Zoubin Ghahramani
Dima
An Infinite Latent Attribute Model for Network Data
Jun 27, 2012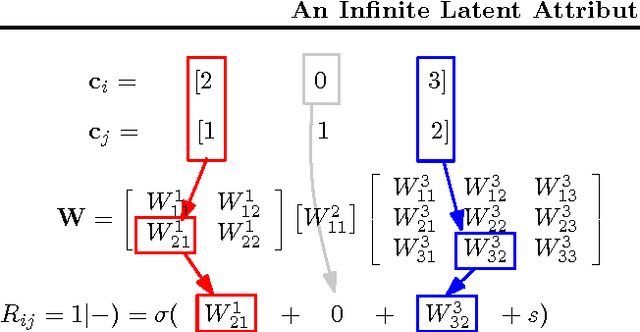
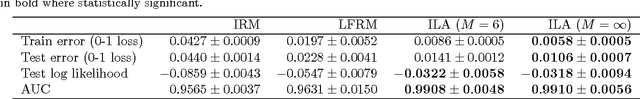

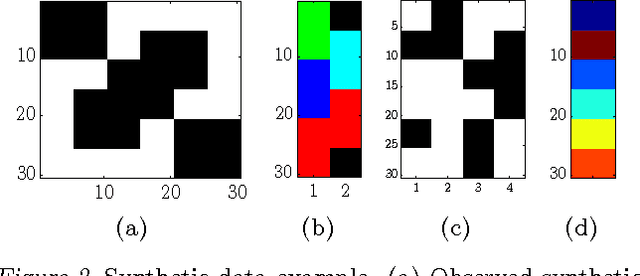

Abstract:Latent variable models for network data extract a summary of the relational structure underlying an observed network. The simplest possible models subdivide nodes of the network into clusters; the probability of a link between any two nodes then depends only on their cluster assignment. Currently available models can be classified by whether clusters are disjoint or are allowed to overlap. These models can explain a "flat" clustering structure. Hierarchical Bayesian models provide a natural approach to capture more complex dependencies. We propose a model in which objects are characterised by a latent feature vector. Each feature is itself partitioned into disjoint groups (subclusters), corresponding to a second layer of hierarchy. In experimental comparisons, the model achieves significantly improved predictive performance on social and biological link prediction tasks. The results indicate that models with a single layer hierarchy over-simplify real networks.
Bayesian Inference for Gaussian Mixed Graph Models
Jun 27, 2012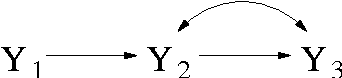
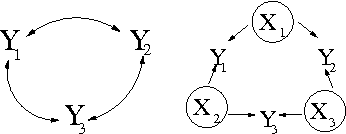
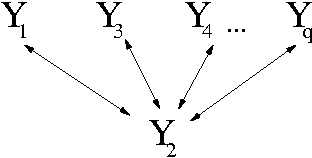
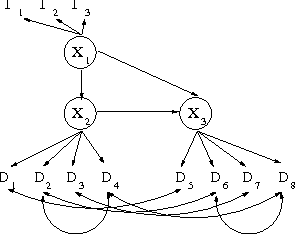
Abstract:We introduce priors and algorithms to perform Bayesian inference in Gaussian models defined by acyclic directed mixed graphs. Such a class of graphs, composed of directed and bi-directed edges, is a representation of conditional independencies that is closed under marginalization and arises naturally from causal models which allow for unmeasured confounding. Monte Carlo methods and a variational approximation for such models are presented. Our algorithms for Bayesian inference allow the evaluation of posterior distributions for several quantities of interest, including causal effects that are not identifiable from data alone but could otherwise be inferred where informative prior knowledge about confounding is available.
Variable noise and dimensionality reduction for sparse Gaussian processes
Jun 27, 2012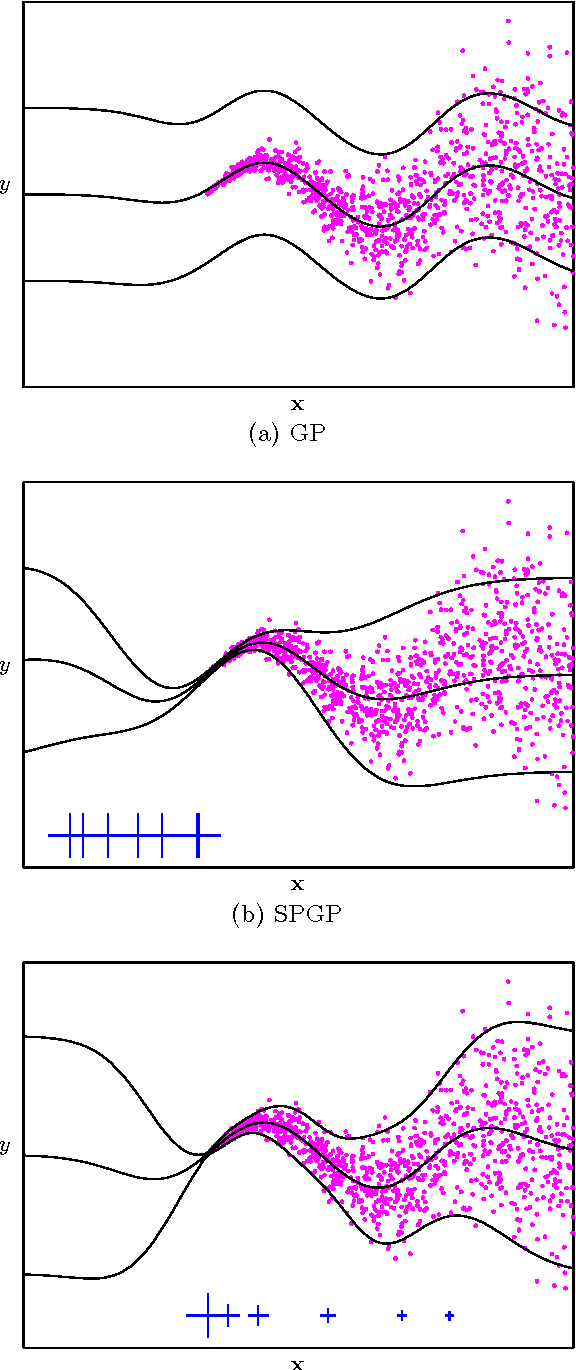
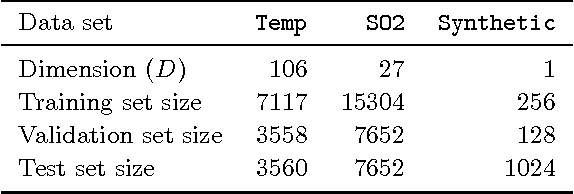
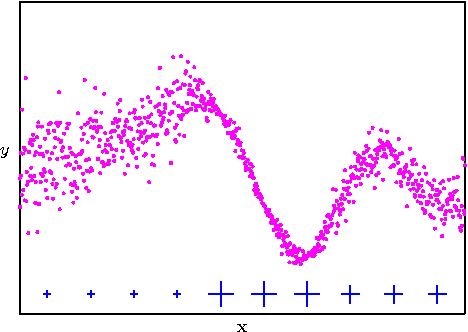
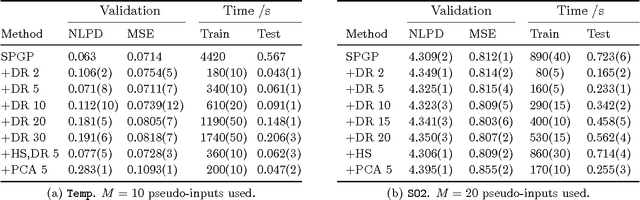
Abstract:The sparse pseudo-input Gaussian process (SPGP) is a new approximation method for speeding up GP regression in the case of a large number of data points N. The approximation is controlled by the gradient optimization of a small set of M `pseudo-inputs', thereby reducing complexity from N^3 to NM^2. One limitation of the SPGP is that this optimization space becomes impractically big for high dimensional data sets. This paper addresses this limitation by performing automatic dimensionality reduction. A projection of the input space to a low dimensional space is learned in a supervised manner, alongside the pseudo-inputs, which now live in this reduced space. The paper also investigates the suitability of the SPGP for modeling data with input-dependent noise. A further extension of the model is made to make it even more powerful in this regard - we learn an uncertainty parameter for each pseudo-input. The combination of sparsity, reduced dimension, and input-dependent noise makes it possible to apply GPs to much larger and more complex data sets than was previously practical. We demonstrate the benefits of these methods on several synthetic and real world problems.
A Non-Parametric Bayesian Method for Inferring Hidden Causes
Jun 27, 2012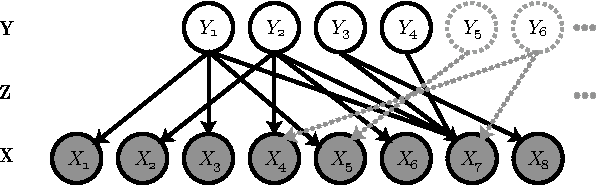
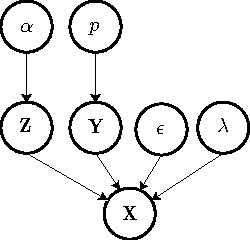
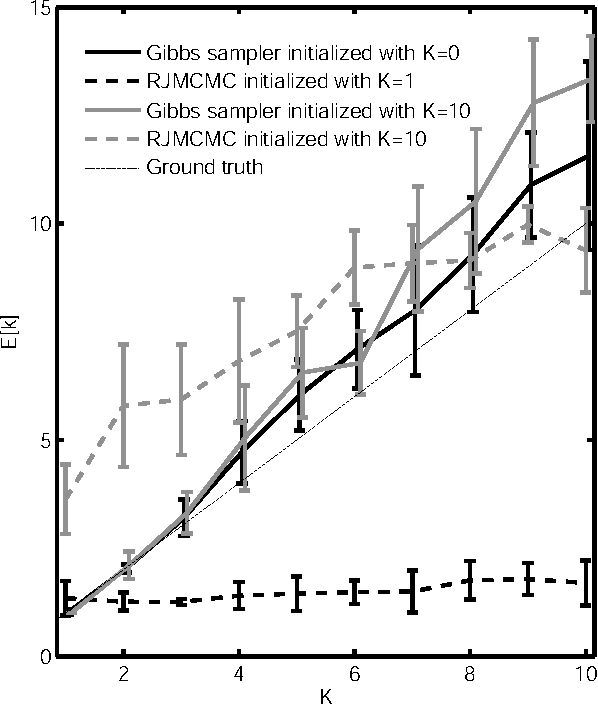
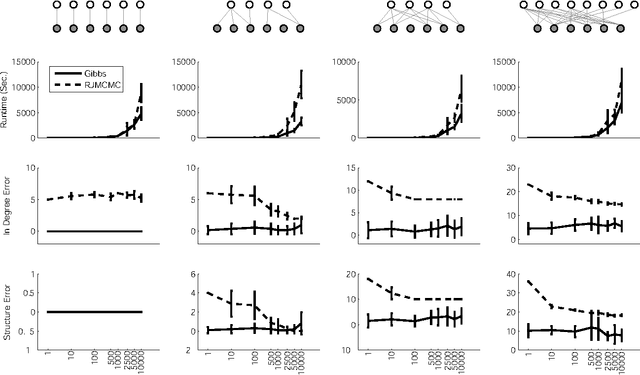
Abstract:We present a non-parametric Bayesian approach to structure learning with hidden causes. Previous Bayesian treatments of this problem define a prior over the number of hidden causes and use algorithms such as reversible jump Markov chain Monte Carlo to move between solutions. In contrast, we assume that the number of hidden causes is unbounded, but only a finite number influence observable variables. This makes it possible to use a Gibbs sampler to approximate the distribution over causal structures. We evaluate the performance of both approaches in discovering hidden causes in simulated data, and use our non-parametric approach to discover hidden causes in a real medical dataset.
Copula-based Kernel Dependency Measures
Jun 18, 2012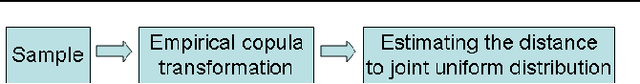
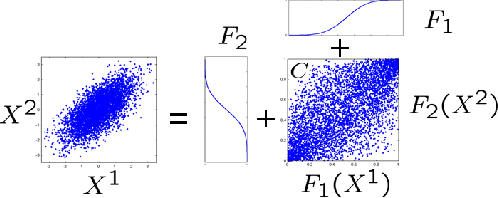
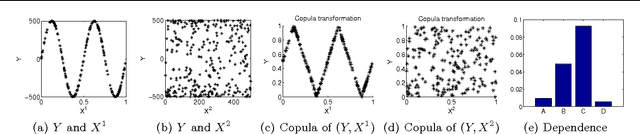
Abstract:The paper presents a new copula based method for measuring dependence between random variables. Our approach extends the Maximum Mean Discrepancy to the copula of the joint distribution. We prove that this approach has several advantageous properties. Similarly to Shannon mutual information, the proposed dependence measure is invariant to any strictly increasing transformation of the marginal variables. This is important in many applications, for example in feature selection. The estimator is consistent, robust to outliers, and uses rank statistics only. We derive upper bounds on the convergence rate and propose independence tests too. We illustrate the theoretical contributions through a series of experiments in feature selection and low-dimensional embedding of distributions.
Correlated Non-Parametric Latent Feature Models
May 09, 2012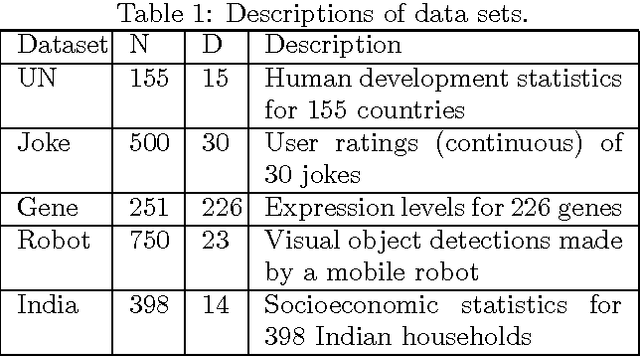
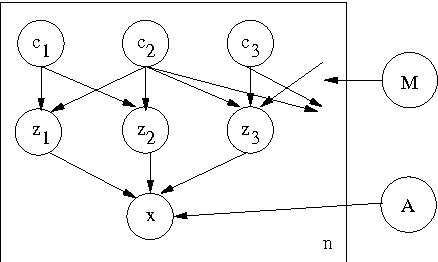
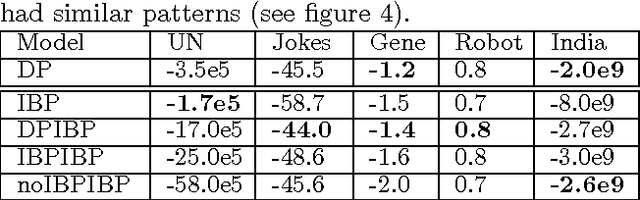
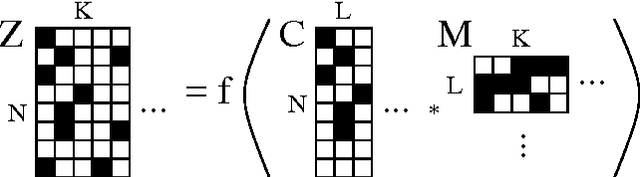
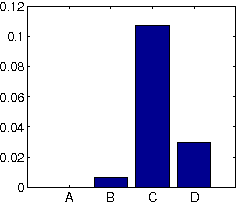
Abstract:We are often interested in explaining data through a set of hidden factors or features. When the number of hidden features is unknown, the Indian Buffet Process (IBP) is a nonparametric latent feature model that does not bound the number of active features in dataset. However, the IBP assumes that all latent features are uncorrelated, making it inadequate for many realworld problems. We introduce a framework for correlated nonparametric feature models, generalising the IBP. We use this framework to generate several specific models and demonstrate applications on realworld datasets.
Bayesian Active Learning for Classification and Preference Learning
Dec 24, 2011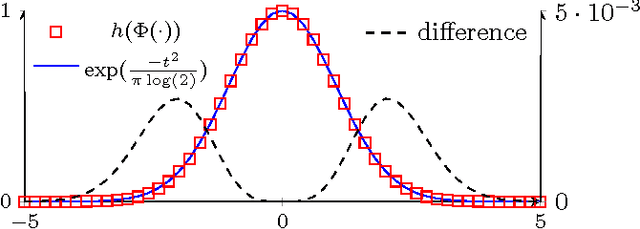
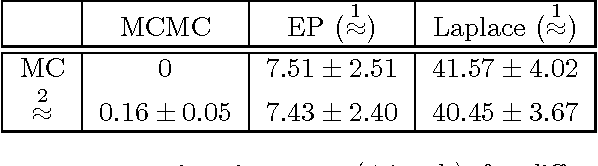
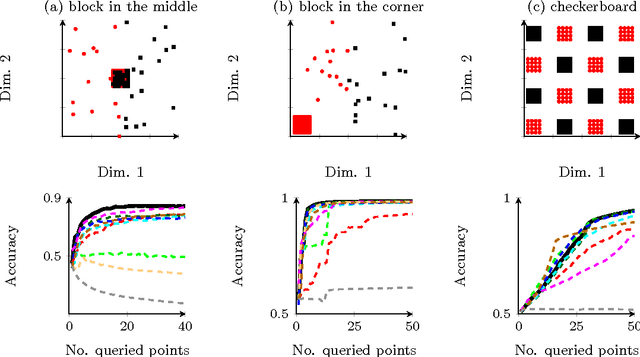
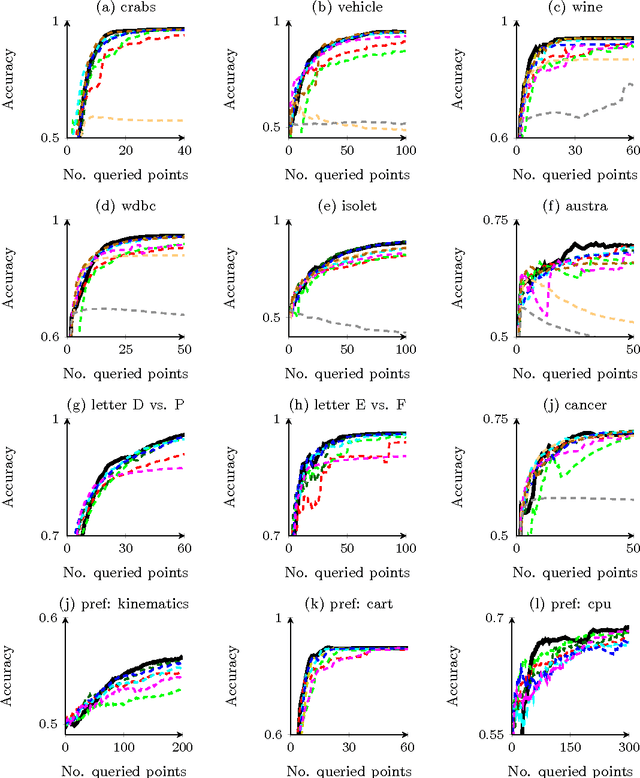
Abstract:Information theoretic active learning has been widely studied for probabilistic models. For simple regression an optimal myopic policy is easily tractable. However, for other tasks and with more complex models, such as classification with nonparametric models, the optimal solution is harder to compute. Current approaches make approximations to achieve tractability. We propose an approach that expresses information gain in terms of predictive entropies, and apply this method to the Gaussian Process Classifier (GPC). Our approach makes minimal approximations to the full information theoretic objective. Our experimental performance compares favourably to many popular active learning algorithms, and has equal or lower computational complexity. We compare well to decision theoretic approaches also, which are privy to more information and require much more computational time. Secondly, by developing further a reformulation of binary preference learning to a classification problem, we extend our algorithm to Gaussian Process preference learning.
Gaussian Process Regression Networks
Oct 19, 2011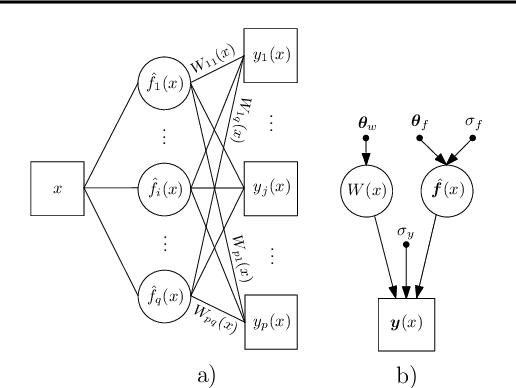
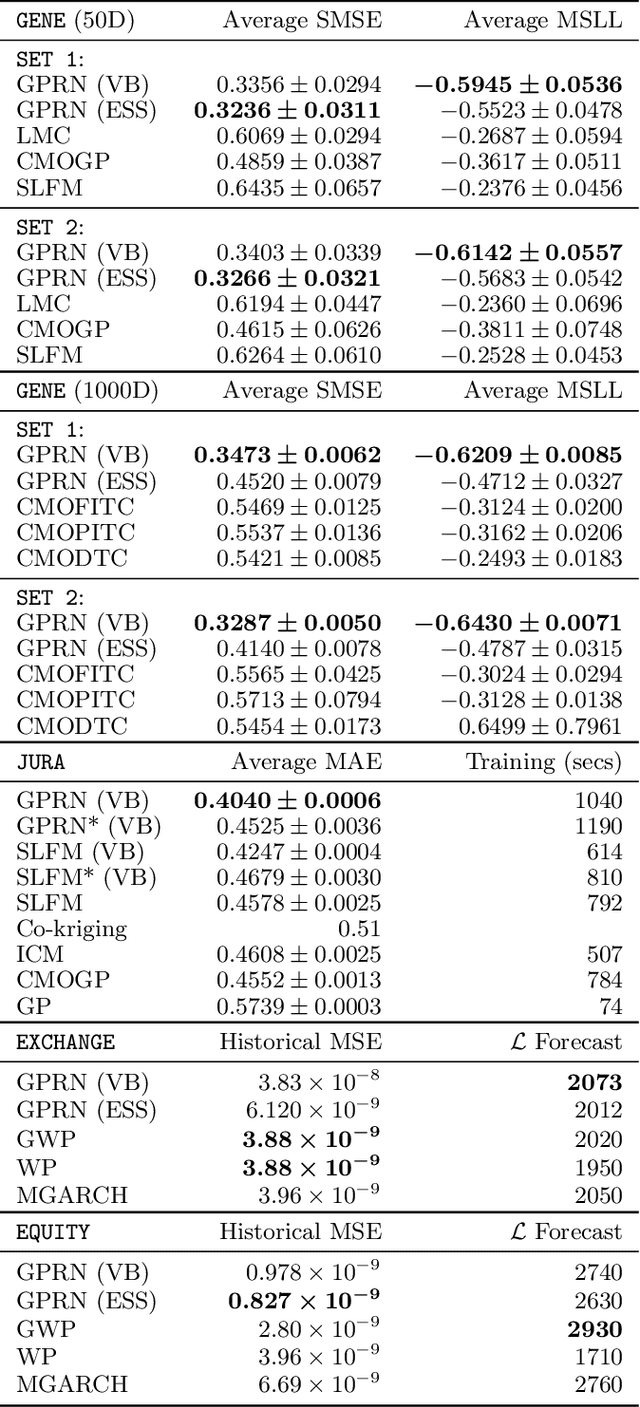
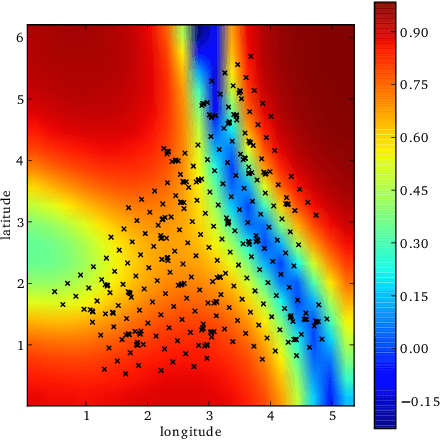
Abstract:We introduce a new regression framework, Gaussian process regression networks (GPRN), which combines the structural properties of Bayesian neural networks with the non-parametric flexibility of Gaussian processes. This model accommodates input dependent signal and noise correlations between multiple response variables, input dependent length-scales and amplitudes, and heavy-tailed predictive distributions. We derive both efficient Markov chain Monte Carlo and variational Bayes inference procedures for this model. We apply GPRN as a multiple output regression and multivariate volatility model, demonstrating substantially improved performance over eight popular multiple output (multi-task) Gaussian process models and three multivariate volatility models on benchmark datasets, including a 1000 dimensional gene expression dataset.
Nonparametric Bayesian sparse factor models with application to gene expression modeling
Jul 28, 2011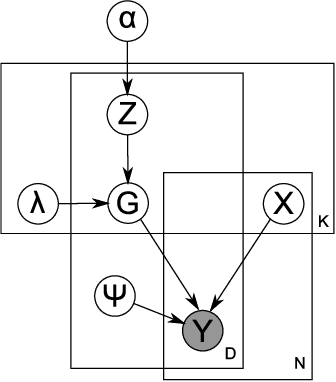
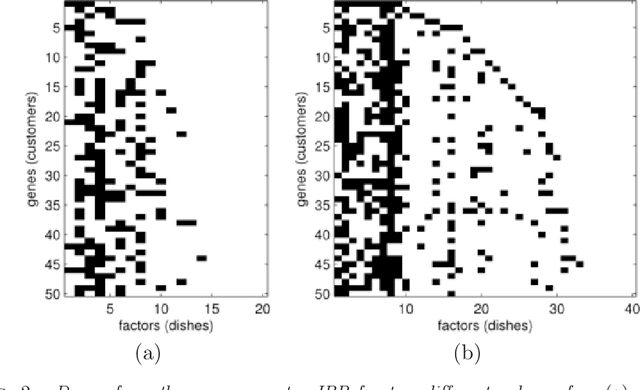
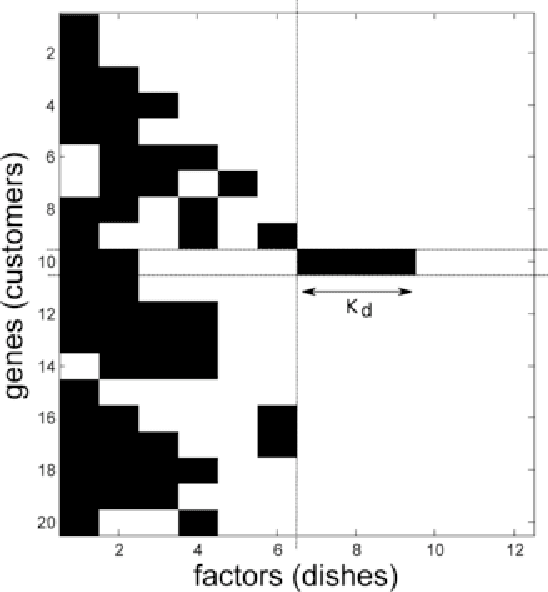
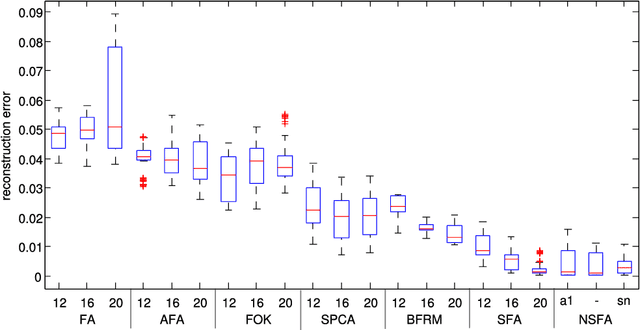
Abstract:A nonparametric Bayesian extension of Factor Analysis (FA) is proposed where observed data $\mathbf{Y}$ is modeled as a linear superposition, $\mathbf{G}$, of a potentially infinite number of hidden factors, $\mathbf{X}$. The Indian Buffet Process (IBP) is used as a prior on $\mathbf{G}$ to incorporate sparsity and to allow the number of latent features to be inferred. The model's utility for modeling gene expression data is investigated using randomly generated data sets based on a known sparse connectivity matrix for E. Coli, and on three biological data sets of increasing complexity.
* Published in at http://dx.doi.org/10.1214/10-AOAS435 the Annals of Applied Statistics (http://www.imstat.org/aoas/) by the Institute of Mathematical Statistics (http://www.imstat.org)
Pitman-Yor Diffusion Trees
Jun 16, 2011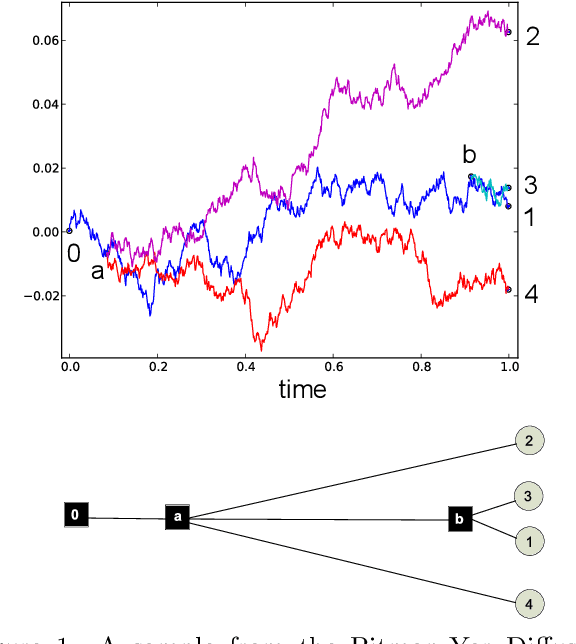
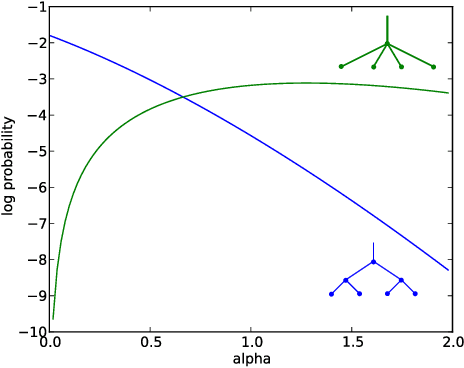
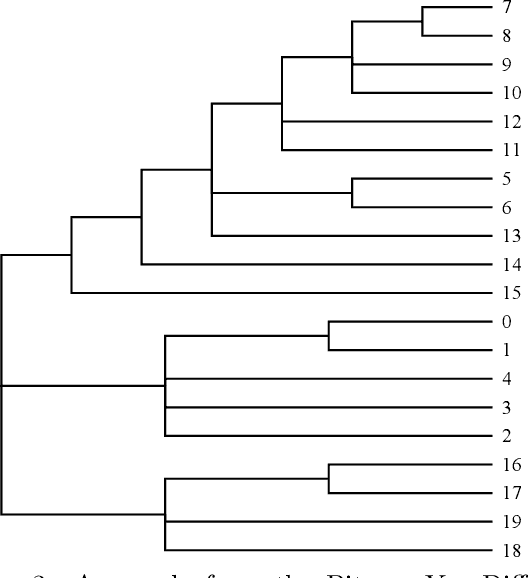
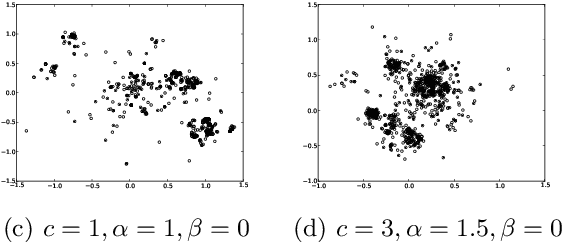
Abstract:We introduce the Pitman Yor Diffusion Tree (PYDT) for hierarchical clustering, a generalization of the Dirichlet Diffusion Tree (Neal, 2001) which removes the restriction to binary branching structure. The generative process is described and shown to result in an exchangeable distribution over data points. We prove some theoretical properties of the model and then present two inference methods: a collapsed MCMC sampler which allows us to model uncertainty over tree structures, and a computationally efficient greedy Bayesian EM search algorithm. Both algorithms use message passing on the tree structure. The utility of the model and algorithms is demonstrated on synthetic and real world data, both continuous and binary.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge