Youbao Tang
TUNA-Net: Task-oriented UNsupervised Adversarial Network for Disease Recognition in Cross-Domain Chest X-rays
Aug 21, 2019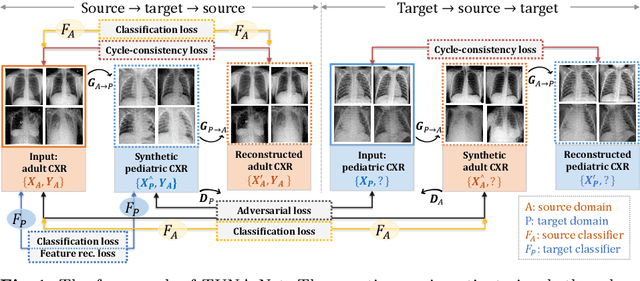
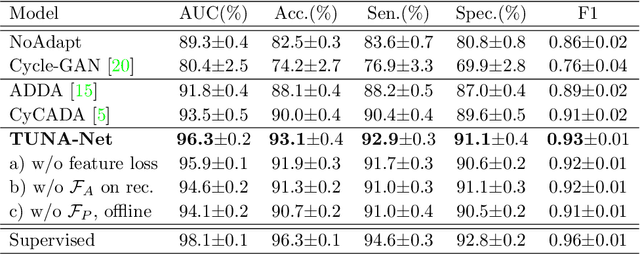
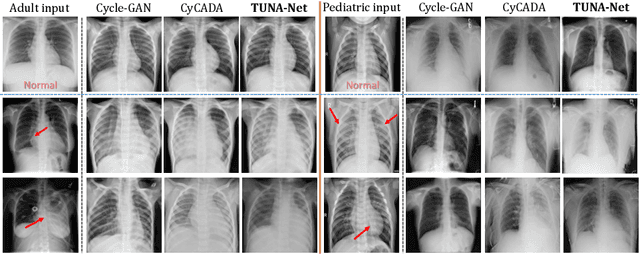
Abstract:In this work, we exploit the unsupervised domain adaptation problem for radiology image interpretation across domains. Specifically, we study how to adapt the disease recognition model from a labeled source domain to an unlabeled target domain, so as to reduce the effort of labeling each new dataset. To address the shortcoming of cross-domain, unpaired image-to-image translation methods which typically ignore class-specific semantics, we propose a task-driven, discriminatively trained, cycle-consistent generative adversarial network, termed TUNA-Net. It is able to preserve 1) low-level details, 2) high-level semantic information and 3) mid-level feature representation during the image-to-image translation process, to favor the target disease recognition task. The TUNA-Net framework is general and can be readily adapted to other learning tasks. We evaluate the proposed framework on two public chest X-ray datasets for pneumonia recognition. The TUNA-Net model can adapt labeled adult chest X-rays in the source domain such that they appear as if they were drawn from pediatric X-rays in the unlabeled target domain, while preserving the disease semantics. Extensive experiments show the superiority of the proposed method as compared to state-of-the-art unsupervised domain adaptation approaches. Notably, TUNA-Net achieves an AUC of 96.3% for pediatric pneumonia classification, which is very close to that of the supervised approach (98.1%), but without the need for labels on the target domain.
MULAN: Multitask Universal Lesion Analysis Network for Joint Lesion Detection, Tagging, and Segmentation
Aug 12, 2019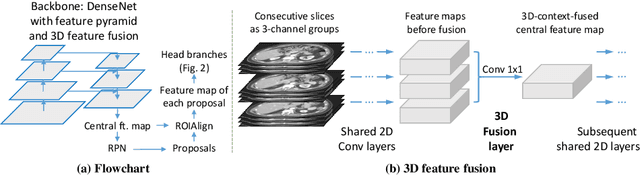
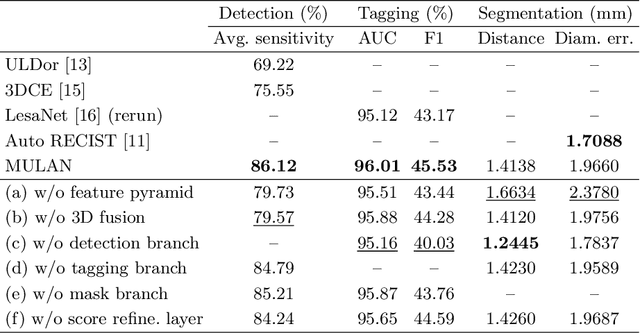
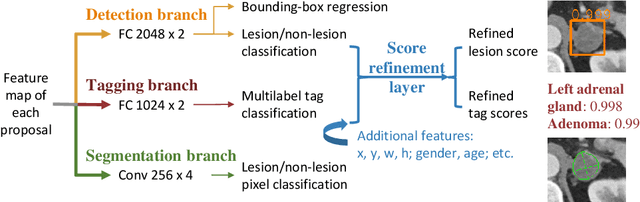
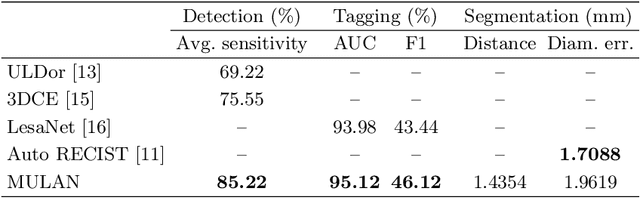
Abstract:When reading medical images such as a computed tomography (CT) scan, radiologists generally search across the image to find lesions, characterize and measure them, and then describe them in the radiological report. To automate this process, we propose a multitask universal lesion analysis network (MULAN) for joint detection, tagging, and segmentation of lesions in a variety of body parts, which greatly extends existing work of single-task lesion analysis on specific body parts. MULAN is based on an improved Mask R-CNN framework with three head branches and a 3D feature fusion strategy. It achieves the state-of-the-art accuracy in the detection and tagging tasks on the DeepLesion dataset, which contains 32K lesions in the whole body. We also analyze the relationship between the three tasks and show that tag predictions can improve detection accuracy via a score refinement layer.
XLSor: A Robust and Accurate Lung Segmentor on Chest X-Rays Using Criss-Cross Attention and Customized Radiorealistic Abnormalities Generation
Apr 19, 2019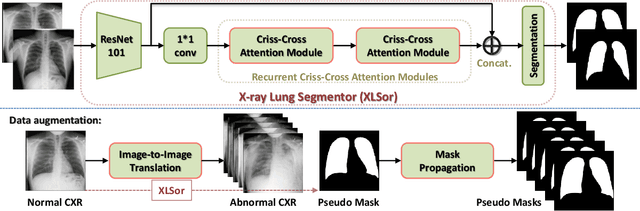
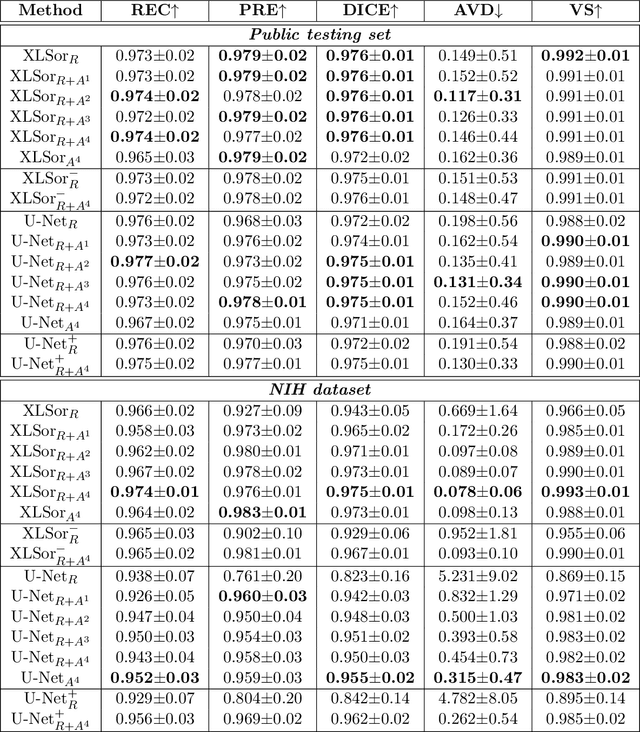
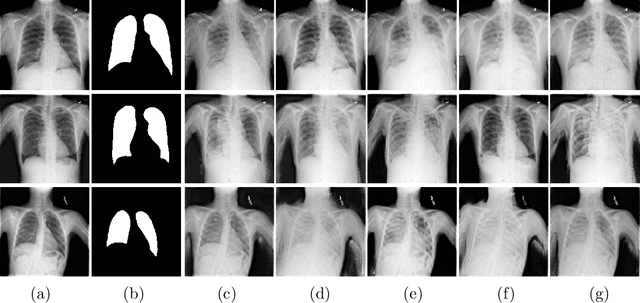
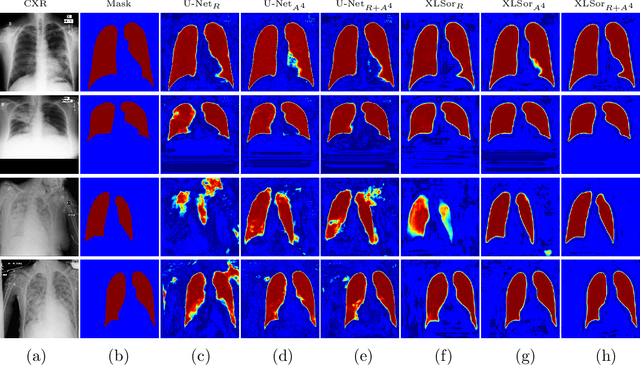
Abstract:This paper proposes a novel framework for lung segmentation in chest X-rays. It consists of two key contributions, a criss-cross attention based segmentation network and radiorealistic chest X-ray image synthesis (i.e. a synthesized radiograph that appears anatomically realistic) for data augmentation. The criss-cross attention modules capture rich global contextual information in both horizontal and vertical directions for all the pixels thus facilitating accurate lung segmentation. To reduce the manual annotation burden and to train a robust lung segmentor that can be adapted to pathological lungs with hazy lung boundaries, an image-to-image translation module is employed to synthesize radiorealistic abnormal CXRs from the source of normal ones for data augmentation. The lung masks of synthetic abnormal CXRs are propagated from the segmentation results of their normal counterparts, and then serve as pseudo masks for robust segmentor training. In addition, we annotate 100 CXRs with lung masks on a more challenging NIH Chest X-ray dataset containing both posterioranterior and anteroposterior views for evaluation. Extensive experiments validate the robustness and effectiveness of the proposed framework. The code and data can be found from https://github.com/rsummers11/CADLab/tree/master/Lung_Segmentation_XLSor .
Abnormal Chest X-ray Identification With Generative Adversarial One-Class Classifier
Mar 05, 2019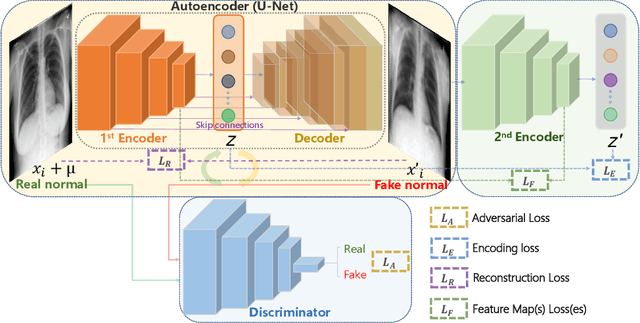
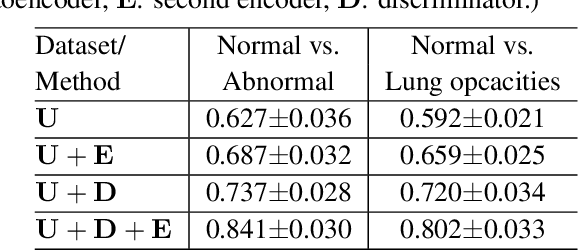
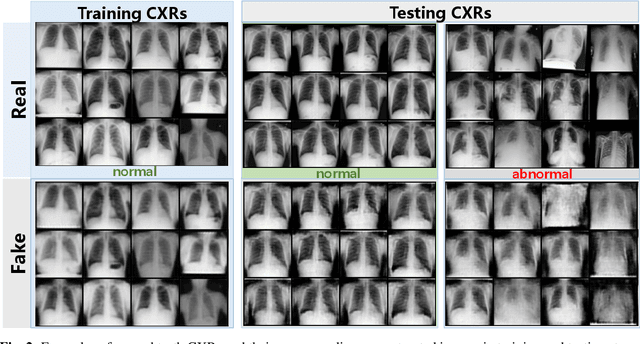
Abstract:Being one of the most common diagnostic imaging tests, chest radiography requires timely reporting of potential findings in the images. In this paper, we propose an end-to-end architecture for abnormal chest X-ray identification using generative adversarial one-class learning. Unlike previous approaches, our method takes only normal chest X-ray images as input. The architecture is composed of three deep neural networks, each of which learned by competing while collaborating among them to model the underlying content structure of the normal chest X-rays. Given a chest X-ray image in the testing phase, if it is normal, the learned architecture can well model and reconstruct the content; if it is abnormal, since the content is unseen in the training phase, the model would perform poorly in its reconstruction. It thus enables distinguishing abnormal chest X-rays from normal ones. Quantitative and qualitative experiments demonstrate the effectiveness and efficiency of our approach, where an AUC of 0.841 is achieved on the challenging NIH Chest X-ray dataset in a one-class learning setting, with the potential in reducing the workload for radiologists.
ULDor: A Universal Lesion Detector for CT Scans with Pseudo Masks and Hard Negative Example Mining
Jan 18, 2019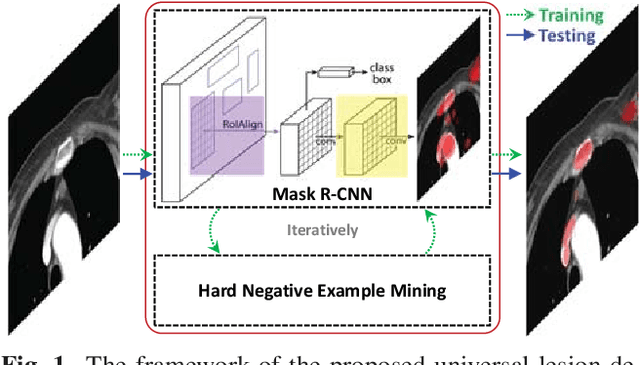
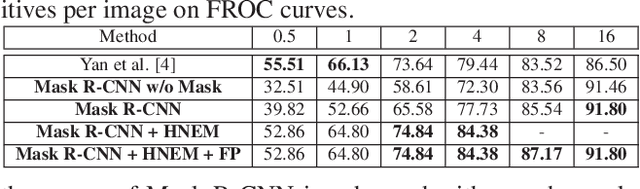
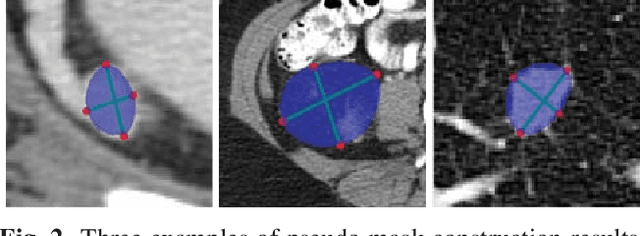
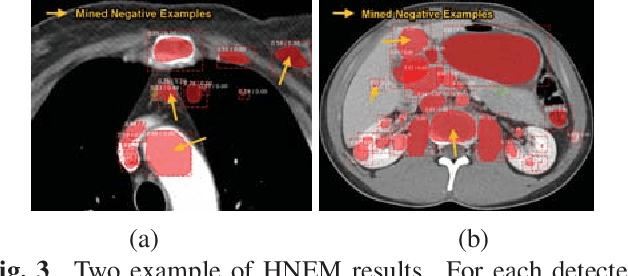
Abstract:Automatic lesion detection from computed tomography (CT) scans is an important task in medical imaging analysis. It is still very challenging due to similar appearances (e.g. intensity and texture) between lesions and other tissues, making it especially difficult to develop a universal lesion detector. Instead of developing a specific-type lesion detector, this work builds a Universal Lesion Detector (ULDor) based on Mask R-CNN, which is able to detect all different kinds of lesions from whole body parts. As a state-of-the-art object detector, Mask R-CNN adds a branch for predicting segmentation masks on each Region of Interest (RoI) to improve the detection performance. However, it is almost impossible to manually annotate a large-scale dataset with pixel-level lesion masks to train the Mask R-CNN for lesion detection. To address this problem, this work constructs a pseudo mask for each lesion region that can be considered as a surrogate of the real mask, based on which the Mask R-CNN is employed for lesion detection. On the other hand, this work proposes a hard negative example mining strategy to reduce the false positives for improving the detection performance. Experimental results on the NIH DeepLesion dataset demonstrate that the ULDor is enhanced using pseudo masks and the proposed hard negative example mining strategy and achieves a sensitivity of 86.21% with five false positives per image.
CT Image Enhancement Using Stacked Generative Adversarial Networks and Transfer Learning for Lesion Segmentation Improvement
Jul 18, 2018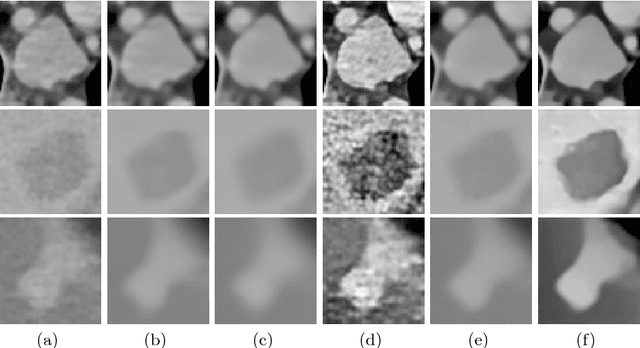
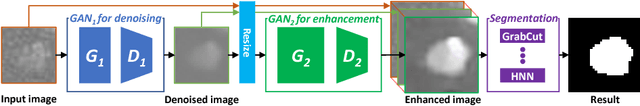
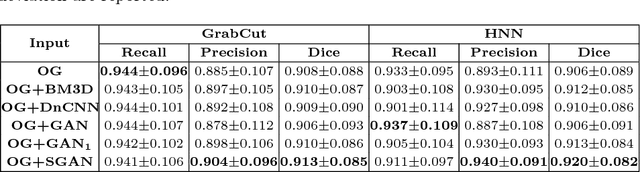

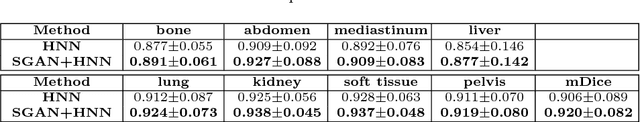
Abstract:Automated lesion segmentation from computed tomography (CT) is an important and challenging task in medical image analysis. While many advancements have been made, there is room for continued improvements. One hurdle is that CT images can exhibit high noise and low contrast, particularly in lower dosages. To address this, we focus on a preprocessing method for CT images that uses stacked generative adversarial networks (SGAN) approach. The first GAN reduces the noise in the CT image and the second GAN generates a higher resolution image with enhanced boundaries and high contrast. To make up for the absence of high quality CT images, we detail how to synthesize a large number of low- and high-quality natural images and use transfer learning with progressively larger amounts of CT images. We apply both the classic GrabCut method and the modern holistically nested network (HNN) to lesion segmentation, testing whether SGAN can yield improved lesion segmentation. Experimental results on the DeepLesion dataset demonstrate that the SGAN enhancements alone can push GrabCut performance over HNN trained on original images. We also demonstrate that HNN + SGAN performs best compared against four other enhancement methods, including when using only a single GAN.
Accurate Weakly-Supervised Deep Lesion Segmentation using Large-Scale Clinical Annotations: Slice-Propagated 3D Mask Generation from 2D RECIST
Jul 02, 2018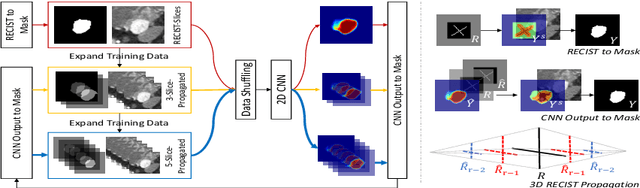
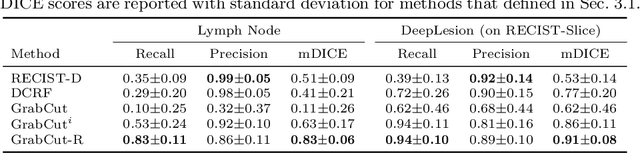
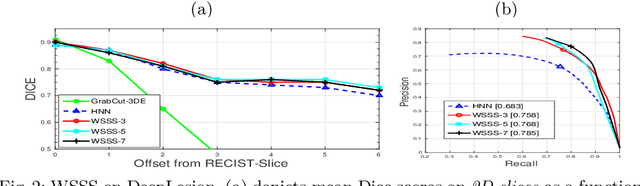
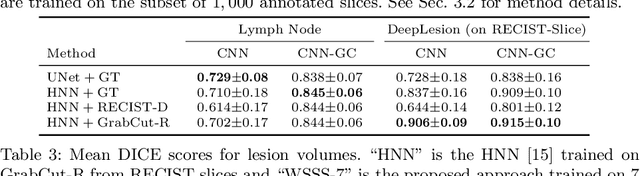
Abstract:Volumetric lesion segmentation from computed tomography (CT) images is a powerful means to precisely assess multiple time-point lesion/tumor changes. However, because manual 3D segmentation is prohibitively time consuming, current practices rely on an imprecise surrogate called response evaluation criteria in solid tumors (RECIST). Despite their coarseness, RECIST markers are commonly found in current hospital picture and archiving systems (PACS), meaning they can provide a potentially powerful, yet extraordinarily challenging, source of weak supervision for full 3D segmentation. Toward this end, we introduce a convolutional neural network (CNN) based weakly supervised slice-propagated segmentation (WSSS) method to 1) generate the initial lesion segmentation on the axial RECIST-slice; 2) learn the data distribution on RECIST-slices; 3) extrapolate to segment the whole lesion slice by slice to finally obtain a volumetric segmentation. To validate the proposed method, we first test its performance on a fully annotated lymph node dataset, where WSSS performs comparably to its fully supervised counterparts. We then test on a comprehensive lesion dataset with 32,735 RECIST marks, where we report a mean Dice score of 92% on RECIST-marked slices and 76% on the entire 3D volumes.
Semi-Automatic RECIST Labeling on CT Scans with Cascaded Convolutional Neural Networks
Jun 25, 2018
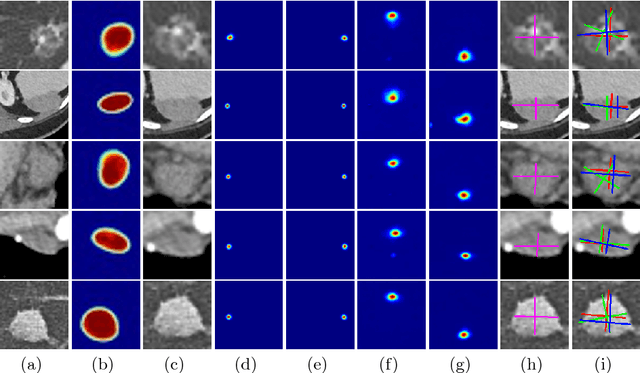
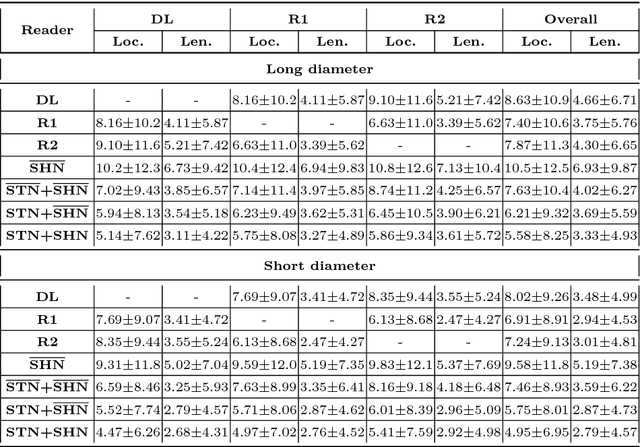
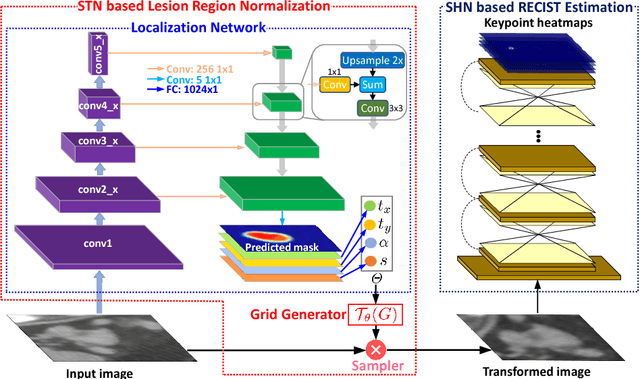

Abstract:Response evaluation criteria in solid tumors (RECIST) is the standard measurement for tumor extent to evaluate treatment responses in cancer patients. As such, RECIST annotations must be accurate. However, RECIST annotations manually labeled by radiologists require professional knowledge and are time-consuming, subjective, and prone to inconsistency among different observers. To alleviate these problems, we propose a cascaded convolutional neural network based method to semi-automatically label RECIST annotations and drastically reduce annotation time. The proposed method consists of two stages: lesion region normalization and RECIST estimation. We employ the spatial transformer network (STN) for lesion region normalization, where a localization network is designed to predict the lesion region and the transformation parameters with a multi-task learning strategy. For RECIST estimation, we adapt the stacked hourglass network (SHN), introducing a relationship constraint loss to improve the estimation precision. STN and SHN can both be learned in an end-to-end fashion. We train our system on the DeepLesion dataset, obtaining a consensus model trained on RECIST annotations performed by multiple radiologists over a multi-year period. Importantly, when judged against the inter-reader variability of two additional radiologist raters, our system performs more stably and with less variability, suggesting that RECIST annotations can be reliably obtained with reduced labor and time.
CT-Realistic Lung Nodule Simulation from 3D Conditional Generative Adversarial Networks for Robust Lung Segmentation
Jun 11, 2018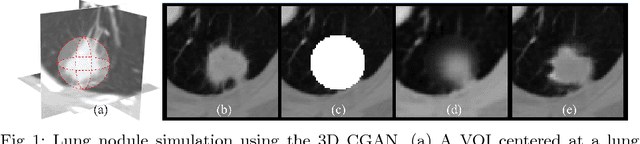
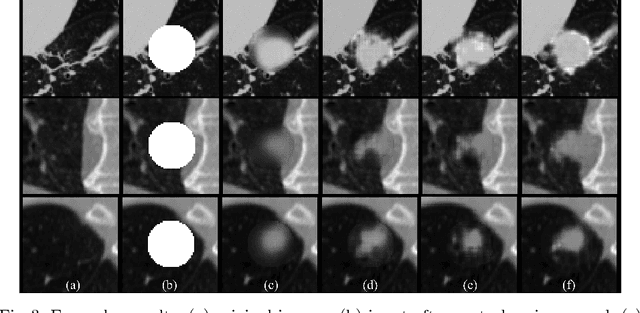
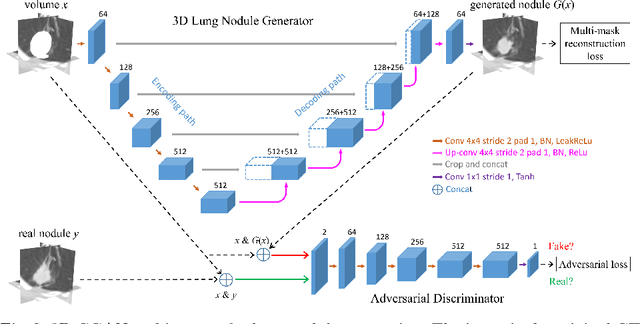

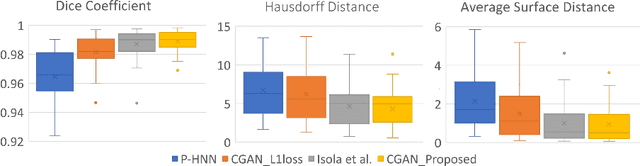
Abstract:Data availability plays a critical role for the performance of deep learning systems. This challenge is especially acute within the medical image domain, particularly when pathologies are involved, due to two factors: 1) limited number of cases, and 2) large variations in location, scale, and appearance. In this work, we investigate whether augmenting a dataset with artificially generated lung nodules can improve the robustness of the progressive holistically nested network (P-HNN) model for pathological lung segmentation of CT scans. To achieve this goal, we develop a 3D generative adversarial network (GAN) that effectively learns lung nodule property distributions in 3D space. In order to embed the nodules within their background context, we condition the GAN based on a volume of interest whose central part containing the nodule has been erased. To further improve realism and blending with the background, we propose a novel multi-mask reconstruction loss. We train our method on over 1000 nodules from the LIDC dataset. Qualitative results demonstrate the effectiveness of our method compared to the state-of-art. We then use our GAN to generate simulated training images where nodules lie on the lung border, which are cases where the published P-HNN model struggles. Qualitative and quantitative results demonstrate that armed with these simulated images, the P-HNN model learns to better segment lung regions under these challenging situations. As a result, our system provides a promising means to help overcome the data paucity that commonly afflicts medical imaging.
Accurate Weakly Supervised Deep Lesion Segmentation on CT Scans: Self-Paced 3D Mask Generation from RECIST
Jan 25, 2018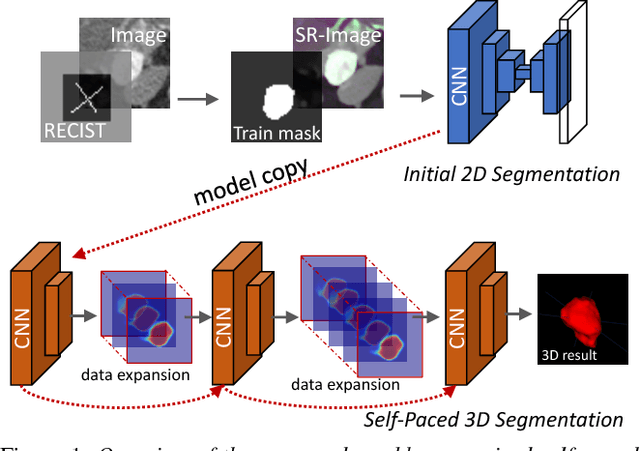
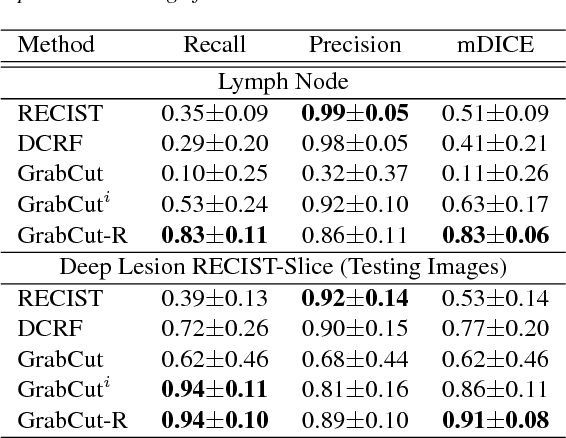
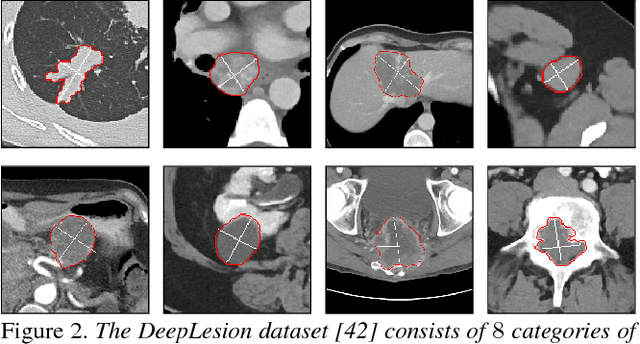
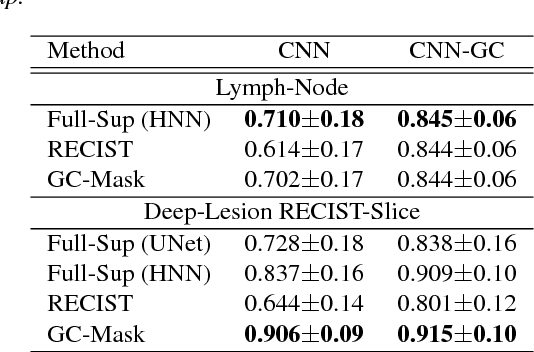
Abstract:Volumetric lesion segmentation via medical imaging is a powerful means to precisely assess multiple time-point lesion/tumor changes. Because manual 3D segmentation is prohibitively time consuming and requires radiological experience, current practices rely on an imprecise surrogate called response evaluation criteria in solid tumors (RECIST). Despite their coarseness, RECIST marks are commonly found in current hospital picture and archiving systems (PACS), meaning they can provide a potentially powerful, yet extraordinarily challenging, source of weak supervision for full 3D segmentation. Toward this end, we introduce a convolutional neural network based weakly supervised self-paced segmentation (WSSS) method to 1) generate the initial lesion segmentation on the axial RECIST-slice; 2) learn the data distribution on RECIST-slices; 3) adapt to segment the whole volume slice by slice to finally obtain a volumetric segmentation. In addition, we explore how super-resolution images (2~5 times beyond the physical CT imaging), generated from a proposed stacked generative adversarial network, can aid the WSSS performance. We employ the DeepLesion dataset, a comprehensive CT-image lesion dataset of 32,735 PACS-bookmarked findings, which include lesions, tumors, and lymph nodes of varying sizes, categories, body regions and surrounding contexts. These are drawn from 10,594 studies of 4,459 patients. We also validate on a lymph-node dataset, where 3D ground truth masks are available for all images. For the DeepLesion dataset, we report mean Dice coefficients of 93% on RECIST-slices and 76% in 3D lesion volumes. We further validate using a subjective user study, where an experienced radiologist accepted our WSSS-generated lesion segmentation results with a high probability of 92.4%.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge