"Image": models, code, and papers
Max-Fusion U-Net for Multi-Modal Pathology Segmentation with Attention and Dynamic Resampling
Sep 05, 2020



Automatic segmentation of multi-sequence (multi-modal) cardiac MR (CMR) images plays a significant role in diagnosis and management for a variety of cardiac diseases. However, the performance of relevant algorithms is significantly affected by the proper fusion of the multi-modal information. Furthermore, particular diseases, such as myocardial infarction, display irregular shapes on images and occupy small regions at random locations. These facts make pathology segmentation of multi-modal CMR images a challenging task. In this paper, we present the Max-Fusion U-Net that achieves improved pathology segmentation performance given aligned multi-modal images of LGE, T2-weighted, and bSSFP modalities. Specifically, modality-specific features are extracted by dedicated encoders. Then they are fused with the pixel-wise maximum operator. Together with the corresponding encoding features, these representations are propagated to decoding layers with U-Net skip-connections. Furthermore, a spatial-attention module is applied in the last decoding layer to encourage the network to focus on those small semantically meaningful pathological regions that trigger relatively high responses by the network neurons. We also use a simple image patch extraction strategy to dynamically resample training examples with varying spacial and batch sizes. With limited GPU memory, this strategy reduces the imbalance of classes and forces the model to focus on regions around the interested pathology. It further improves segmentation accuracy and reduces the mis-classification of pathology. We evaluate our methods using the Myocardial pathology segmentation (MyoPS) combining the multi-sequence CMR dataset which involves three modalities. Extensive experiments demonstrate the effectiveness of the proposed model which outperforms the related baselines.
* 13 pages, 7 figures, conference paper
Out-of-Distribution Generalization with Maximal Invariant Predictor
Aug 04, 2020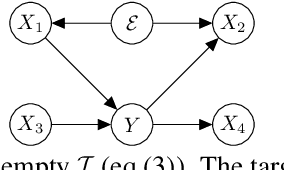
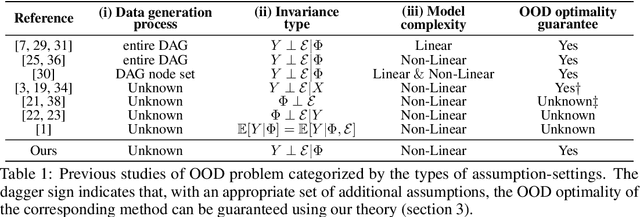
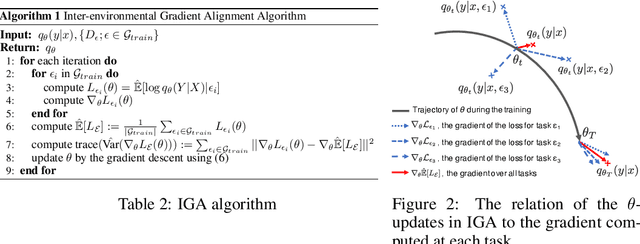


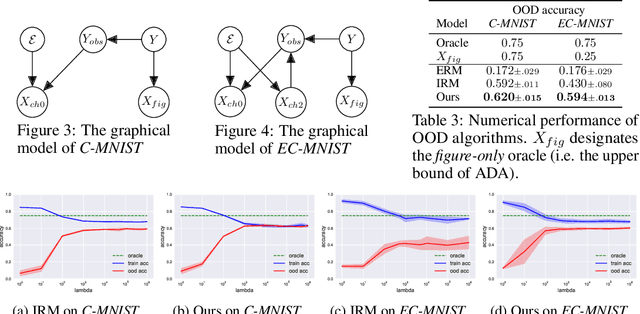
Out-of-Distribution (OOD) generalization problem is a problem of seeking the predictor function whose performance in the worst environments is optimal. This paper makes two contributions to OOD problem. We first use the basic results of probability to prove maximal Invariant Predictor(MIP) condition, a theoretical result that can be used to identify the OOD optimal solution. We then use our MIP to derive inner-environmental Gradient Alignment(IGA) algorithm that can be used to help seek the OOD optimal predictor. Previous studies that have investigated the theoretical aspect of the OOD-problem use strong structural assumptions such as causal DAG. However, in cases involving image datasets, for example, the identification of hidden structural relations is itself a difficult problem. Our theoretical results are different from those of many previous studies in that it can be applied to cases in which the underlying structure of a dataset is difficult to analyze. We present an extensive comparison of previous theoretical approaches to the OODproblems based on the assumptions they make. We also present an extension of the colored-MNIST that can more accurately represent the pathological OOD situation than the original version, and demonstrate the superiority of IGA over previous methods on both the original and the extended version of Colored-MNIST.
Unsupervised Pose Flow Learning for Pose Guided Synthesis
Sep 30, 2019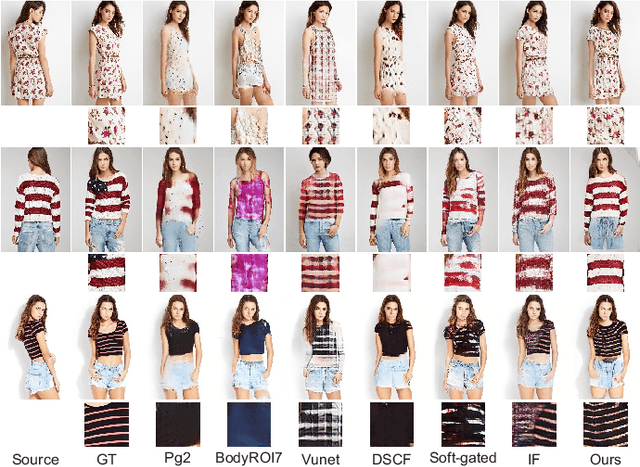
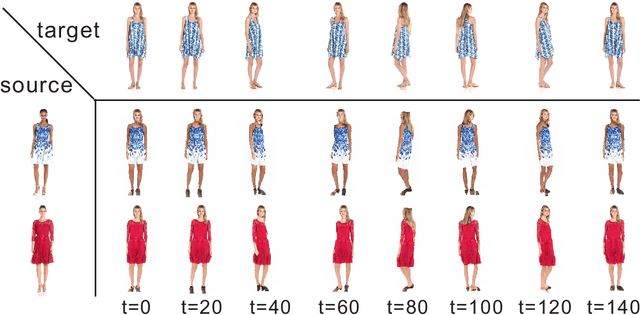


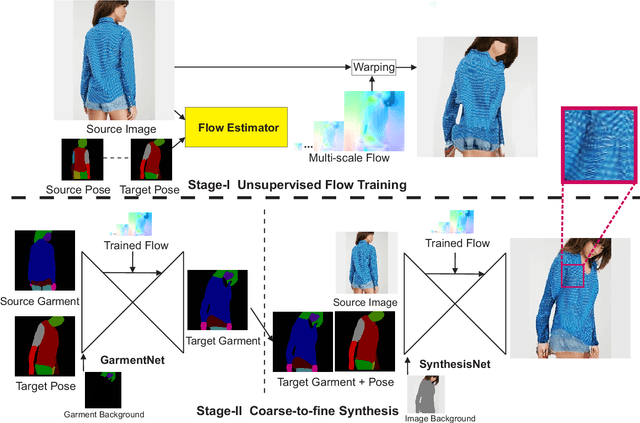
Pose guided synthesis aims to generate a new image in an arbitrary target pose while preserving the appearance details from the source image. Existing approaches rely on either hard-coded spatial transformations or 3D body modeling. They often overlook complex non-rigid pose deformation or unmatched occluded regions, thus fail to effectively preserve appearance information. In this paper, we propose an unsupervised pose flow learning scheme that learns to transfer the appearance details from the source image. Based on such learned pose flow, we proposed GarmentNet and SynthesisNet, both of which use multi-scale feature-domain alignment for coarse-to-fine synthesis. Experiments on the DeepFashion, MVC dataset and additional real-world datasets demonstrate that our approach compares favorably with the state-of-the-art methods and generalizes to unseen poses and clothing styles.
Character-independent font identification
Jan 24, 2020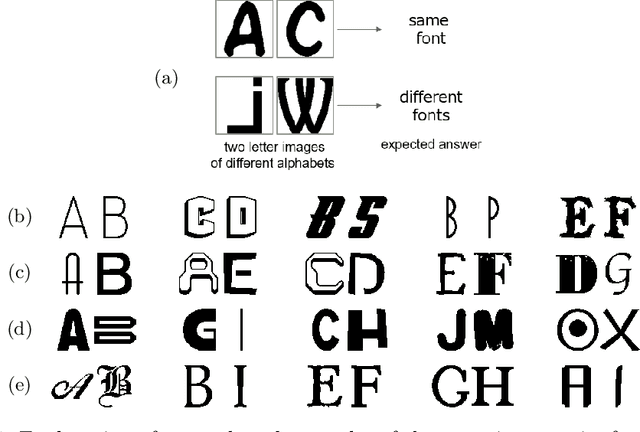
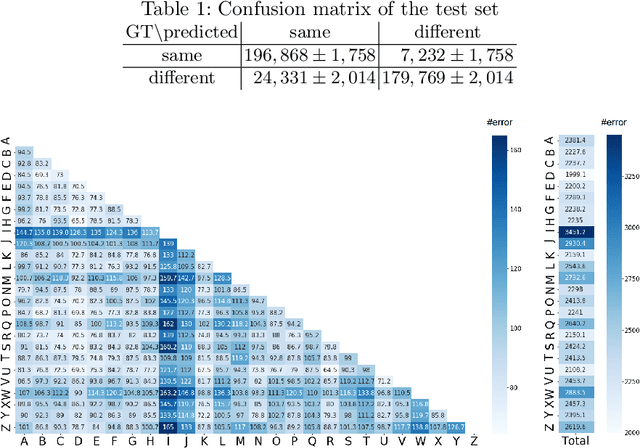
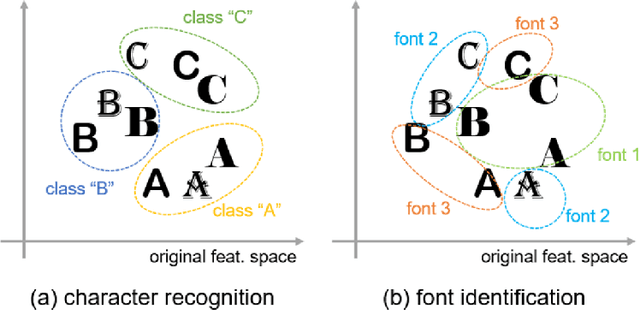
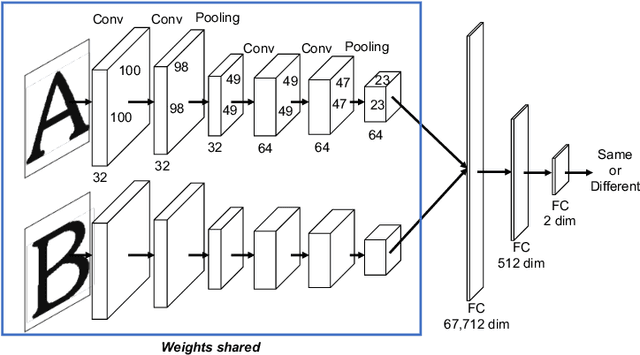
There are a countless number of fonts with various shapes and styles. In addition, there are many fonts that only have subtle differences in features. Due to this, font identification is a difficult task. In this paper, we propose a method of determining if any two characters are from the same font or not. This is difficult due to the difference between fonts typically being smaller than the difference between alphabet classes. Additionally, the proposed method can be used with fonts regardless of whether they exist in the training or not. In order to accomplish this, we use a Convolutional Neural Network (CNN) trained with various font image pairs. In the experiment, the network is trained on image pairs of various fonts. We then evaluate the model on a different set of fonts that are unseen by the network. The evaluation is performed with an accuracy of 92.27%. Moreover, we analyzed the relationship between character classes and font identification accuracy.
ODE-based Deep Network for MRI Reconstruction
Dec 27, 2019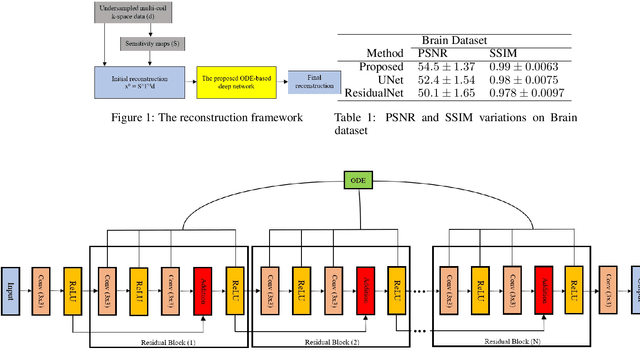
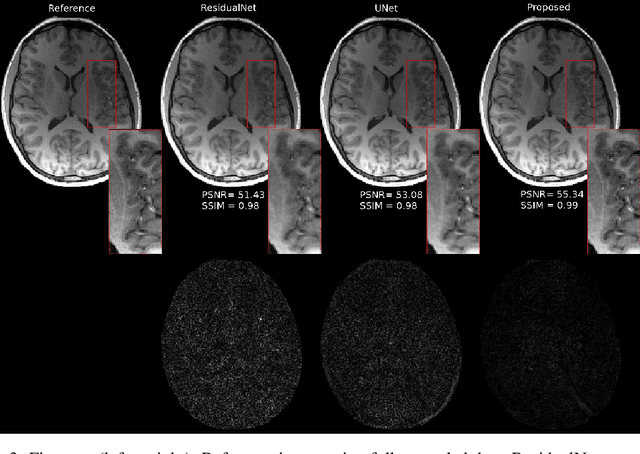
Fast data acquisition in Magnetic Resonance Imaging (MRI) is vastly in demand and scan time directly depends on the number of acquired k-space samples. The data-driven methods based on deep neural networks have resulted in promising improvements, compared to the conventional methods, in image reconstruction algorithms. The connection between deep neural network and Ordinary Differential Equation (ODE) has been observed and studied recently. The studies show that different residual networks can be interpreted as Euler discretization of an ODE. In this paper, we propose an ODE-based deep network for MRI reconstruction to enable the rapid acquisition of MR images with improved image quality. Our results with undersampled data demonstrate that our method can deliver higher quality images in comparison to the reconstruction methods based on the standard UNet network and Residual network.
M3D-GAN: Multi-Modal Multi-Domain Translation with Universal Attention
Jul 09, 2019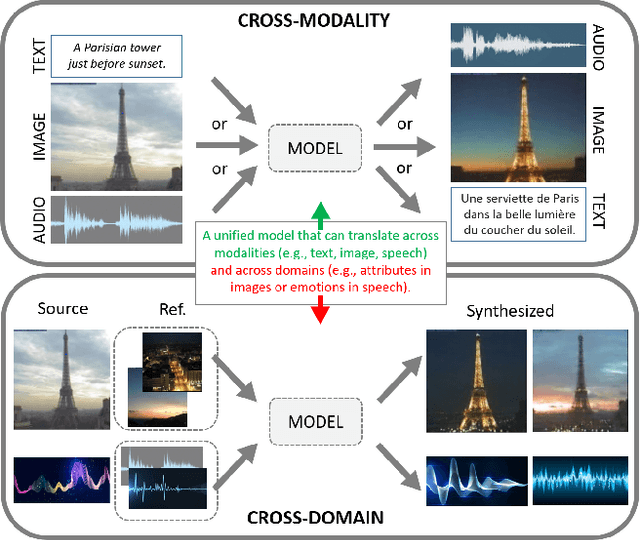
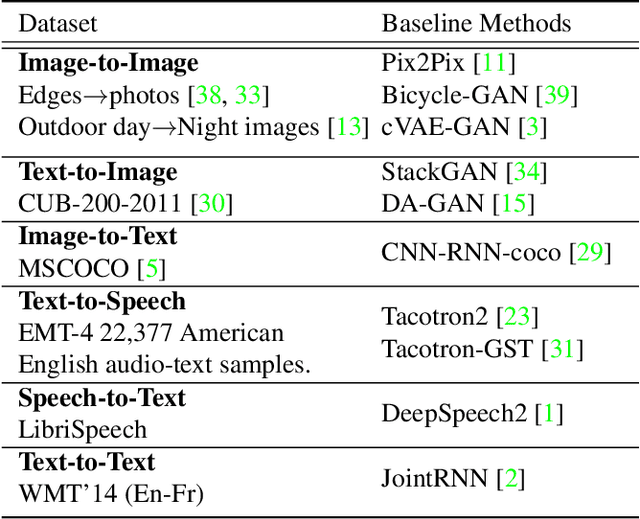

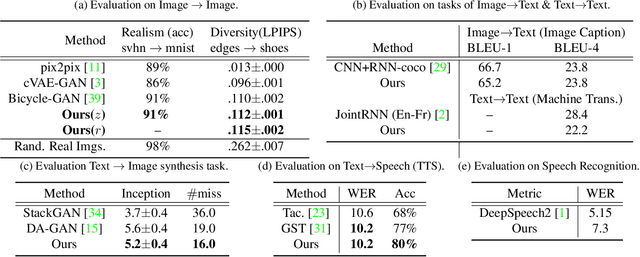
Generative adversarial networks have led to significant advances in cross-modal/domain translation. However, typically these networks are designed for a specific task (e.g., dialogue generation or image synthesis, but not both). We present a unified model, M3D-GAN, that can translate across a wide range of modalities (e.g., text, image, and speech) and domains (e.g., attributes in images or emotions in speech). Our model consists of modality subnets that convert data from different modalities into unified representations, and a unified computing body where data from different modalities share the same network architecture. We introduce a universal attention module that is jointly trained with the whole network and learns to encode a large range of domain information into a highly structured latent space. We use this to control synthesis in novel ways, such as producing diverse realistic pictures from a sketch or varying the emotion of synthesized speech. We evaluate our approach on extensive benchmark tasks, including image-to-image, text-to-image, image captioning, text-to-speech, speech recognition, and machine translation. Our results show state-of-the-art performance on some of the tasks.
Optimal input configuration of dynamic contrast enhanced MRI in convolutional neural networks for liver segmentation
Aug 22, 2019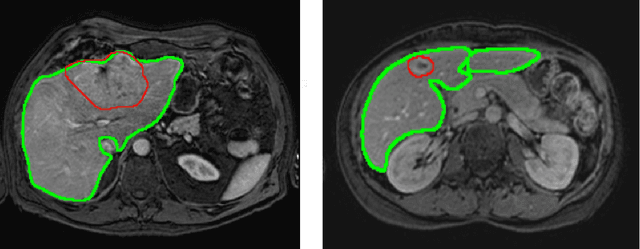
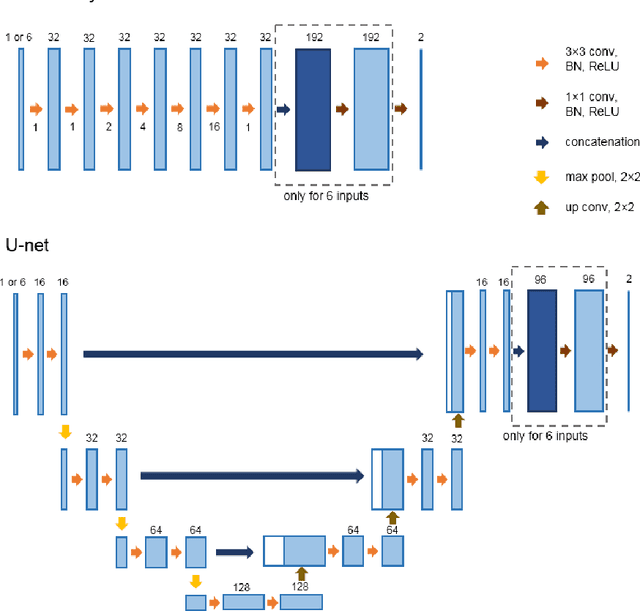
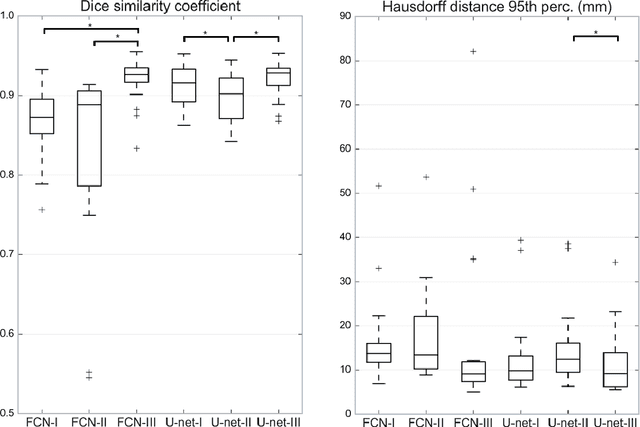
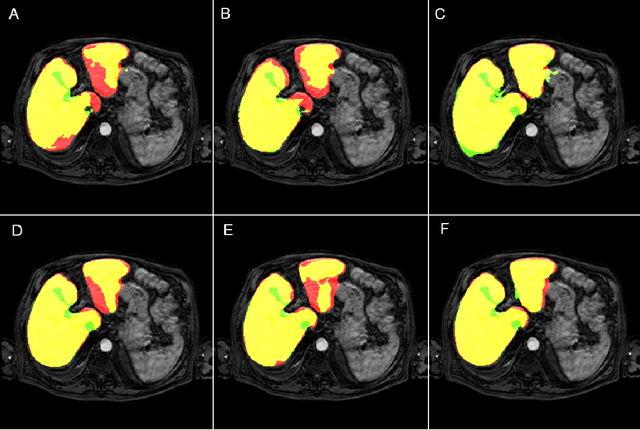
Most MRI liver segmentation methods use a structural 3D scan as input, such as a T1 or T2 weighted scan. Segmentation performance may be improved by utilizing both structural and functional information, as contained in dynamic contrast enhanced (DCE) MR series. Dynamic information can be incorporated in a segmentation method based on convolutional neural networks in a number of ways. In this study, the optimal input configuration of DCE MR images for convolutional neural networks (CNNs) is studied. The performance of three different input configurations for CNNs is studied for a liver segmentation task. The three configurations are I) one phase image of the DCE-MR series as input image; II) the separate phases of the DCE-MR as input images; and III) the separate phases of the DCE-MR as channels of one input image. The three input configurations are fed into a dilated fully convolutional network and into a small U-net. The CNNs were trained using 19 annotated DCE-MR series and tested on another 19 annotated DCE-MR series. The performance of the three input configurations for both networks is evaluated against manual annotations. The results show that both neural networks perform better when the separate phases of the DCE-MR series are used as channels of an input image in comparison to one phase as input image or the separate phases as input images. No significant difference between the performances of the two network architectures was found for the separate phases as channels of an input image.
A Deep Convolutional Neural Network for COVID-19 Detection Using Chest X-Rays
Apr 30, 2020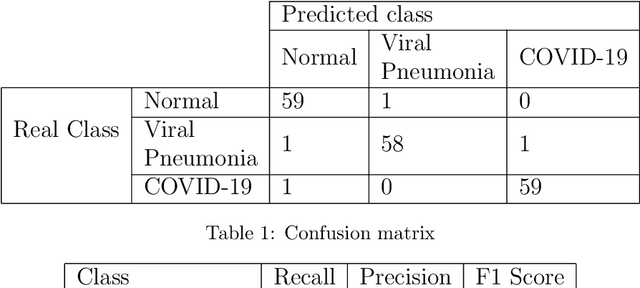
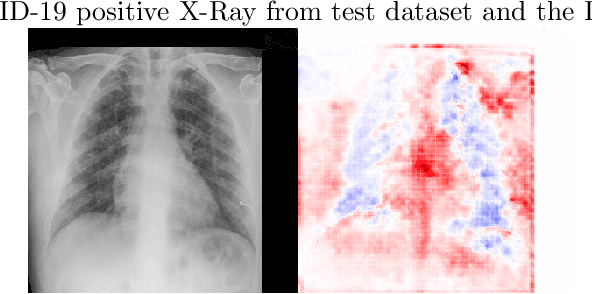
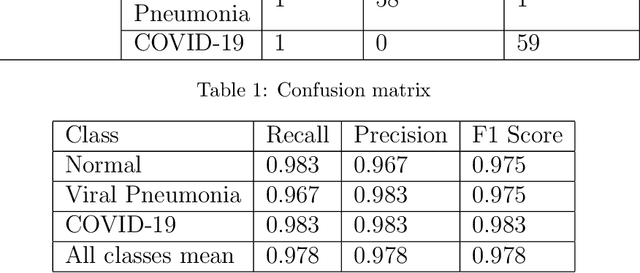
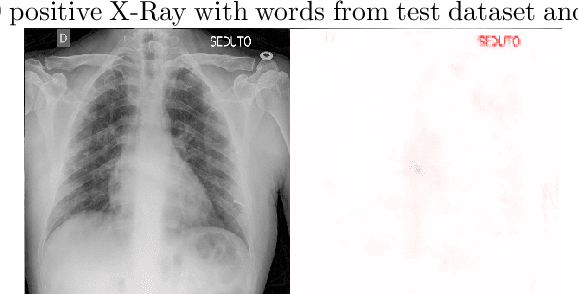
We present an image classifier based on the CheXNet and a transfer learning stage to classify chest X-Ray images according to three labels: COVID-19, viral pneumonia and normal. CheXNet is a DenseNet121 that has been trained twice, firstly on ImageNet and then, for classification of pneumonia and other 13 chest diseases, over a large chest X-Ray database (ChestX- ray14). The proposed network reached a test accuracy of 97.8% and, for the COVID-19 class, of 98.3%. In order to clarify the modus operandi of the network, we used Layer Wise Relevance Propagation (LRP) to generate heat maps, indicating an analytical path for future research on diagnosis.
SARS-CoV-2 virus RNA sequence classification and geographical analysis with convolutional neural networks approach
Jul 09, 2020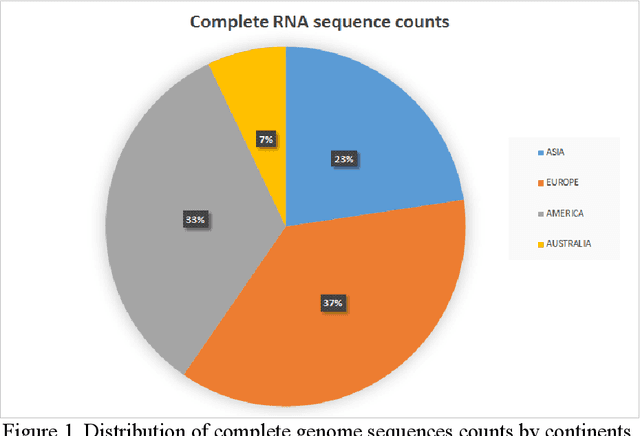

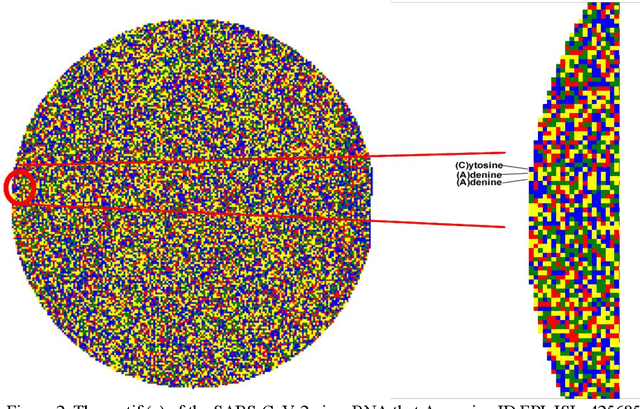
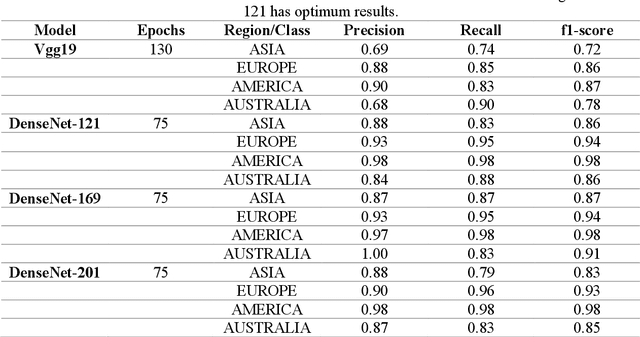
Covid-19 infection, which spread to the whole world in December 2019 and is still active, caused more than 250 thousand deaths in the world today. Researches on this subject have been focused on analyzing the genetic structure of the virus, developing vaccines, the course of the disease, and its source. In this study, RNA sequences belonging to the SARS-CoV-2 virus are transformed into gene motifs with two basic image processing algorithms and classified with the convolutional neural network (CNN) models. The CNN models achieved an average of 98% Area Under Curve(AUC) value was achieved in RNA sequences classified as Asia, Europe, America, and Oceania. The resulting artificial neural network model was used for phylogenetic analysis of the variant of the virus isolated in Turkey. The classification results reached were compared with gene alignment values in the GISAID database, where SARS-CoV-2 virus records are kept all over the world. Our experimental results have revealed that now the detection of the geographic distribution of the virus with the CNN models might serve as an efficient method.
Handwritten and Machine printed OCR for Geez Numbers Using Artificial Neural Network
Nov 15, 2019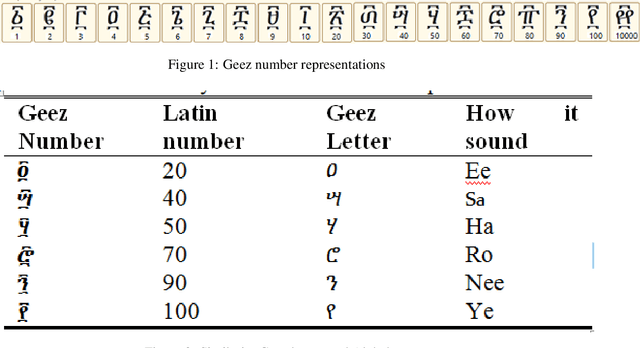

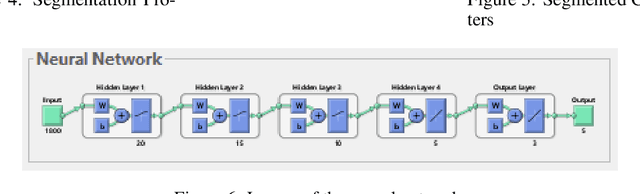
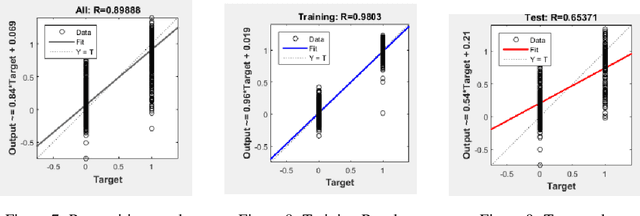
Researches have been done on Ethiopic scripts. However studies excluded the Geez numbers from the studies because of different reasons. This paper presents offline handwritten and machine printed Geez number recognition using feed forward back propagation artificial neural network. On this study, different Geez image characters were collected from google image search and three persons are instructed to write the numbers using pencil. In total we have collected 560 numbers of characters. We have used 460 of the characters for training and 100 are used for testing. Accordingly we have achieved overall all classification ~89:88%
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge