Arindam Dutta
Towards Source-Free Machine Unlearning
Aug 20, 2025



Abstract:As machine learning becomes more pervasive and data privacy regulations evolve, the ability to remove private or copyrighted information from trained models is becoming an increasingly critical requirement. Existing unlearning methods often rely on the assumption of having access to the entire training dataset during the forgetting process. However, this assumption may not hold true in practical scenarios where the original training data may not be accessible, i.e., the source-free setting. To address this challenge, we focus on the source-free unlearning scenario, where an unlearning algorithm must be capable of removing specific data from a trained model without requiring access to the original training dataset. Building on recent work, we present a method that can estimate the Hessian of the unknown remaining training data, a crucial component required for efficient unlearning. Leveraging this estimation technique, our method enables efficient zero-shot unlearning while providing robust theoretical guarantees on the unlearning performance, while maintaining performance on the remaining data. Extensive experiments over a wide range of datasets verify the efficacy of our method.
Leveraging Synthetic Adult Datasets for Unsupervised Infant Pose Estimation
Apr 08, 2025Abstract:Human pose estimation is a critical tool across a variety of healthcare applications. Despite significant progress in pose estimation algorithms targeting adults, such developments for infants remain limited. Existing algorithms for infant pose estimation, despite achieving commendable performance, depend on fully supervised approaches that require large amounts of labeled data. These algorithms also struggle with poor generalizability under distribution shifts. To address these challenges, we introduce SHIFT: Leveraging SyntHetic Adult Datasets for Unsupervised InFanT Pose Estimation, which leverages the pseudo-labeling-based Mean-Teacher framework to compensate for the lack of labeled data and addresses distribution shifts by enforcing consistency between the student and the teacher pseudo-labels. Additionally, to penalize implausible predictions obtained from the mean-teacher framework, we incorporate an infant manifold pose prior. To enhance SHIFT's self-occlusion perception ability, we propose a novel visibility consistency module for improved alignment of the predicted poses with the original image. Extensive experiments on multiple benchmarks show that SHIFT significantly outperforms existing state-of-the-art unsupervised domain adaptation (UDA) pose estimation methods by 5% and supervised infant pose estimation methods by a margin of 16%. The project page is available at: https://sarosijbose.github.io/SHIFT.
Uncertainty-Aware Diffusion Guided Refinement of 3D Scenes
Mar 19, 2025Abstract:Reconstructing 3D scenes from a single image is a fundamentally ill-posed task due to the severely under-constrained nature of the problem. Consequently, when the scene is rendered from novel camera views, existing single image to 3D reconstruction methods render incoherent and blurry views. This problem is exacerbated when the unseen regions are far away from the input camera. In this work, we address these inherent limitations in existing single image-to-3D scene feedforward networks. To alleviate the poor performance due to insufficient information beyond the input image's view, we leverage a strong generative prior in the form of a pre-trained latent video diffusion model, for iterative refinement of a coarse scene represented by optimizable Gaussian parameters. To ensure that the style and texture of the generated images align with that of the input image, we incorporate on-the-fly Fourier-style transfer between the generated images and the input image. Additionally, we design a semantic uncertainty quantification module that calculates the per-pixel entropy and yields uncertainty maps used to guide the refinement process from the most confident pixels while discarding the remaining highly uncertain ones. We conduct extensive experiments on real-world scene datasets, including in-domain RealEstate-10K and out-of-domain KITTI-v2, showing that our approach can provide more realistic and high-fidelity novel view synthesis results compared to existing state-of-the-art methods.
CHROME: Clothed Human Reconstruction with Occlusion-Resilience and Multiview-Consistency from a Single Image
Mar 19, 2025



Abstract:Reconstructing clothed humans from a single image is a fundamental task in computer vision with wide-ranging applications. Although existing monocular clothed human reconstruction solutions have shown promising results, they often rely on the assumption that the human subject is in an occlusion-free environment. Thus, when encountering in-the-wild occluded images, these algorithms produce multiview inconsistent and fragmented reconstructions. Additionally, most algorithms for monocular 3D human reconstruction leverage geometric priors such as SMPL annotations for training and inference, which are extremely challenging to acquire in real-world applications. To address these limitations, we propose CHROME: Clothed Human Reconstruction with Occlusion-Resilience and Multiview-ConsistEncy from a Single Image, a novel pipeline designed to reconstruct occlusion-resilient 3D humans with multiview consistency from a single occluded image, without requiring either ground-truth geometric prior annotations or 3D supervision. Specifically, CHROME leverages a multiview diffusion model to first synthesize occlusion-free human images from the occluded input, compatible with off-the-shelf pose control to explicitly enforce cross-view consistency during synthesis. A 3D reconstruction model is then trained to predict a set of 3D Gaussians conditioned on both the occluded input and synthesized views, aligning cross-view details to produce a cohesive and accurate 3D representation. CHROME achieves significant improvements in terms of both novel view synthesis (upto 3 db PSNR) and geometric reconstruction under challenging conditions.
Unsupervised Domain Adaptation for Occlusion Resilient Human Pose Estimation
Jan 06, 2025



Abstract:Occlusions are a significant challenge to human pose estimation algorithms, often resulting in inaccurate and anatomically implausible poses. Although current occlusion-robust human pose estimation algorithms exhibit impressive performance on existing datasets, their success is largely attributed to supervised training and the availability of additional information, such as multiple views or temporal continuity. Furthermore, these algorithms typically suffer from performance degradation under distribution shifts. While existing domain adaptive human pose estimation algorithms address this bottleneck, they tend to perform suboptimally when the target domain images are occluded, a common occurrence in real-life scenarios. To address these challenges, we propose OR-POSE: Unsupervised Domain Adaptation for Occlusion Resilient Human POSE Estimation. OR-POSE is an innovative unsupervised domain adaptation algorithm which effectively mitigates domain shifts and overcomes occlusion challenges by employing the mean teacher framework for iterative pseudo-label refinement. Additionally, OR-POSE reinforces realistic pose prediction by leveraging a learned human pose prior which incorporates the anatomical constraints of humans in the adaptation process. Lastly, OR-POSE avoids overfitting to inaccurate pseudo labels generated from heavily occluded images by employing a novel visibility-based curriculum learning approach. This enables the model to gradually transition from training samples with relatively less occlusion to more challenging, heavily occluded samples. Extensive experiments show that OR-POSE outperforms existing analogous state-of-the-art algorithms by $\sim$ 7% on challenging occluded human pose estimation datasets.
Unfair Alignment: Examining Safety Alignment Across Vision Encoder Layers in Vision-Language Models
Nov 06, 2024



Abstract:Vision-language models (VLMs) have improved significantly in multi-modal tasks, but their more complex architecture makes their safety alignment more challenging than the alignment of large language models (LLMs). In this paper, we reveal an unfair distribution of safety across the layers of VLM's vision encoder, with earlier and middle layers being disproportionately vulnerable to malicious inputs compared to the more robust final layers. This 'cross-layer' vulnerability stems from the model's inability to generalize its safety training from the default architectural settings used during training to unseen or out-of-distribution scenarios, leaving certain layers exposed. We conduct a comprehensive analysis by projecting activations from various intermediate layers and demonstrate that these layers are more likely to generate harmful outputs when exposed to malicious inputs. Our experiments with LLaVA-1.5 and Llama 3.2 show discrepancies in attack success rates and toxicity scores across layers, indicating that current safety alignment strategies focused on a single default layer are insufficient.
Multi-modal Pose Diffuser: A Multimodal Generative Conditional Pose Prior
Oct 18, 2024Abstract:The Skinned Multi-Person Linear (SMPL) model plays a crucial role in 3D human pose estimation, providing a streamlined yet effective representation of the human body. However, ensuring the validity of SMPL configurations during tasks such as human mesh regression remains a significant challenge , highlighting the necessity for a robust human pose prior capable of discerning realistic human poses. To address this, we introduce MOPED: \underline{M}ulti-m\underline{O}dal \underline{P}os\underline{E} \underline{D}iffuser. MOPED is the first method to leverage a novel multi-modal conditional diffusion model as a prior for SMPL pose parameters. Our method offers powerful unconditional pose generation with the ability to condition on multi-modal inputs such as images and text. This capability enhances the applicability of our approach by incorporating additional context often overlooked in traditional pose priors. Extensive experiments across three distinct tasks-pose estimation, pose denoising, and pose completion-demonstrate that our multi-modal diffusion model-based prior significantly outperforms existing methods. These results indicate that our model captures a broader spectrum of plausible human poses.
POSTURE: Pose Guided Unsupervised Domain Adaptation for Human Body Part Segmentation
Jul 04, 2024



Abstract:Existing algorithms for human body part segmentation have shown promising results on challenging datasets, primarily relying on end-to-end supervision. However, these algorithms exhibit severe performance drops in the face of domain shifts, leading to inaccurate segmentation masks. To tackle this issue, we introduce POSTURE: \underline{Po}se Guided Un\underline{s}upervised Domain Adap\underline{t}ation for H\underline{u}man Body Pa\underline{r}t S\underline{e}gmentation - an innovative pseudo-labelling approach designed to improve segmentation performance on the unlabeled target data. Distinct from conventional domain adaptive methods for general semantic segmentation, POSTURE stands out by considering the underlying structure of the human body and uses anatomical guidance from pose keypoints to drive the adaptation process. This strong inductive prior translates to impressive performance improvements, averaging 8\% over existing state-of-the-art domain adaptive semantic segmentation methods across three benchmark datasets. Furthermore, the inherent flexibility of our proposed approach facilitates seamless extension to source-free settings (SF-POSTURE), effectively mitigating potential privacy and computational concerns, with negligible drop in performance.
TEMP3D: Temporally Continuous 3D Human Pose Estimation Under Occlusions
Dec 24, 2023
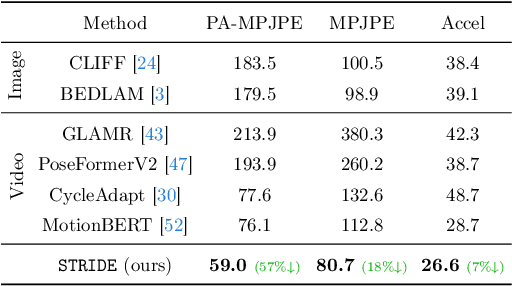
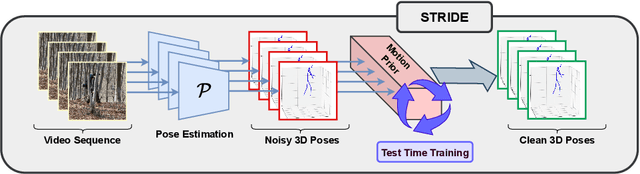
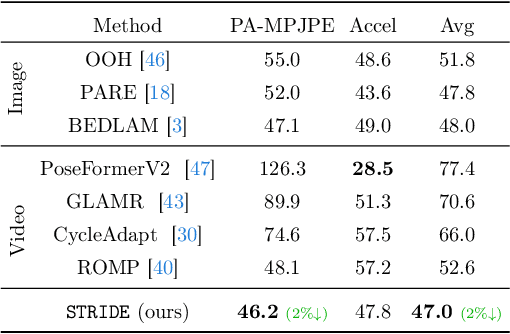
Abstract:Existing 3D human pose estimation methods perform remarkably well in both monocular and multi-view settings. However, their efficacy diminishes significantly in the presence of heavy occlusions, which limits their practical utility. For video sequences, temporal continuity can help infer accurate poses, especially in heavily occluded frames. In this paper, we aim to leverage this potential of temporal continuity through human motion priors, coupled with large-scale pre-training on 3D poses and self-supervised learning, to enhance 3D pose estimation in a given video sequence. This leads to a temporally continuous 3D pose estimate on unlabelled in-the-wild videos, which may contain occlusions, while exclusively relying on pre-trained 3D pose models. We propose an unsupervised method named TEMP3D that aligns a motion prior model on a given in-the-wild video using existing SOTA single image-based 3D pose estimation methods to give temporally continuous output under occlusions. To evaluate our method, we test it on the Occluded Human3.6M dataset, our custom-built dataset which contains significantly large (up to 100%) human body occlusions incorporated into the Human3.6M dataset. We achieve SOTA results on Occluded Human3.6M and the OcMotion dataset while maintaining competitive performance on non-occluded data. URL: https://sites.google.com/ucr.edu/temp3d
Active Learning Guided Federated Online Adaptation: Applications in Medical Image Segmentation
Dec 08, 2023Abstract:Data privacy, storage, and distribution shifts are major bottlenecks in medical image analysis. Data cannot be shared across patients, physicians, and facilities due to privacy concerns, usually requiring each patient's data to be analyzed in a discreet setting at a near real-time pace. However, one would like to take advantage of the accumulated knowledge across healthcare facilities as the computational systems analyze data of more and more patients while incorporating feedback provided by physicians to improve accuracy. Motivated by these, we propose a method for medical image segmentation that adapts to each incoming data batch (online adaptation), incorporates physician feedback through active learning, and assimilates knowledge across facilities in a federated setup. Combining an online adaptation scheme at test time with an efficient sampling strategy with budgeted annotation helps bridge the gap between the source and the incoming stream of target domain data. A federated setup allows collaborative aggregation of knowledge across distinct distributed models without needing to share the data across different models. This facilitates the improvement of performance over time by accumulating knowledge across users. Towards achieving these goals, we propose a computationally amicable, privacy-preserving image segmentation technique \textbf{DrFRODA} that uses federated learning to adapt the model in an online manner with feedback from doctors in the loop. Our experiments on publicly available datasets show that the proposed distributed active learning-based online adaptation method outperforms unsupervised online adaptation methods and shows competitive results with offline active learning-based adaptation methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge