cancer detection
Cancer detection using Artificial Intelligence (AI) involves leveraging advanced machine learning algorithms and techniques to identify and diagnose cancer from various medical data sources. The goal is to enhance early detection, improve diagnostic accuracy, and potentially reduce the need for invasive procedures.
Papers and Code
Mitigating Multi-Sequence 3D Prostate MRI Data Scarcity through Domain Adaptation using Locally-Trained Latent Diffusion Models for Prostate Cancer Detection
Jul 08, 2025Objective: Latent diffusion models (LDMs) could mitigate data scarcity challenges affecting machine learning development for medical image interpretation. The recent CCELLA LDM improved prostate cancer detection performance using synthetic MRI for classifier training but was limited to the axial T2-weighted (AxT2) sequence, did not investigate inter-institutional domain shift, and prioritized radiology over histopathology outcomes. We propose CCELLA++ to address these limitations and improve clinical utility. Methods: CCELLA++ expands CCELLA for simultaneous biparametric prostate MRI (bpMRI) generation, including the AxT2, high b-value diffusion series (HighB) and apparent diffusion coefficient map (ADC). Domain adaptation was investigated by pretraining classifiers on real or LDM-generated synthetic data from an internal institution, followed with fine-tuning on progressively smaller fractions of an out-of-distribution, external dataset. Results: CCELLA++ improved 3D FID for HighB and ADC but not AxT2 (0.013, 0.012, 0.063 respectively) sequences compared to CCELLA (0.060). Classifier pretraining with CCELLA++ bpMRI outperformed real bpMRI in AP and AUC for all domain adaptation scenarios. CCELLA++ pretraining achieved highest classifier performance below 50% (n=665) external dataset volume. Conclusion: Synthetic bpMRI generated by our method can improve downstream classifier generalization and performance beyond real bpMRI or CCELLA-generated AxT2-only images. Future work should seek to quantify medical image sample quality, balance multi-sequence LDM training, and condition the LDM with additional information. Significance: The proposed CCELLA++ LDM can generate synthetic bpMRI that outperforms real data for domain adaptation with a limited target institution dataset. Our code is available at https://github.com/grabkeem/CCELLA-plus-plus
Synthetic Data-Driven Multi-Architecture Framework for Automated Polyp Segmentation Through Integrated Detection and Mask Generation
Aug 08, 2025Colonoscopy is a vital tool for the early diagnosis of colorectal cancer, which is one of the main causes of cancer-related mortality globally; hence, it is deemed an essential technique for the prevention and early detection of colorectal cancer. The research introduces a unique multidirectional architectural framework to automate polyp detection within colonoscopy images while helping resolve limited healthcare dataset sizes and annotation complexities. The research implements a comprehensive system that delivers synthetic data generation through Stable Diffusion enhancements together with detection and segmentation algorithms. This detection approach combines Faster R-CNN for initial object localization while the Segment Anything Model (SAM) refines the segmentation masks. The faster R-CNN detection algorithm achieved a recall of 93.08% combined with a precision of 88.97% and an F1 score of 90.98%.SAM is then used to generate the image mask. The research evaluated five state-of-the-art segmentation models that included U-Net, PSPNet, FPN, LinkNet, and MANet using ResNet34 as a base model. The results demonstrate the superior performance of FPN with the highest scores of PSNR (7.205893) and SSIM (0.492381), while UNet excels in recall (84.85%) and LinkNet shows balanced performance in IoU (64.20%) and Dice score (77.53%).
Model Compression Engine for Wearable Devices Skin Cancer Diagnosis
Jul 23, 2025Skin cancer is one of the most prevalent and preventable types of cancer, yet its early detection remains a challenge, particularly in resource-limited settings where access to specialized healthcare is scarce. This study proposes an AI-driven diagnostic tool optimized for embedded systems to address this gap. Using transfer learning with the MobileNetV2 architecture, the model was adapted for binary classification of skin lesions into "Skin Cancer" and "Other." The TensorRT framework was employed to compress and optimize the model for deployment on the NVIDIA Jetson Orin Nano, balancing performance with energy efficiency. Comprehensive evaluations were conducted across multiple benchmarks, including model size, inference speed, throughput, and power consumption. The optimized models maintained their performance, achieving an F1-Score of 87.18% with a precision of 93.18% and recall of 81.91%. Post-compression results showed reductions in model size of up to 0.41, along with improvements in inference speed and throughput, and a decrease in energy consumption of up to 0.93 in INT8 precision. These findings validate the feasibility of deploying high-performing, energy-efficient diagnostic tools on resource-constrained edge devices. Beyond skin cancer detection, the methodologies applied in this research have broader applications in other medical diagnostics and domains requiring accessible, efficient AI solutions. This study underscores the potential of optimized AI systems to revolutionize healthcare diagnostics, thereby bridging the divide between advanced technology and underserved regions.
OCELOT 2023: Cell Detection from Cell-Tissue Interaction Challenge
Sep 11, 2025Pathologists routinely alternate between different magnifications when examining Whole-Slide Images, allowing them to evaluate both broad tissue morphology and intricate cellular details to form comprehensive diagnoses. However, existing deep learning-based cell detection models struggle to replicate these behaviors and learn the interdependent semantics between structures at different magnifications. A key barrier in the field is the lack of datasets with multi-scale overlapping cell and tissue annotations. The OCELOT 2023 challenge was initiated to gather insights from the community to validate the hypothesis that understanding cell and tissue (cell-tissue) interactions is crucial for achieving human-level performance, and to accelerate the research in this field. The challenge dataset includes overlapping cell detection and tissue segmentation annotations from six organs, comprising 673 pairs sourced from 306 The Cancer Genome Atlas (TCGA) Whole-Slide Images with hematoxylin and eosin staining, divided into training, validation, and test subsets. Participants presented models that significantly enhanced the understanding of cell-tissue relationships. Top entries achieved up to a 7.99 increase in F1-score on the test set compared to the baseline cell-only model that did not incorporate cell-tissue relationships. This is a substantial improvement in performance over traditional cell-only detection methods, demonstrating the need for incorporating multi-scale semantics into the models. This paper provides a comparative analysis of the methods used by participants, highlighting innovative strategies implemented in the OCELOT 2023 challenge.
* This is the accepted manuscript of an article published in Medical Image Analysis (Elsevier). The final version is available at: https://doi.org/10.1016/j.media.2025.103751
Improving Diagnostic Accuracy for Oral Cancer with inpainting Synthesis Lesions Generated Using Diffusion Models
Aug 08, 2025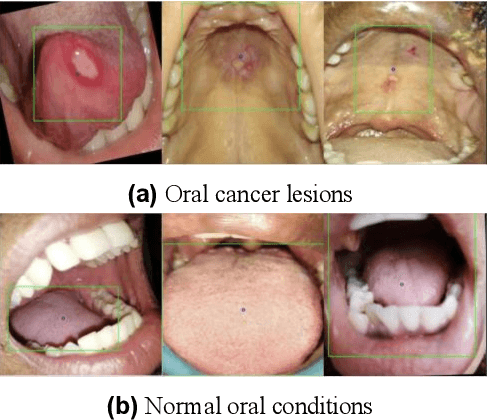
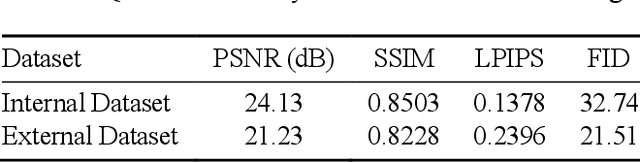
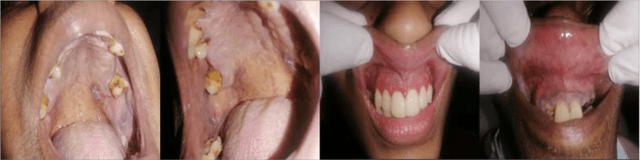
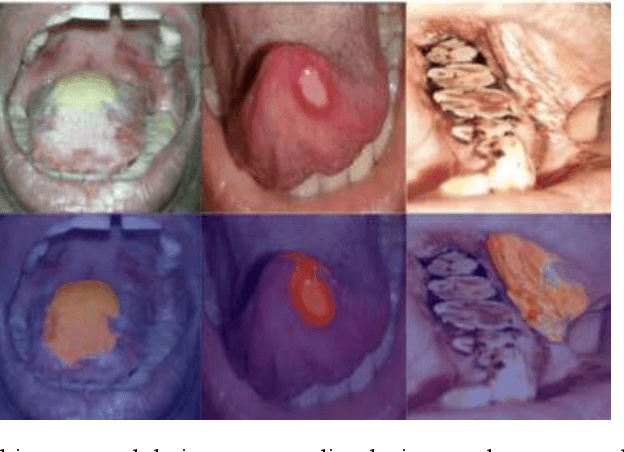
In oral cancer diagnostics, the limited availability of annotated datasets frequently constrains the performance of diagnostic models, particularly due to the variability and insufficiency of training data. To address these challenges, this study proposed a novel approach to enhance diagnostic accuracy by synthesizing realistic oral cancer lesions using an inpainting technique with a fine-tuned diffusion model. We compiled a comprehensive dataset from multiple sources, featuring a variety of oral cancer images. Our method generated synthetic lesions that exhibit a high degree of visual fidelity to actual lesions, thereby significantly enhancing the performance of diagnostic algorithms. The results show that our classification model achieved a diagnostic accuracy of 0.97 in differentiating between cancerous and non-cancerous tissues, while our detection model accurately identified lesion locations with 0.85 accuracy. This method validates the potential for synthetic image generation in medical diagnostics and paves the way for further research into extending these methods to other types of cancer diagnostics.
A computationally frugal open-source foundation model for thoracic disease detection in lung cancer screening programs
Jul 02, 2025Low-dose computed tomography (LDCT) imaging employed in lung cancer screening (LCS) programs is increasing in uptake worldwide. LCS programs herald a generational opportunity to simultaneously detect cancer and non-cancer-related early-stage lung disease. Yet these efforts are hampered by a shortage of radiologists to interpret scans at scale. Here, we present TANGERINE, a computationally frugal, open-source vision foundation model for volumetric LDCT analysis. Designed for broad accessibility and rapid adaptation, TANGERINE can be fine-tuned off the shelf for a wide range of disease-specific tasks with limited computational resources and training data. Relative to models trained from scratch, TANGERINE demonstrates fast convergence during fine-tuning, thereby requiring significantly fewer GPU hours, and displays strong label efficiency, achieving comparable or superior performance with a fraction of fine-tuning data. Pretrained using self-supervised learning on over 98,000 thoracic LDCTs, including the UK's largest LCS initiative to date and 27 public datasets, TANGERINE achieves state-of-the-art performance across 14 disease classification tasks, including lung cancer and multiple respiratory diseases, while generalising robustly across diverse clinical centres. By extending a masked autoencoder framework to 3D imaging, TANGERINE offers a scalable solution for LDCT analysis, departing from recent closed, resource-intensive models by combining architectural simplicity, public availability, and modest computational requirements. Its accessible, open-source lightweight design lays the foundation for rapid integration into next-generation medical imaging tools that could transform LCS initiatives, allowing them to pivot from a singular focus on lung cancer detection to comprehensive respiratory disease management in high-risk populations.
A Hybrid CNN-VSSM model for Multi-View, Multi-Task Mammography Analysis: Robust Diagnosis with Attention-Based Fusion
Jul 22, 2025Early and accurate interpretation of screening mammograms is essential for effective breast cancer detection, yet it remains a complex challenge due to subtle imaging findings and diagnostic ambiguity. Many existing AI approaches fall short by focusing on single view inputs or single-task outputs, limiting their clinical utility. To address these limitations, we propose a novel multi-view, multitask hybrid deep learning framework that processes all four standard mammography views and jointly predicts diagnostic labels and BI-RADS scores for each breast. Our architecture integrates a hybrid CNN VSSM backbone, combining convolutional encoders for rich local feature extraction with Visual State Space Models (VSSMs) to capture global contextual dependencies. To improve robustness and interpretability, we incorporate a gated attention-based fusion module that dynamically weights information across views, effectively handling cases with missing data. We conduct extensive experiments across diagnostic tasks of varying complexity, benchmarking our proposed hybrid models against baseline CNN architectures and VSSM models in both single task and multi task learning settings. Across all tasks, the hybrid models consistently outperform the baselines. In the binary BI-RADS 1 vs. 5 classification task, the shared hybrid model achieves an AUC of 0.9967 and an F1 score of 0.9830. For the more challenging ternary classification, it attains an F1 score of 0.7790, while in the five-class BI-RADS task, the best F1 score reaches 0.4904. These results highlight the effectiveness of the proposed hybrid framework and underscore both the potential and limitations of multitask learning for improving diagnostic performance and enabling clinically meaningful mammography analysis.
Attention-Enhanced Deep Learning Ensemble for Breast Density Classification in Mammography
Jul 08, 2025Breast density assessment is a crucial component of mammographic interpretation, with high breast density (BI-RADS categories C and D) representing both a significant risk factor for developing breast cancer and a technical challenge for tumor detection. This study proposes an automated deep learning system for robust binary classification of breast density (low: A/B vs. high: C/D) using the VinDr-Mammo dataset. We implemented and compared four advanced convolutional neural networks: ResNet18, ResNet50, EfficientNet-B0, and DenseNet121, each enhanced with channel attention mechanisms. To address the inherent class imbalance, we developed a novel Combined Focal Label Smoothing Loss function that integrates focal loss, label smoothing, and class-balanced weighting. Our preprocessing pipeline incorporated advanced techniques, including contrast-limited adaptive histogram equalization (CLAHE) and comprehensive data augmentation. The individual models were combined through an optimized ensemble voting approach, achieving superior performance (AUC: 0.963, F1-score: 0.952) compared to any single model. This system demonstrates significant potential to standardize density assessments in clinical practice, potentially improving screening efficiency and early cancer detection rates while reducing inter-observer variability among radiologists.
MyGO: Make your Goals Obvious, Avoiding Semantic Confusion in Prostate Cancer Lesion Region Segmentation
Jul 23, 2025Early diagnosis and accurate identification of lesion location and progression in prostate cancer (PCa) are critical for assisting clinicians in formulating effective treatment strategies. However, due to the high semantic homogeneity between lesion and non-lesion areas, existing medical image segmentation methods often struggle to accurately comprehend lesion semantics, resulting in the problem of semantic confusion. To address this challenge, we propose a novel Pixel Anchor Module, which guides the model to discover a sparse set of feature anchors that serve to capture and interpret global contextual information. This mechanism enhances the model's nonlinear representation capacity and improves segmentation accuracy within lesion regions. Moreover, we design a self-attention-based Top_k selection strategy to further refine the identification of these feature anchors, and incorporate a focal loss function to mitigate class imbalance, thereby facilitating more precise semantic interpretation across diverse regions. Our method achieves state-of-the-art performance on the PI-CAI dataset, demonstrating 69.73% IoU and 74.32% Dice scores, and significantly improving prostate cancer lesion detection.
An Artificial Intelligence Model for Early Stage Breast Cancer Detection from Biopsy Images
May 24, 2025Accurate identification of breast cancer types plays a critical role in guiding treatment decisions and improving patient outcomes. This paper presents an artificial intelligence enabled tool designed to aid in the identification of breast cancer types using histopathological biopsy images. Traditionally additional tests have to be done on women who are detected with breast cancer to find out the types of cancer it is to give the necessary cure. Those tests are not only invasive but also delay the initiation of treatment and increase patient burden. The proposed model utilizes a convolutional neural network (CNN) architecture to distinguish between benign and malignant tissues as well as accurate subclassification of breast cancer types. By preprocessing the images to reduce noise and enhance features, the model achieves reliable levels of classification performance. Experimental results on such datasets demonstrate the model's effectiveness, outperforming several existing solutions in terms of accuracy, precision, recall, and F1-score. The study emphasizes the potential of deep learning techniques in clinical diagnostics and offers a promising tool to assist pathologists in breast cancer classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge