo settings.
Improved Multi-Shot Diffusion-Weighted MRI with Zero-Shot Self-Supervised Learning Reconstruction
Aug 09, 2023



Abstract:Diffusion MRI is commonly performed using echo-planar imaging (EPI) due to its rapid acquisition time. However, the resolution of diffusion-weighted images is often limited by magnetic field inhomogeneity-related artifacts and blurring induced by T2- and T2*-relaxation effects. To address these limitations, multi-shot EPI (msEPI) combined with parallel imaging techniques is frequently employed. Nevertheless, reconstructing msEPI can be challenging due to phase variation between multiple shots. In this study, we introduce a novel msEPI reconstruction approach called zero-MIRID (zero-shot self-supervised learning of Multi-shot Image Reconstruction for Improved Diffusion MRI). This method jointly reconstructs msEPI data by incorporating deep learning-based image regularization techniques. The network incorporates CNN denoisers in both k- and image-spaces, while leveraging virtual coils to enhance image reconstruction conditioning. By employing a self-supervised learning technique and dividing sampled data into three groups, the proposed approach achieves superior results compared to the state-of-the-art parallel imaging method, as demonstrated in an in-vivo experiment.
Zero-DeepSub: Zero-Shot Deep Subspace Reconstruction for Rapid Multiparametric Quantitative MRI Using 3D-QALAS
Jul 04, 2023Abstract:Purpose: To develop and evaluate methods for 1) reconstructing 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) time-series images using a low-rank subspace method, which enables accurate and rapid T1 and T2 mapping, and 2) improving the fidelity of subspace QALAS by combining scan-specific deep-learning-based reconstruction and subspace modeling. Methods: A low-rank subspace method for 3D-QALAS (i.e., subspace QALAS) and zero-shot deep-learning subspace method (i.e., Zero-DeepSub) were proposed for rapid and high fidelity T1 and T2 mapping and time-resolved imaging using 3D-QALAS. Using an ISMRM/NIST system phantom, the accuracy of the T1 and T2 maps estimated using the proposed methods was evaluated by comparing them with reference techniques. The reconstruction performance of the proposed subspace QALAS using Zero-DeepSub was evaluated in vivo and compared with conventional QALAS at high reduction factors of up to 9-fold. Results: Phantom experiments showed that subspace QALAS had good linearity with respect to the reference methods while reducing biases compared to conventional QALAS, especially for T2 maps. Moreover, in vivo results demonstrated that subspace QALAS had better g-factor maps and could reduce voxel blurring, noise, and artifacts compared to conventional QALAS and showed robust performance at up to 9-fold acceleration with Zero-DeepSub, which enabled whole-brain T1, T2, and PD mapping at 1 mm isotropic resolution within 2 min of scan time. Conclusion: The proposed subspace QALAS along with Zero-DeepSub enabled high fidelity and rapid whole-brain multiparametric quantification and time-resolved imaging.
SSL-QALAS: Self-Supervised Learning for Rapid Multiparameter Estimation in Quantitative MRI Using 3D-QALAS
Feb 28, 2023Abstract:Purpose: To develop and evaluate a method for rapid estimation of multiparametric T1, T2, proton density (PD), and inversion efficiency (IE) maps from 3D-quantification using an interleaved Look-Locker acquisition sequence with T2 preparation pulse (3D-QALAS) measurements using self-supervised learning (SSL) without the need for an external dictionary. Methods: A SSL-based QALAS mapping method (SSL-QALAS) was developed for rapid and dictionary-free estimation of multiparametric maps from 3D-QALAS measurements. The accuracy of the reconstructed quantitative maps using dictionary matching and SSL-QALAS was evaluated by comparing the estimated T1 and T2 values with those obtained from the reference methods on an ISMRM/NIST phantom. The SSL-QALAS and the dictionary matching methods were also compared in vivo, and generalizability was evaluated by comparing the scan-specific, pre-trained, and transfer learning models. Results: Phantom experiments showed that both the dictionary matching and SSL-QALAS methods produced T1 and T2 estimates that had a strong linear agreement with the reference values in the ISMRM/NIST phantom. Further, SSL-QALAS showed similar performance with dictionary matching in reconstructing the T1, T2, PD, and IE maps on in vivo data. Rapid reconstruction of multiparametric maps was enabled by inferring the data using a pre-trained SSL-QALAS model within 10 s. Fast scan-specific tuning was also demonstrated by fine-tuning the pre-trained model with the target subject's data within 15 min. Conclusion: The proposed SSL-QALAS method enabled rapid reconstruction of multiparametric maps from 3D-QALAS measurements without an external dictionary or labeled ground-truth training data.
3D-EPI Blip-Up/Down Acquisition with CAIPI and Joint Hankel Structured Low-Rank Reconstruction for Rapid Distortion-Free High-Resolution T2* Mapping
Dec 01, 2022Abstract:Purpose: This work aims to develop a novel distortion-free 3D-EPI acquisition and image reconstruction technique for fast and robust, high-resolution, whole-brain imaging as well as quantitative T2* mapping. Methods: 3D-Blip-Up and -Down Acquisition (3D-BUDA) sequence is designed for both single- and multi-echo 3D GRE-EPI imaging using multiple shots with blip-up and -down readouts to encode B0 field map information. Complementary k-space coverage is achieved using controlled aliasing in parallel imaging (CAIPI) sampling across the shots. For image reconstruction, an iterative hard-thresholding algorithm is employed to minimize the cost function that combines field map information informed parallel imaging with the structured low-rank constraint for multi-shot 3D-BUDA data. Extending 3D-BUDA to multi-echo imaging permits T2* mapping. For this, we propose constructing a joint Hankel matrix along both echo and shot dimensions to improve the reconstruction. Results: Experimental results on in vivo multi-echo data demonstrate that, by performing joint reconstruction along with both echo and shot dimensions, reconstruction accuracy is improved compared to standard 3D-BUDA reconstruction. CAIPI sampling is further shown to enhance the image quality. For T2* mapping, T2* values from 3D-Joint-CAIPI-BUDA and reference multi-echo GRE are within limits of agreement as quantified by Bland-Altman analysis. Conclusions: The proposed technique enables rapid 3D distortion-free high-resolution imaging and T2* mapping. Specifically, 3D-BUDA enables 1-mm isotropic whole-brain imaging in 22 s at 3 T and 9 s on a 7 T scanner. The combination of multi-echo 3D-BUDA with CAIPI acquisition and joint reconstruction enables distortion-free whole-brain T2* mapping in 47 s at 1.1x1.1x1.0 mm3 resolution.
SRNR: Training neural networks for Super-Resolution MRI using Noisy high-resolution Reference data
Nov 10, 2022



Abstract:Neural network (NN) based approaches for super-resolution MRI typically require high-SNR high-resolution reference data acquired in many subjects, which is time consuming and a barrier to feasible and accessible implementation. We propose to train NNs for Super-Resolution using Noisy Reference data (SRNR), leveraging the mechanism of the classic NN-based denoising method Noise2Noise. We systematically demonstrate that results from NNs trained using noisy and high-SNR references are similar for both simulated and empirical data. SRNR suggests a smaller number of repetitions of high-resolution reference data can be used to simplify the training data preparation for super-resolution MRI.
Time-efficient, High Resolution 3T Whole Brain Quantitative Relaxometry using 3D-QALAS with Wave-CAIPI Readouts
Nov 08, 2022



Abstract:Purpose: Volumetric, high resolution, quantitative mapping of brain tissues relaxation properties is hindered by long acquisition times and SNR challenges. This study, for the first time, combines the time efficient wave-CAIPI readouts into the 3D-QALAS acquisition scheme, enabling full brain quantitative T1, T2 and PD maps at 1.15 isotropic voxels in only 3 minutes. Methods: Wave-CAIPI readouts were embedded in the standard 3d-QALAS encoding scheme, enabling full brain quantitative parameter maps (T1, T2 and PD) at acceleration factors of R=3x2 with minimum SNR loss due to g-factor penalties. The quantitative maps using the accelerated protocol were quantitatively compared against those obtained from conventional 3D-QALAS sequence using GRAPPA acceleration of R=2 in the ISMRM NIST phantom, and ten healthy volunteers. To show the feasibility of the proposed methods in clinical settings, the accelerated wave-CAIPI 3D-QALAS sequence was also employed in pediatric patients undergoing clinical MRI examinations. Results: When tested in both the ISMRM/NIST phantom and 7 healthy volunteers, the quantitative maps using the accelerated protocol showed excellent agreement against those obtained from conventional 3D-QALAS at R=2. Conclusion: 3D-QALAS enhanced with wave-CAIPI readouts enables time-efficient, full brain quantitative T1, T2 and PD mapping at 1.15 in 3 minutes at R=3x2 acceleration. When tested on the NIST phantom and 7 healthy volunteers, the quantitative maps obtained from the accelerated wave-CAIPI 3D-QALAS protocol showed very similar values to those obtained from the standard 3D-QALAS (R=2) protocol, alluding to the robustness and reliability of the proposed methods. This study also shows that the accelerated protocol can be effectively employed in pediatric patient populations, making high-quality high-resolution full brain quantitative imaging feasible in clinical settings.
Latent Signal Models: Learning Compact Representations of Signal Evolution for Improved Time-Resolved, Multi-contrast MRI
Aug 27, 2022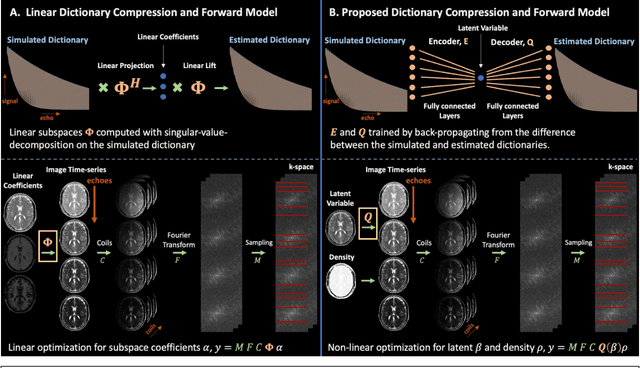
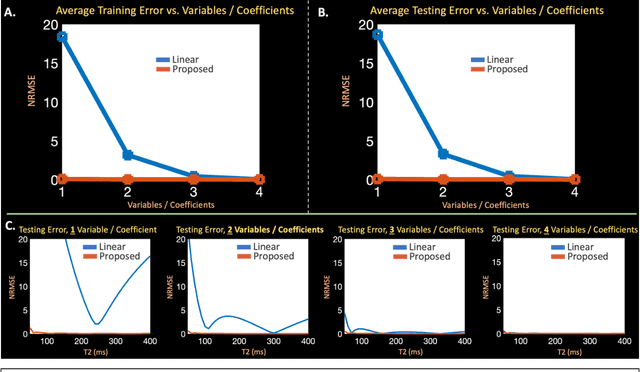
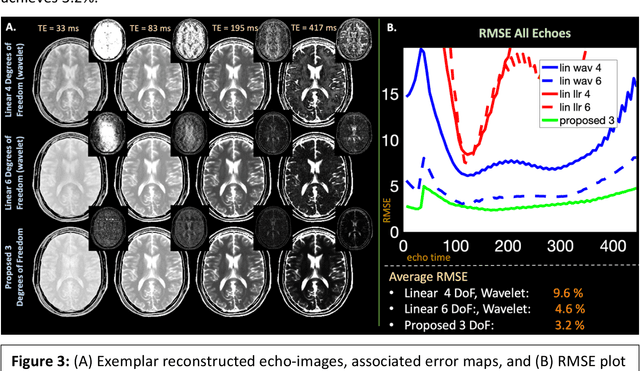
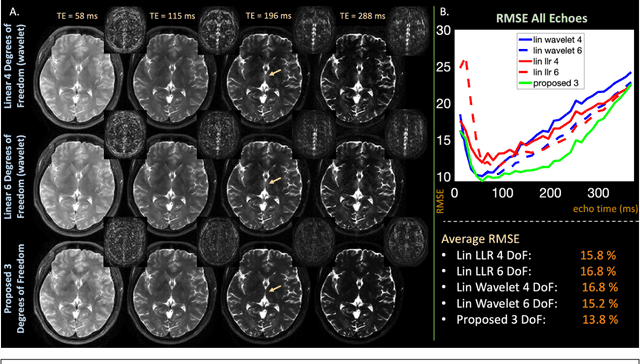
Abstract:Purpose: Training auto-encoders on simulated signal evolution and inserting the decoder into the forward model improves reconstructions through more compact, Bloch-equation-based representations of signal in comparison to linear subspaces. Methods: Building on model-based nonlinear and linear subspace techniques that enable reconstruction of signal dynamics, we train auto-encoders on dictionaries of simulated signal evolution to learn more compact, non-linear, latent representations. The proposed Latent Signal Model framework inserts the decoder portion of the auto-encoder into the forward model and directly reconstructs the latent representation. Latent Signal Models essentially serve as a proxy for fast and feasible differentiation through the Bloch-equations used to simulate signal. This work performs experiments in the context of T2-shuffling, gradient echo EPTI, and MPRAGE-shuffling. We compare how efficiently auto-encoders represent signal evolution in comparison to linear subspaces. Simulation and in-vivo experiments then evaluate if reducing degrees of freedom by inserting the decoder into the forward model improves reconstructions in comparison to subspace constraints. Results: An auto-encoder with one real latent variable represents FSE, EPTI, and MPRAGE signal evolution as well as linear subspaces characterized by four basis vectors. In simulated/in-vivo T2-shuffling and in-vivo EPTI experiments, the proposed framework achieves consistent quantitative NRMSE and qualitative improvement over linear approaches. From qualitative evaluation, the proposed approach yields images with reduced blurring and noise amplification in MPRAGE shuffling experiments. Conclusion: Directly solving for non-linear latent representations of signal evolution improves time-resolved MRI reconstructions through reduced degrees of freedom.
High-Dimensional Sparse Bayesian Learning without Covariance Matrices
Feb 25, 2022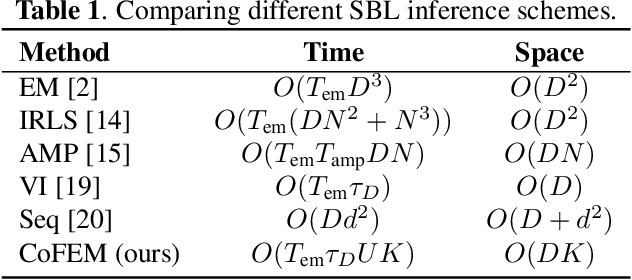
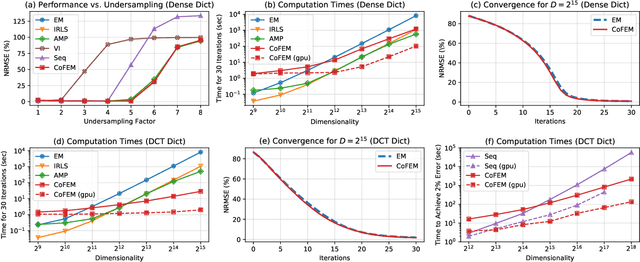
Abstract:Sparse Bayesian learning (SBL) is a powerful framework for tackling the sparse coding problem. However, the most popular inference algorithms for SBL become too expensive for high-dimensional settings, due to the need to store and compute a large covariance matrix. We introduce a new inference scheme that avoids explicit construction of the covariance matrix by solving multiple linear systems in parallel to obtain the posterior moments for SBL. Our approach couples a little-known diagonal estimation result from numerical linear algebra with the conjugate gradient algorithm. On several simulations, our method scales better than existing approaches in computation time and memory, especially for structured dictionaries capable of fast matrix-vector multiplication.
* 5 pages
Wave-Encoded Model-based Deep Learning for Highly Accelerated Imaging with Joint Reconstruction
Feb 06, 2022



Abstract:Purpose: To propose a wave-encoded model-based deep learning (wave-MoDL) strategy for highly accelerated 3D imaging and joint multi-contrast image reconstruction, and further extend this to enable rapid quantitative imaging using an interleaved look-locker acquisition sequence with T2 preparation pulse (3D-QALAS). Method: Recently introduced MoDL technique successfully incorporates convolutional neural network (CNN)-based regularizers into physics-based parallel imaging reconstruction using a small number of network parameters. Wave-CAIPI is an emerging parallel imaging method that accelerates the imaging speed by employing sinusoidal gradients in the phase- and slice-encoding directions during the readout to take better advantage of 3D coil sensitivity profiles. In wave-MoDL, we propose to combine the wave-encoding strategy with unrolled network constraints to accelerate the acquisition speed while enforcing wave-encoded data consistency. We further extend wave-MoDL to reconstruct multi-contrast data with controlled aliasing in parallel imaging (CAIPI) sampling patterns to leverage similarity between multiple images to improve the reconstruction quality. Result: Wave-MoDL enables a 47-second MPRAGE acquisition at 1 mm resolution at 16-fold acceleration. For quantitative imaging, wave-MoDL permits a 2-minute acquisition for T1, T2, and proton density mapping at 1 mm resolution at 12-fold acceleration, from which contrast weighted images can be synthesized as well. Conclusion: Wave-MoDL allows rapid MR acquisition and high-fidelity image reconstruction and may facilitate clinical and neuroscientific applications by incorporating unrolled neural networks into wave-CAIPI reconstruction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge