Xiang Dai
Reliable Thinking with Images
Feb 16, 2026Abstract:As a multimodal extension of Chain-of-Thought (CoT), Thinking with Images (TWI) has recently emerged as a promising avenue to enhance the reasoning capability of Multi-modal Large Language Models (MLLMs), which generates interleaved CoT by incorporating visual cues into the textual reasoning process. However, the success of existing TWI methods heavily relies on the assumption that interleaved image-text CoTs are faultless, which is easily violated in real-world scenarios due to the complexity of multimodal understanding. In this paper, we reveal and study a highly-practical yet under-explored problem in TWI, termed Noisy Thinking (NT). Specifically, NT refers to the imperfect visual cues mining and answer reasoning process. As the saying goes, ``One mistake leads to another'', erroneous interleaved CoT would cause error accumulation, thus significantly degrading the performance of MLLMs. To solve the NT problem, we propose a novel method dubbed Reliable Thinking with Images (RTWI). In brief, RTWI estimates the reliability of visual cues and textual CoT in a unified text-centric manner and accordingly employs robust filtering and voting modules to prevent NT from contaminating the final answer. Extensive experiments on seven benchmarks verify the effectiveness of RTWI against NT.
Can AI Extract Antecedent Factors of Human Trust in AI? An Application of Information Extraction for Scientific Literature in Behavioural and Computer Sciences
Dec 16, 2024



Abstract:Information extraction from the scientific literature is one of the main techniques to transform unstructured knowledge hidden in the text into structured data which can then be used for decision-making in down-stream tasks. One such area is Trust in AI, where factors contributing to human trust in artificial intelligence applications are studied. The relationships of these factors with human trust in such applications are complex. We hence explore this space from the lens of information extraction where, with the input of domain experts, we carefully design annotation guidelines, create the first annotated English dataset in this domain, investigate an LLM-guided annotation, and benchmark it with state-of-the-art methods using large language models in named entity and relation extraction. Our results indicate that this problem requires supervised learning which may not be currently feasible with prompt-based LLMs.
A Critical Look at Meta-evaluating Summarisation Evaluation Metrics
Sep 29, 2024Abstract:Effective summarisation evaluation metrics enable researchers and practitioners to compare different summarisation systems efficiently. Estimating the effectiveness of an automatic evaluation metric, termed meta-evaluation, is a critically important research question. In this position paper, we review recent meta-evaluation practices for summarisation evaluation metrics and find that (1) evaluation metrics are primarily meta-evaluated on datasets consisting of examples from news summarisation datasets, and (2) there has been a noticeable shift in research focus towards evaluating the faithfulness of generated summaries. We argue that the time is ripe to build more diverse benchmarks that enable the development of more robust evaluation metrics and analyze the generalization ability of existing evaluation metrics. In addition, we call for research focusing on user-centric quality dimensions that consider the generated summary's communicative goal and the role of summarisation in the workflow.
MultiADE: A Multi-domain Benchmark for Adverse Drug Event Extraction
May 28, 2024



Abstract:Objective. Active adverse event surveillance monitors Adverse Drug Events (ADE) from different data sources, such as electronic health records, medical literature, social media and search engine logs. Over years, many datasets are created, and shared tasks are organised to facilitate active adverse event surveillance. However, most-if not all-datasets or shared tasks focus on extracting ADEs from a particular type of text. Domain generalisation-the ability of a machine learning model to perform well on new, unseen domains (text types)-is under-explored. Given the rapid advancements in natural language processing, one unanswered question is how far we are from having a single ADE extraction model that are effective on various types of text, such as scientific literature and social media posts}. Methods. We contribute to answering this question by building a multi-domain benchmark for adverse drug event extraction, which we named MultiADE. The new benchmark comprises several existing datasets sampled from different text types and our newly created dataset-CADECv2, which is an extension of CADEC (Karimi, et al., 2015), covering online posts regarding more diverse drugs than CADEC. Our new dataset is carefully annotated by human annotators following detailed annotation guidelines. Conclusion. Our benchmark results show that the generalisation of the trained models is far from perfect, making it infeasible to be deployed to process different types of text. In addition, although intermediate transfer learning is a promising approach to utilising existing resources, further investigation is needed on methods of domain adaptation, particularly cost-effective methods to select useful training instances.
Identifying Health Risks from Family History: A Survey of Natural Language Processing Techniques
Mar 15, 2024Abstract:Electronic health records include information on patients' status and medical history, which could cover the history of diseases and disorders that could be hereditary. One important use of family history information is in precision health, where the goal is to keep the population healthy with preventative measures. Natural Language Processing (NLP) and machine learning techniques can assist with identifying information that could assist health professionals in identifying health risks before a condition is developed in their later years, saving lives and reducing healthcare costs. We survey the literature on the techniques from the NLP field that have been developed to utilise digital health records to identify risks of familial diseases. We highlight that rule-based methods are heavily investigated and are still actively used for family history extraction. Still, more recent efforts have been put into building neural models based on large-scale pre-trained language models. In addition to the areas where NLP has successfully been utilised, we also identify the areas where more research is needed to unlock the value of patients' records regarding data collection, task formulation and downstream applications.
Tensorial tomographic Fourier Ptychography with applications to muscle tissue imaging
May 14, 2023Abstract:We report Tensorial tomographic Fourier Ptychography (ToFu), a new non-scanning label-free tomographic microscopy method for simultaneous imaging of quantitative phase and anisotropic specimen information in 3D. Built upon Fourier Ptychography, a quantitative phase imaging technique, ToFu additionally highlights the vectorial nature of light. The imaging setup consists of a standard microscope equipped with an LED matrix, a polarization generator, and a polarization-sensitive camera. Permittivity tensors of anisotropic samples are computationally recovered from polarized intensity measurements across three dimensions. We demonstrate ToFu's efficiency through volumetric reconstructions of refractive index, birefringence, and orientation for various validation samples, as well as tissue samples from muscle fibers and diseased heart tissue. Our reconstructions of muscle fibers resolve their 3D fine-filament structure and yield consistent morphological measurements compared to gold-standard second harmonic generation scanning confocal microscope images found in the literature. Additionally, we demonstrate reconstructions of a heart tissue sample that carries important polarization information for detecting cardiac amyloidosis.
Detecting Entities in the Astrophysics Literature: A Comparison of Word-based and Span-based Entity Recognition Methods
Nov 24, 2022



Abstract:Information Extraction from scientific literature can be challenging due to the highly specialised nature of such text. We describe our entity recognition methods developed as part of the DEAL (Detecting Entities in the Astrophysics Literature) shared task. The aim of the task is to build a system that can identify Named Entities in a dataset composed by scholarly articles from astrophysics literature. We planned our participation such that it enables us to conduct an empirical comparison between word-based tagging and span-based classification methods. When evaluated on two hidden test sets provided by the organizer, our best-performing submission achieved $F_1$ scores of 0.8307 (validation phase) and 0.7990 (testing phase).
An Exploration of Hierarchical Attention Transformers for Efficient Long Document Classification
Oct 11, 2022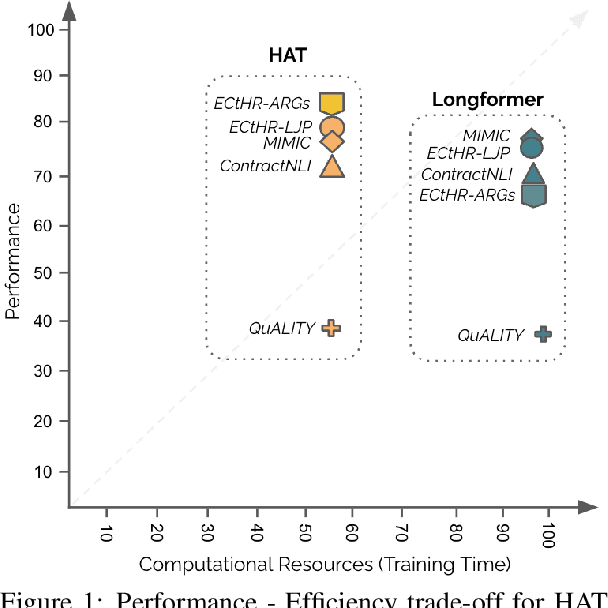
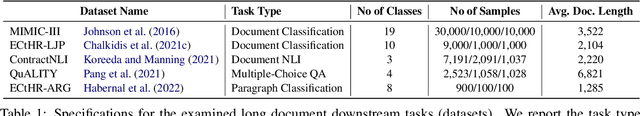
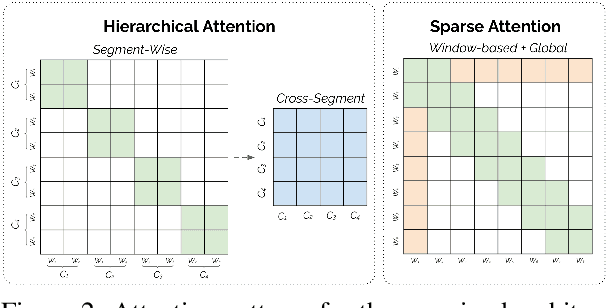
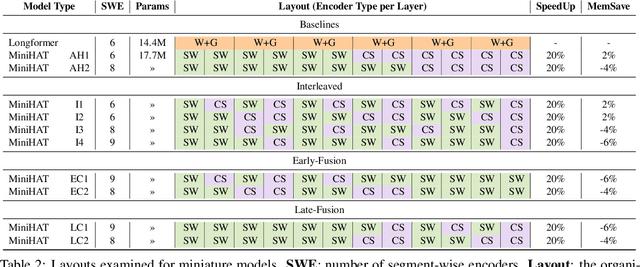
Abstract:Non-hierarchical sparse attention Transformer-based models, such as Longformer and Big Bird, are popular approaches to working with long documents. There are clear benefits to these approaches compared to the original Transformer in terms of efficiency, but Hierarchical Attention Transformer (HAT) models are a vastly understudied alternative. We develop and release fully pre-trained HAT models that use segment-wise followed by cross-segment encoders and compare them with Longformer models and partially pre-trained HATs. In several long document downstream classification tasks, our best HAT model outperforms equally-sized Longformer models while using 10-20% less GPU memory and processing documents 40-45% faster. In a series of ablation studies, we find that HATs perform best with cross-segment contextualization throughout the model than alternative configurations that implement either early or late cross-segment contextualization. Our code is on GitHub: https://github.com/coastalcph/hierarchical-transformers.
Tensorial tomographic differential phase-contrast microscopy
Apr 25, 2022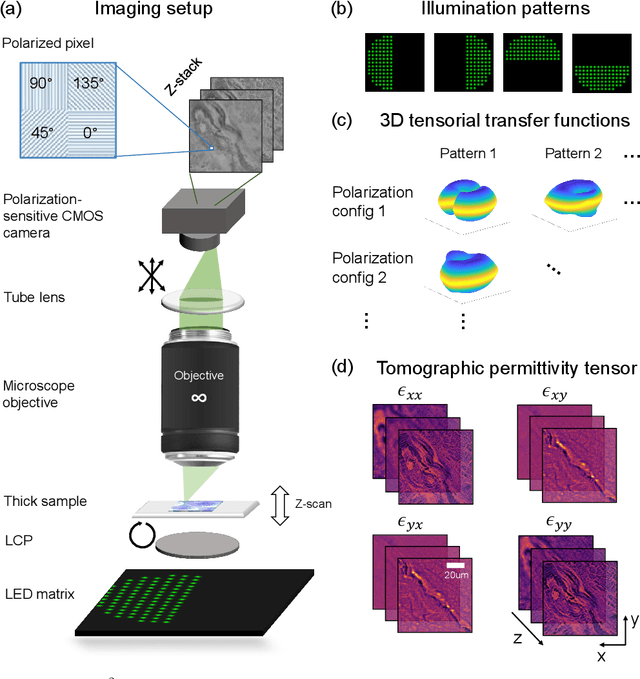
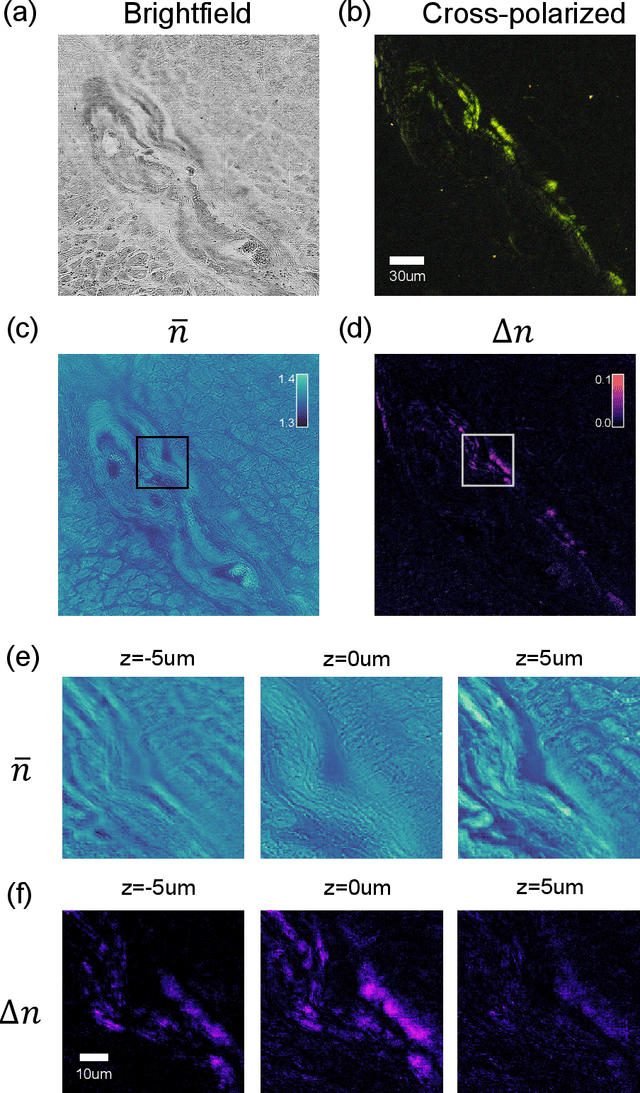
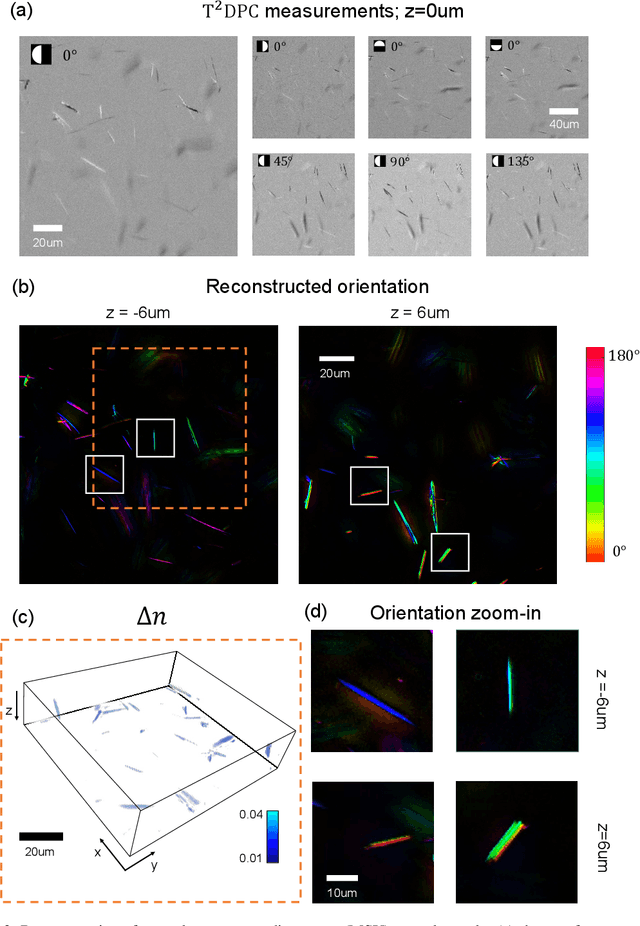
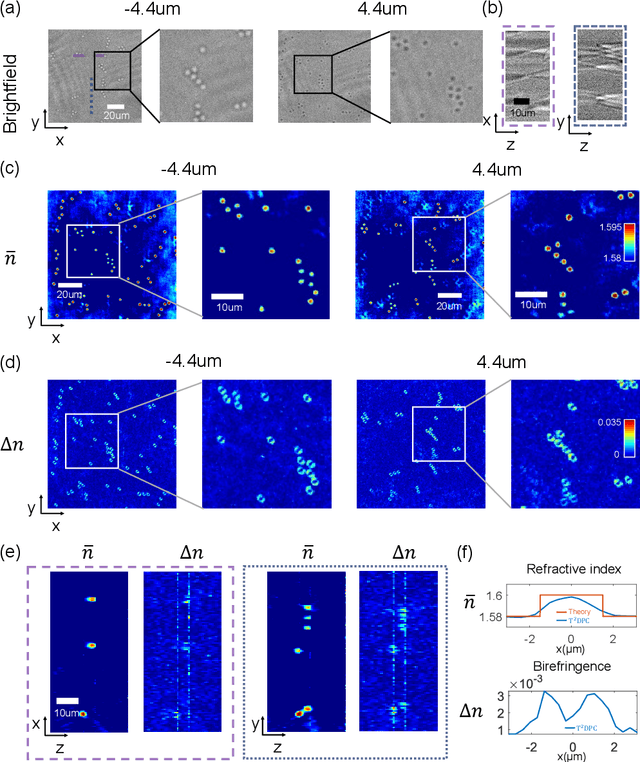

Abstract:We report Tensorial Tomographic Differential Phase-Contrast microscopy (T2DPC), a quantitative label-free tomographic imaging method for simultaneous measurement of phase and anisotropy. T2DPC extends differential phase-contrast microscopy, a quantitative phase imaging technique, to highlight the vectorial nature of light. The method solves for permittivity tensor of anisotropic samples from intensity measurements acquired with a standard microscope equipped with an LED matrix, a circular polarizer, and a polarization-sensitive camera. We demonstrate accurate volumetric reconstructions of refractive index, birefringence, and orientation for various validation samples, and show that the reconstructed polarization structures of a biological specimen are predictive of pathology.
Revisiting Transformer-based Models for Long Document Classification
Apr 14, 2022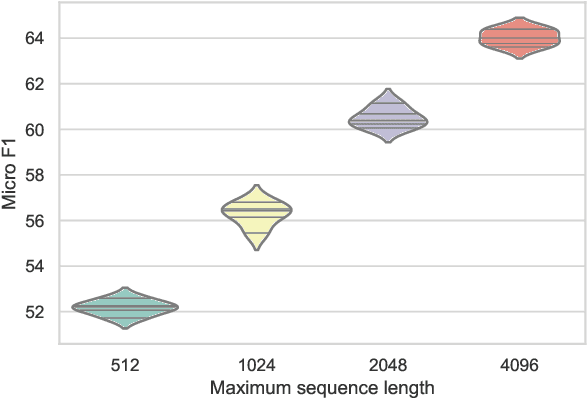
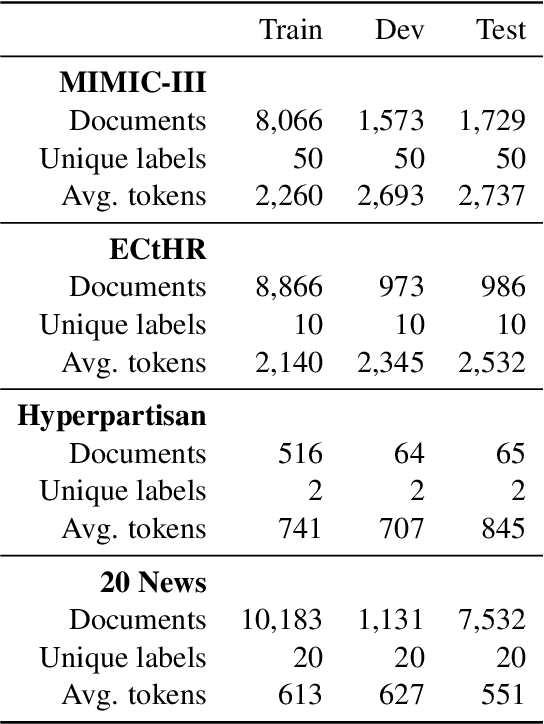
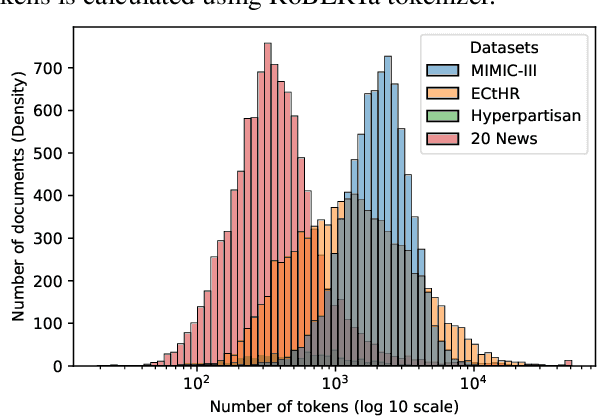
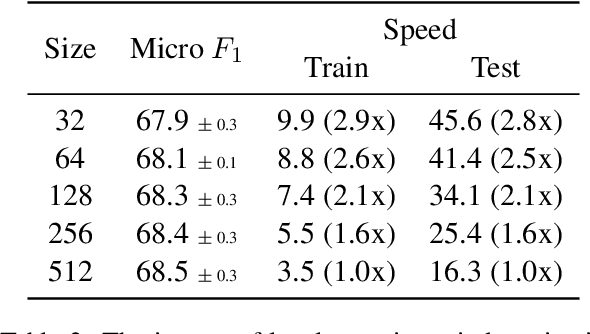
Abstract:The recent literature in text classification is biased towards short text sequences (e.g., sentences or paragraphs). In real-world applications, multi-page multi-paragraph documents are common and they cannot be efficiently encoded by vanilla Transformer-based models. We compare different Transformer-based Long Document Classification (TrLDC) approaches that aim to mitigate the computational overhead of vanilla transformers to encode much longer text, namely sparse attention and hierarchical encoding methods. We examine several aspects of sparse attention (e.g., size of local attention window, use of global attention) and hierarchical (e.g., document splitting strategy) transformers on four document classification datasets covering different domains. We observe a clear benefit from being able to process longer text, and, based on our results, we derive practical advice of applying Transformer-based models on long document classification tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge