"Image": models, code, and papers
Binary Document Image Super Resolution for Improved Readability and OCR Performance
Dec 06, 2018

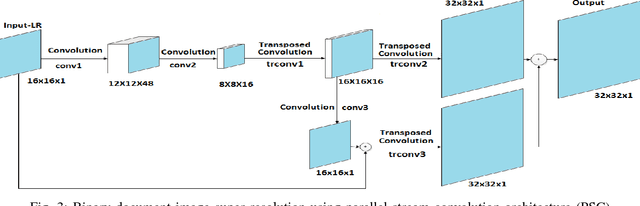
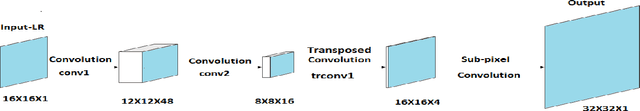
There is a need for information retrieval from large collections of low-resolution (LR) binary document images, which can be found in digital libraries across the world, where the high-resolution (HR) counterpart is not available. This gives rise to the problem of binary document image super-resolution (BDISR). The objective of this paper is to address the interesting and challenging problem of super resolution of binary Tamil document images for improved readability and better optical character recognition (OCR). We propose multiple deep neural network architectures to address this problem and analyze their performance. The proposed models are all single image super-resolution techniques, which learn a generalized spatial correspondence between the LR and HR binary document images. We employ convolutional layers for feature extraction followed by transposed convolution and sub-pixel convolution layers for upscaling the features. Since the outputs of the neural networks are gray scale, we utilize the advantage of power law transformation as a post-processing technique to improve the character level pixel connectivity. The performance of our models is evaluated by comparing the OCR accuracies and the mean opinion scores given by human evaluators on LR images and the corresponding model-generated HR images.
Robust Segmentation of Optic Disc and Cup from Fundus Images Using Deep Neural Networks
Dec 13, 2020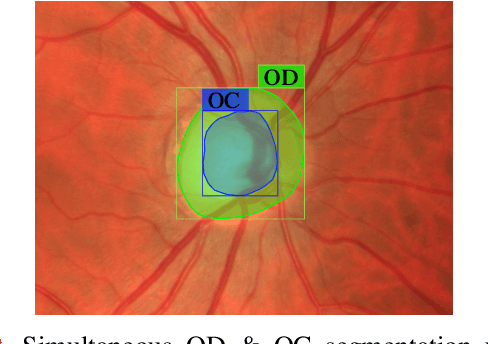
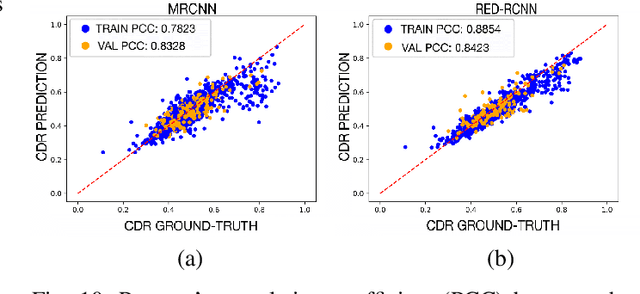
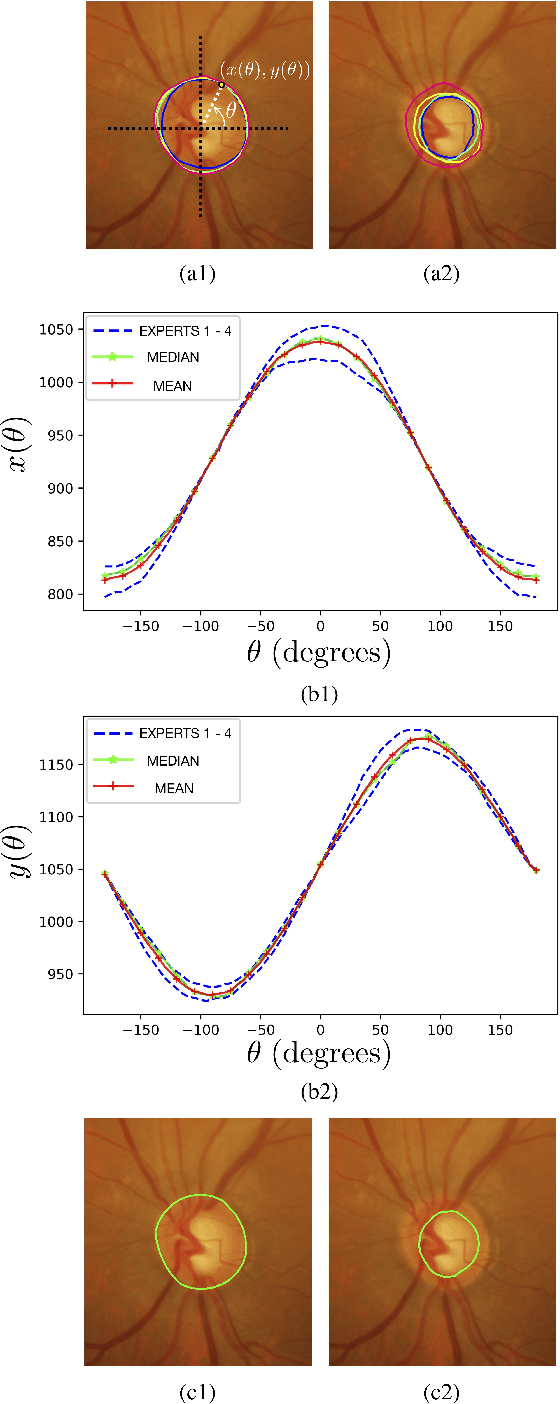
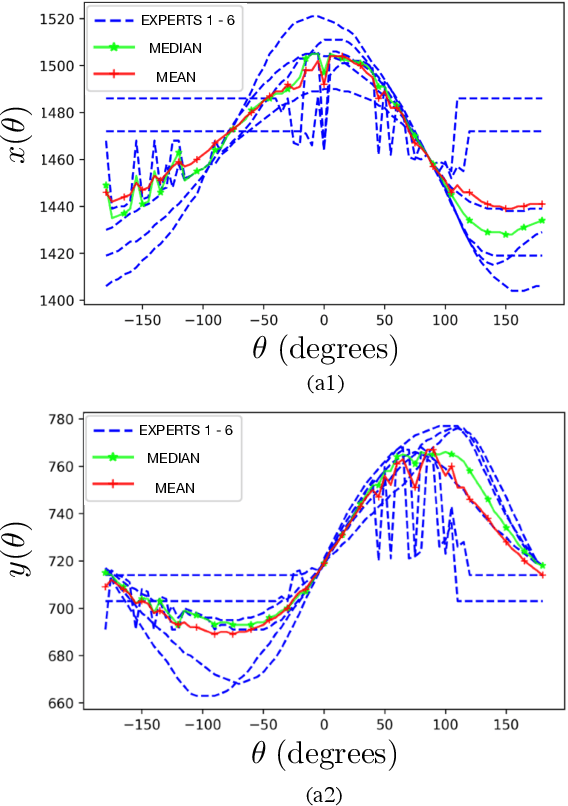
Optic disc (OD) and optic cup (OC) are regions of prominent clinical interest in a retinal fundus image. They are the primary indicators of a glaucomatous condition. With the advent and success of deep learning for healthcare research, several approaches have been proposed for the segmentation of important features in retinal fundus images. We propose a novel approach for the simultaneous segmentation of the OD and OC using a residual encoder-decoder network (REDNet) based regional convolutional neural network (RCNN). The RED-RCNN is motivated by the Mask RCNN (MRCNN). Performance comparisons with the state-of-the-art techniques and extensive validations on standard publicly available fundus image datasets show that RED-RCNN has superior performance compared with MRCNN. RED-RCNN results in Sensitivity, Specificity, Accuracy, Precision, Dice and Jaccard indices of 95.64%, 99.9%, 99.82%, 95.68%, 95.64%, 91.65%, respectively, for OD segmentation, and 91.44%, 99.87%, 99.83%, 85.67%, 87.48%, 78.09%, respectively, for OC segmentation. Further, we perform two-stage glaucoma severity grading using the cup-to-disc ratio (CDR) computed based on the obtained OD/OC segmentation. The superior segmentation performance of RED-RCNN over MRCNN translates to higher accuracy in glaucoma severity grading.
Lookahead optimizer improves the performance of Convolutional Autoencoders for reconstruction of natural images
Dec 03, 2020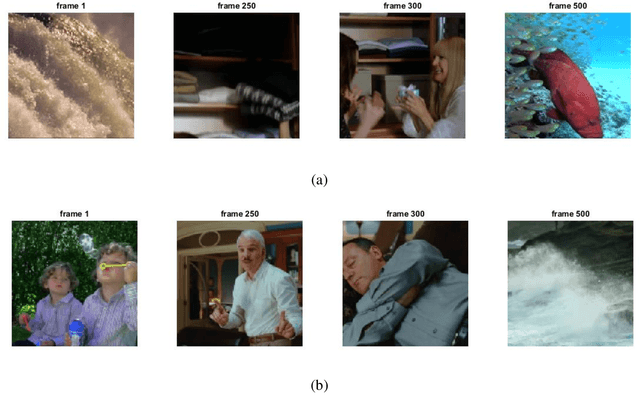
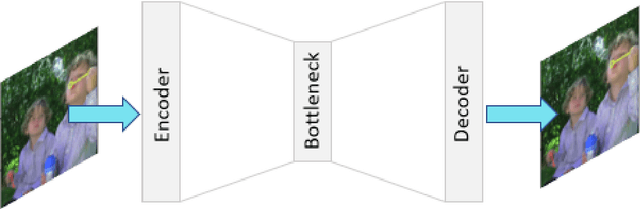
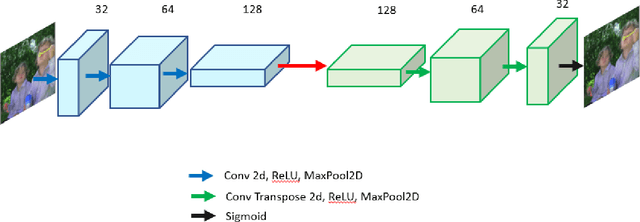
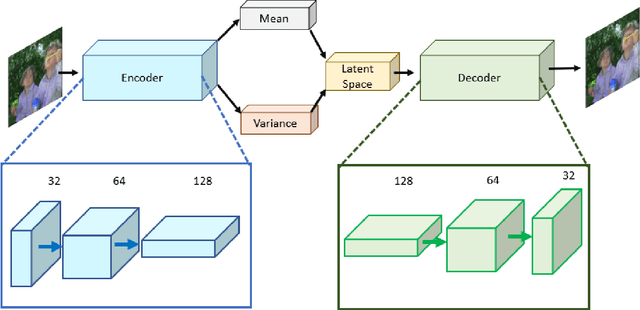
Autoencoders are a class of artificial neural networks which have gained a lot of attention in the recent past. Using the encoder block of an autoencoder the input image can be compressed into a meaningful representation. Then a decoder is employed to reconstruct the compressed representation back to a version which looks like the input image. It has plenty of applications in the field of data compression and denoising. Another version of Autoencoders (AE) exist, called Variational AE (VAE) which acts as a generative model like GAN. Recently, an optimizer was introduced which is known as lookahead optimizer which significantly enhances the performances of Adam as well as SGD. In this paper, we implement Convolutional Autoencoders (CAE) and Convolutional Variational Autoencoders (CVAE) with lookahead optimizer (with Adam) and compare them with the Adam (only) optimizer counterparts. For this purpose, we have used a movie dataset comprising of natural images for the former case and CIFAR100 for the latter case. We show that lookahead optimizer (with Adam) improves the performance of CAEs for reconstruction of natural images.
Fully Automatic Wound Segmentation with Deep Convolutional Neural Networks
Oct 12, 2020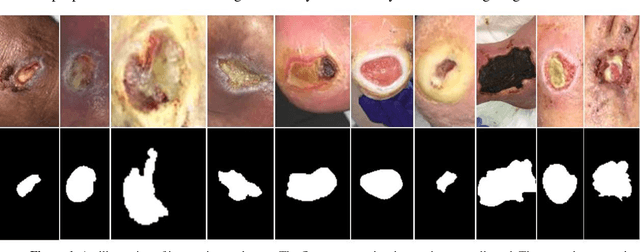
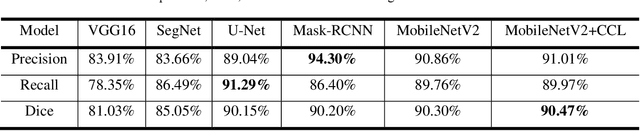
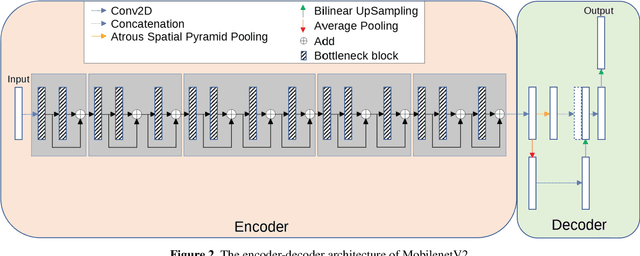
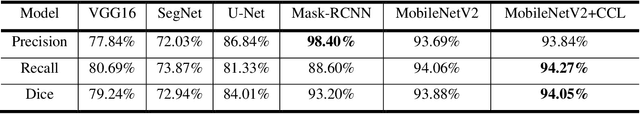
Acute and chronic wounds have varying etiologies and are an economic burden to healthcare systems around the world. The advanced wound care market is expected to exceed $22 billion by 2024. Wound care professionals rely heavily on images and image documentation for proper diagnosis and treatment. Unfortunately lack of expertise can lead to improper diagnosis of wound etiology and inaccurate wound management and documentation. Fully automatic segmentation of wound areas in natural images is an important part of the diagnosis and care protocol since it is crucial to measure the area of the wound and provide quantitative parameters in the treatment. Various deep learning models have gained success in image analysis including semantic segmentation. Particularly, MobileNetV2 stands out among others due to its lightweight architecture and uncompromised performance. This manuscript proposes a novel convolutional framework based on MobileNetV2 and connected component labelling to segment wound regions from natural images. We build an annotated wound image dataset consisting of 1,109 foot ulcer images from 889 patients to train and test the deep learning models. We demonstrate the effectiveness and mobility of our method by conducting comprehensive experiments and analyses on various segmentation neural networks.
Learning to Transfer Visual Effects from Videos to Images
Dec 03, 2020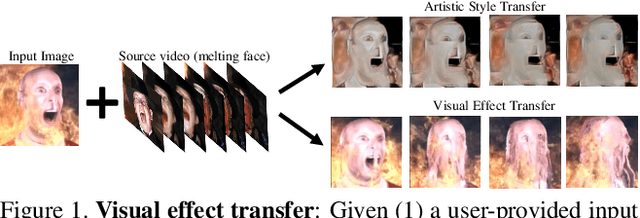
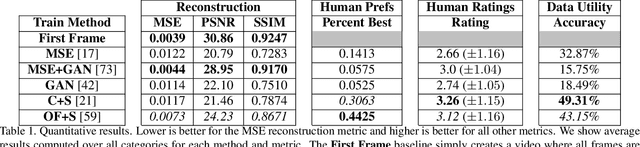
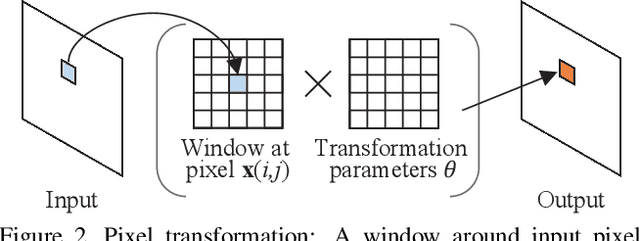
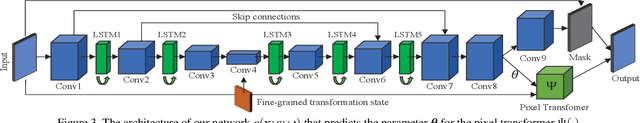
We study the problem of animating images by transferring spatio-temporal visual effects (such as melting) from a collection of videos. We tackle two primary challenges in visual effect transfer: 1) how to capture the effect we wish to distill; and 2) how to ensure that only the effect, rather than content or artistic style, is transferred from the source videos to the input image. To address the first challenge, we evaluate five loss functions; the most promising one encourages the generated animations to have similar optical flow and texture motions as the source videos. To address the second challenge, we only allow our model to move existing image pixels from the previous frame, rather than predicting unconstrained pixel values. This forces any visual effects to occur using the input image's pixels, preventing unwanted artistic style or content from the source video from appearing in the output. We evaluate our method in objective and subjective settings, and show interesting qualitative results which demonstrate objects undergoing atypical transformations, such as making a face melt or a deer bloom.
Evaluation of Multi-Slice Inputs to Convolutional Neural Networks for Medical Image Segmentation
Dec 22, 2019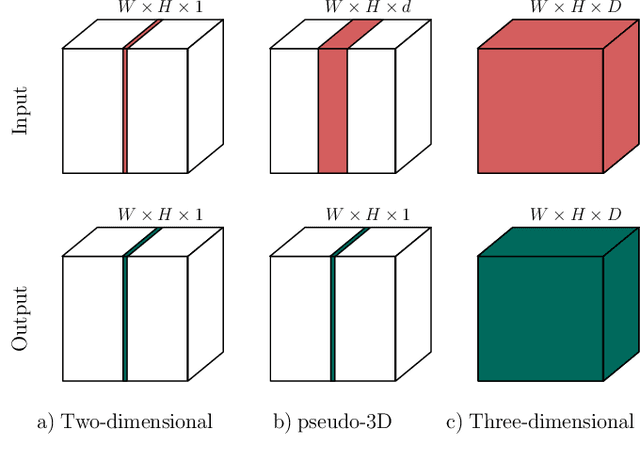
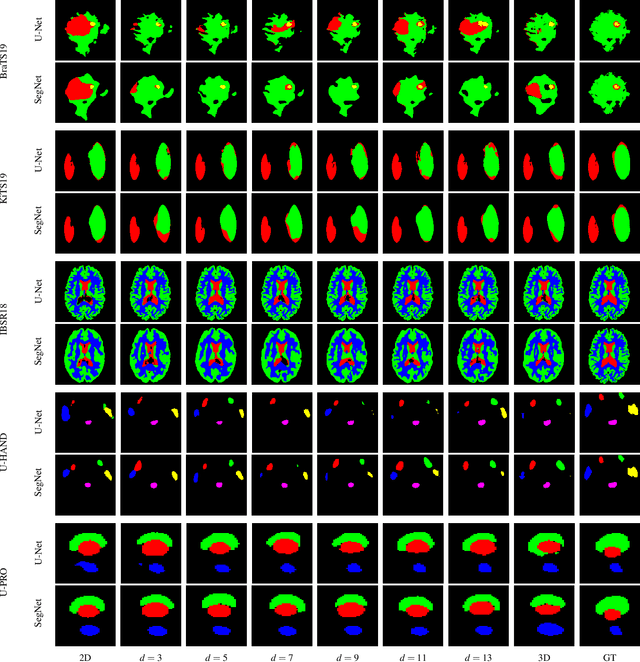
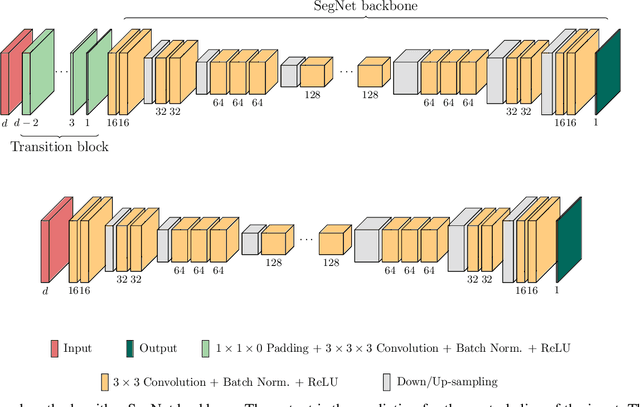
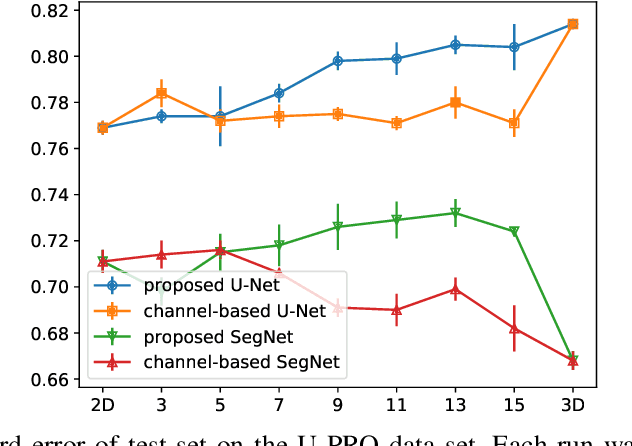
When using Convolutional Neural Networks (CNNs) for segmentation of organs and lesions in medical images, the conventional approach is to work with inputs and outputs either as single slice (2D) or whole volumes (3D). One common alternative, in this study denoted as pseudo-3D, is to use a stack of adjacent slices as input and produce a prediction for at least the central slice. This approach gives the network the possibility to capture 3D spatial information, with only a minor additional computational cost. In this study, we systematically evaluate the segmentation performance and computational costs of this pseudo-3D approach as a function of the number of input slices, and compare the results to conventional end-to-end 2D and 3D CNNs. The standard pseudo-3D method regards the neighboring slices as multiple input image channels. We additionally evaluate a simple approach where the input stack is a volumetric input that is repeatably convolved in 3D to obtain a 2D feature map. This 2D map is in turn fed into a standard 2D network. We conducted experiments using two different CNN backbone architectures and on five diverse data sets covering different anatomical regions, imaging modalities, and segmentation tasks. We found that while both pseudo-3D methods can process a large number of slices at once and still be computationally much more efficient than fully 3D CNNs, a significant improvement over a regular 2D CNN was only observed for one of the five data sets. An analysis of the structural properties of the segmentation masks revealed no relations to the segmentation performance with respect to the number of input slices. The conclusion is therefore that in the general case, multi-slice inputs appear to not significantly improve segmentation results over using 2D or 3D CNNs.
Holistic 3D Human and Scene Mesh Estimation from Single View Images
Dec 02, 2020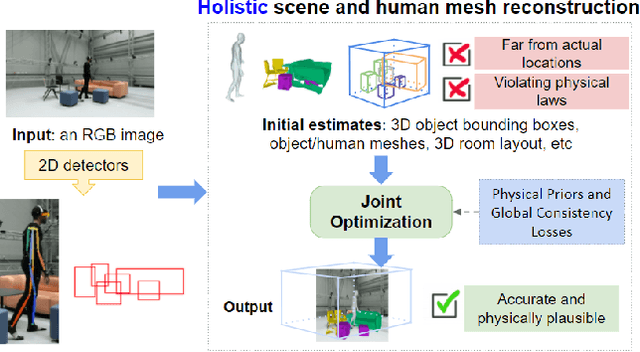
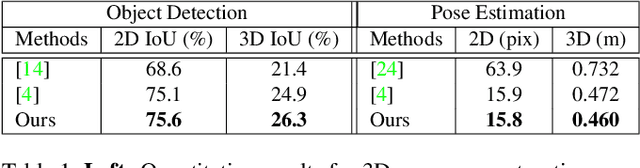
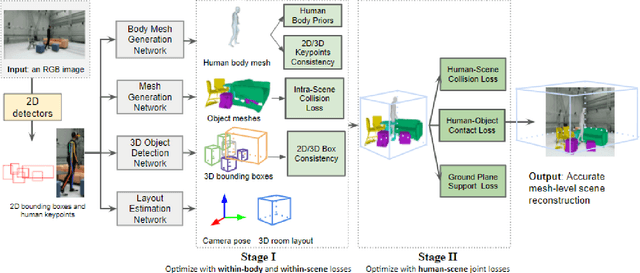
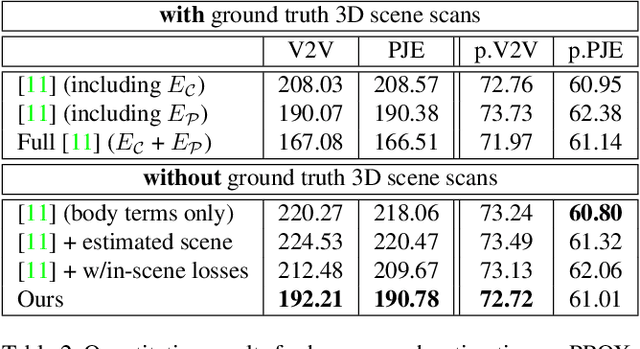
The 3D world limits the human body pose and the human body pose conveys information about the surrounding objects. Indeed, from a single image of a person placed in an indoor scene, we as humans are adept at resolving ambiguities of the human pose and room layout through our knowledge of the physical laws and prior perception of the plausible object and human poses. However, few computer vision models fully leverage this fact. In this work, we propose an end-to-end trainable model that perceives the 3D scene from a single RGB image, estimates the camera pose and the room layout, and reconstructs both human body and object meshes. By imposing a set of comprehensive and sophisticated losses on all aspects of the estimations, we show that our model outperforms existing human body mesh methods and indoor scene reconstruction methods. To the best of our knowledge, this is the first model that outputs both object and human predictions at the mesh level, and performs joint optimization on the scene and human poses.
Impact of Facial Tattoos and Paintings on Face Recognition Systems
Mar 17, 2021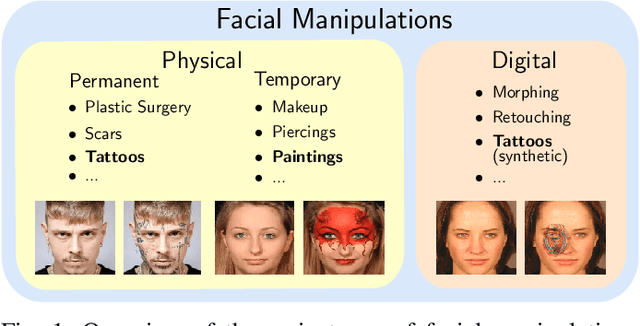
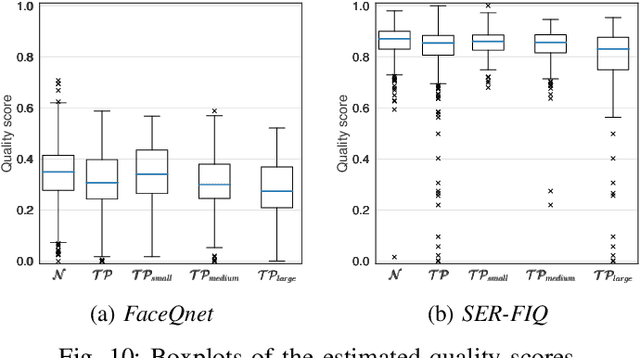
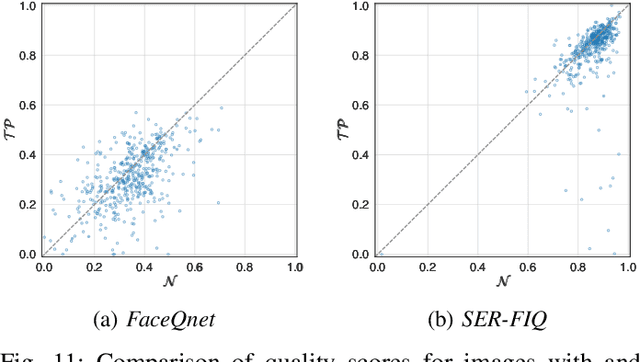

In the past years, face recognition technologies have shown impressive recognition performance, mainly due to recent developments in deep convolutional neural networks. Notwithstanding those improvements, several challenges which affect the performance of face recognition systems remain. In this work, we investigate the impact that facial tattoos and paintings have on current face recognition systems. To this end, we first collected an appropriate database containing image-pairs of individuals with and without facial tattoos or paintings. The assembled database was used to evaluate how facial tattoos and paintings affect the detection, quality estimation, as well as the feature extraction and comparison modules of a face recognition system. The impact on these modules was evaluated using state-of-the-art open-source and commercial systems. The obtained results show that facial tattoos and paintings affect all the tested modules, especially for images where a large area of the face is covered with tattoos or paintings. Our work is an initial case-study and indicates a need to design algorithms which are robust to the visual changes caused by facial tattoos and paintings.
Efficient, high-performance pancreatic segmentation using multi-scale feature extraction
Sep 02, 2020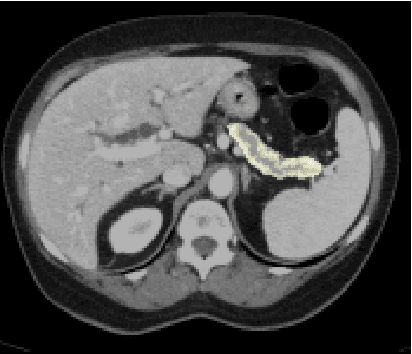
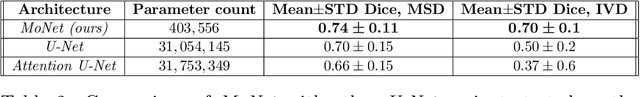

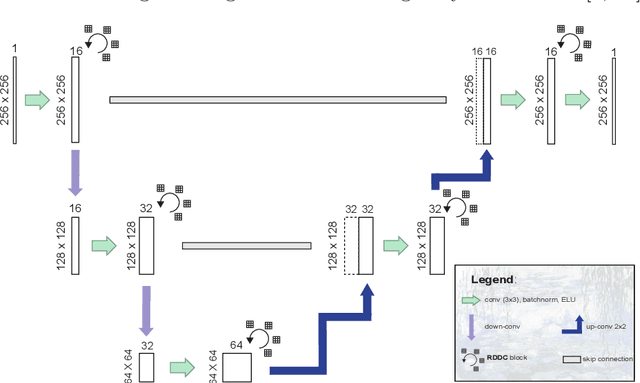

For artificial intelligence-based image analysis methods to reach clinical applicability, the development of high-performance algorithms is crucial. For example, existent segmentation algorithms based on natural images are neither efficient in their parameter use nor optimized for medical imaging. Here we present MoNet, a highly optimized neural-network-based pancreatic segmentation algorithm focused on achieving high performance by efficient multi-scale image feature utilization.
A Unified Framework for Generic, Query-Focused, Privacy Preserving and Update Summarization using Submodular Information Measures
Oct 12, 2020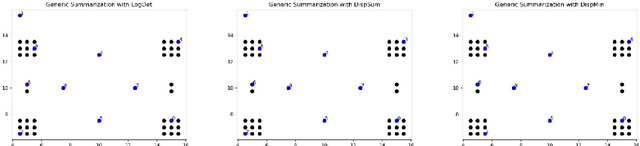
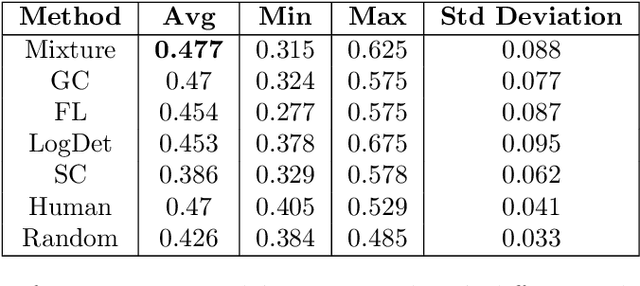
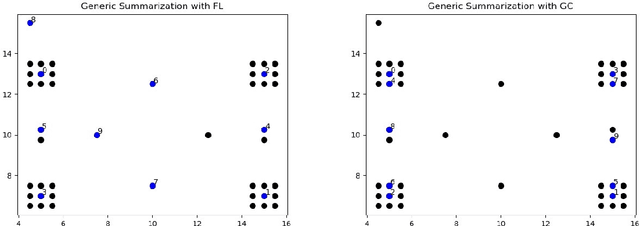
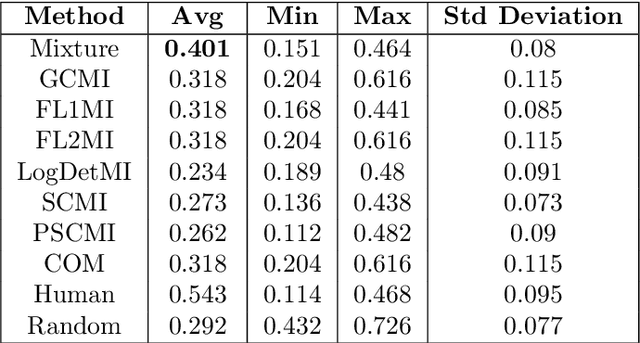
We study submodular information measures as a rich framework for generic, query-focused, privacy sensitive, and update summarization tasks. While past work generally treats these problems differently ({\em e.g.}, different models are often used for generic and query-focused summarization), the submodular information measures allow us to study each of these problems via a unified approach. We first show that several previous query-focused and update summarization techniques have, unknowingly, used various instantiations of the aforesaid submodular information measures, providing evidence for the benefit and naturalness of these models. We then carefully study and demonstrate the modelling capabilities of the proposed functions in different settings and empirically verify our findings on both a synthetic dataset and an existing real-world image collection dataset (that has been extended by adding concept annotations to each image making it suitable for this task) and will be publicly released. We employ a max-margin framework to learn a mixture model built using the proposed instantiations of submodular information measures and demonstrate the effectiveness of our approach. While our experiments are in the context of image summarization, our framework is generic and can be easily extended to other summarization settings (e.g., videos or documents).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge