Yikun Yang
Step 3.5 Flash: Open Frontier-Level Intelligence with 11B Active Parameters
Feb 11, 2026Abstract:We introduce Step 3.5 Flash, a sparse Mixture-of-Experts (MoE) model that bridges frontier-level agentic intelligence and computational efficiency. We focus on what matters most when building agents: sharp reasoning and fast, reliable execution. Step 3.5 Flash pairs a 196B-parameter foundation with 11B active parameters for efficient inference. It is optimized with interleaved 3:1 sliding-window/full attention and Multi-Token Prediction (MTP-3) to reduce the latency and cost of multi-round agentic interactions. To reach frontier-level intelligence, we design a scalable reinforcement learning framework that combines verifiable signals with preference feedback, while remaining stable under large-scale off-policy training, enabling consistent self-improvement across mathematics, code, and tool use. Step 3.5 Flash demonstrates strong performance across agent, coding, and math tasks, achieving 85.4% on IMO-AnswerBench, 86.4% on LiveCodeBench-v6 (2024.08-2025.05), 88.2% on tau2-Bench, 69.0% on BrowseComp (with context management), and 51.0% on Terminal-Bench 2.0, comparable to frontier models such as GPT-5.2 xHigh and Gemini 3.0 Pro. By redefining the efficiency frontier, Step 3.5 Flash provides a high-density foundation for deploying sophisticated agents in real-world industrial environments.
Unified Unbiased Variance Estimation for MMD: Robust Finite-Sample Performance with Imbalanced Data and Exact Acceleration under Null and Alternative Hypotheses
Jan 20, 2026Abstract:The maximum mean discrepancy (MMD) is a kernel-based nonparametric statistic for two-sample testing, whose inferential accuracy depends critically on variance characterization. Existing work provides various finite-sample estimators of the MMD variance, often differing under the null and alternative hypotheses and across balanced or imbalanced sampling schemes. In this paper, we study the variance of the MMD statistic through its U-statistic representation and Hoeffding decomposition, and establish a unified finite-sample characterization covering different hypotheses and sample configurations. Building on this analysis, we propose an exact acceleration method for the univariate case under the Laplacian kernel, which reduces the overall computational complexity from $\mathcal O(n^2)$ to $\mathcal O(n \log n)$.
ConvPath: A Software Tool for Lung Adenocarcinoma Digital Pathological Image Analysis Aided by Convolutional Neural Network
Sep 20, 2018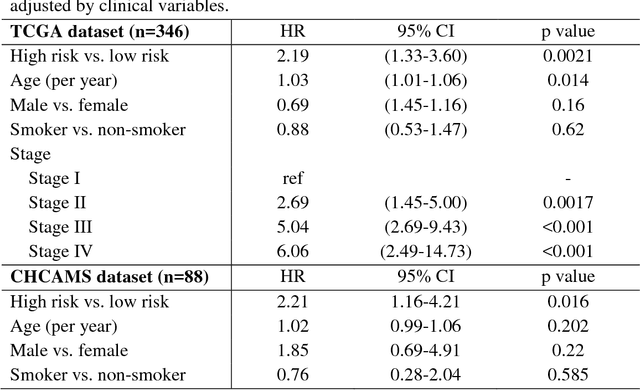
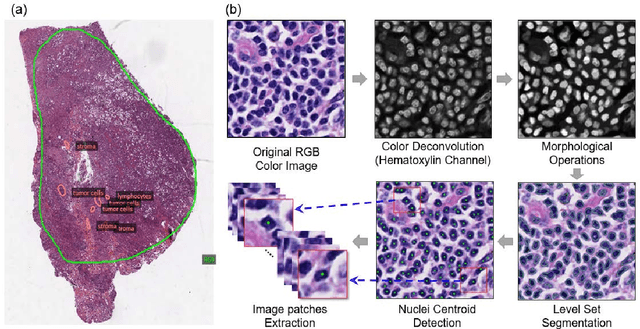
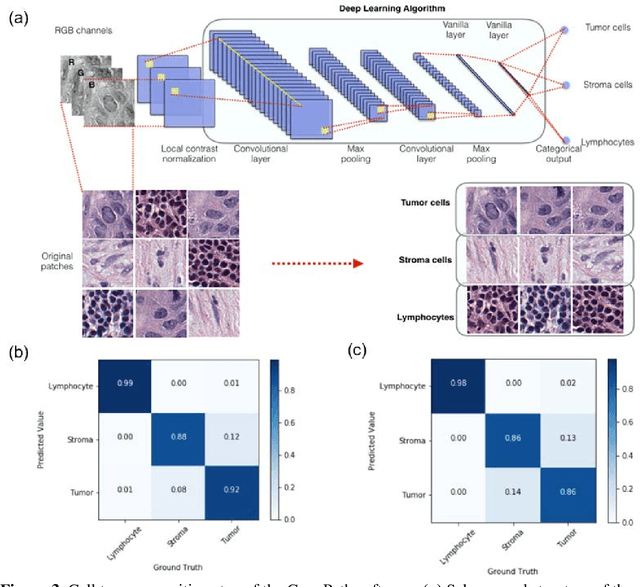
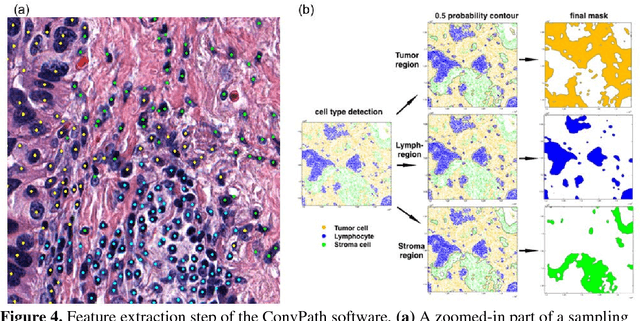
Abstract:The spatial distributions of different types of cells could reveal a cancer cell growth pattern, its relationships with the tumor microenvironment and the immune response of the body, all of which represent key hallmarks of cancer. However, manually recognizing and localizing all the cells in pathology slides are almost impossible. In this study, we developed an automated cell type classification pipeline, ConvPath, which includes nuclei segmentation, convolutional neural network-based tumor, stromal and lymphocytes classification, and extraction of tumor microenvironment related features for lung cancer pathology images. The overall classification accuracy is 92.9% and 90.1% in training and independent testing datasets, respectively. By identifying cells and classifying cell types, this pipeline can convert a pathology image into a spatial map of tumor, stromal and lymphocyte cells. From this spatial map, we can extracted features that characterize the tumor micro-environment. Based on these features, we developed an image feature-based prognostic model and validated the model in two independent cohorts. The predicted risk group serves as an independent prognostic factor, after adjusting for clinical variables that include age, gender, smoking status, and stage.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge