Weijie Gan
Dose-aware Diffusion Model for 3D Ultra Low-dose PET Imaging
Nov 07, 2023Abstract:As PET imaging is accompanied by substantial radiation exposure and cancer risk, reducing radiation dose in PET scans is an important topic. Recently, diffusion models have emerged as the new state-of-the-art generative model to generate high-quality samples and have demonstrated strong potential for various tasks in medical imaging. However, it is difficult to extend diffusion models for 3D image reconstructions due to the memory burden. Directly stacking 2D slices together to create 3D image volumes would results in severe inconsistencies between slices. Previous works tried to either applying a penalty term along the z-axis to remove inconsistencies or reconstructing the 3D image volumes with 2 pre-trained perpendicular 2D diffusion models. Nonetheless, these previous methods failed to produce satisfactory results in challenging cases for PET image denoising. In addition to administered dose, the noise-levels in PET images are affected by several other factors in clinical settings, such as scan time, patient size, and weight, etc. Therefore, a method to simultaneously denoise PET images with different noise-levels is needed. Here, we proposed a dose-aware diffusion model for 3D low-dose PET imaging (DDPET) to address these challenges. The proposed DDPET method was tested on 295 patients from three different medical institutions globally with different low-dose levels. These patient data were acquired on three different commercial PET scanners, including Siemens Vision Quadra, Siemens mCT, and United Imaging Healthcare uExplorere. The proposed method demonstrated superior performance over previously proposed diffusion models for 3D imaging problems as well as models proposed for noise-aware medical image denoising. Code is available at: xxx.
A Structured Pruning Algorithm for Model-based Deep Learning
Nov 03, 2023



Abstract:There is a growing interest in model-based deep learning (MBDL) for solving imaging inverse problems. MBDL networks can be seen as iterative algorithms that estimate the desired image using a physical measurement model and a learned image prior specified using a convolutional neural net (CNNs). The iterative nature of MBDL networks increases the test-time computational complexity, which limits their applicability in certain large-scale applications. We address this issue by presenting structured pruning algorithm for model-based deep learning (SPADE) as the first structured pruning algorithm for MBDL networks. SPADE reduces the computational complexity of CNNs used within MBDL networks by pruning its non-essential weights. We propose three distinct strategies to fine-tune the pruned MBDL networks to minimize the performance loss. Each fine-tuning strategy has a unique benefit that depends on the presence of a pre-trained model and a high-quality ground truth. We validate SPADE on two distinct inverse problems, namely compressed sensing MRI and image super-resolution. Our results highlight that MBDL models pruned by SPADE can achieve substantial speed up in testing time while maintaining competitive performance.
PtychoDV: Vision Transformer-Based Deep Unrolling Network for Ptychographic Image Reconstruction
Oct 11, 2023Abstract:Ptychography is an imaging technique that captures multiple overlapping snapshots of a sample, illuminated coherently by a moving localized probe. The image recovery from ptychographic data is generally achieved via an iterative algorithm that solves a nonlinear phase-field problem derived from measured diffraction patterns. However, these approaches have high computational cost. In this paper, we introduce PtychoDV, a novel deep model-based network designed for efficient, high-quality ptychographic image reconstruction. PtychoDV comprises a vision transformer that generates an initial image from the set of raw measurements, taking into consideration their mutual correlations. This is followed by a deep unrolling network that refines the initial image using learnable convolutional priors and the ptychography measurement model. Experimental results on simulated data demonstrate that PtychoDV is capable of outperforming existing deep learning methods for this problem, and significantly reduces computational cost compared to iterative methodologies, while maintaining competitive performance.
A Plug-and-Play Image Registration Network
Oct 06, 2023



Abstract:Deformable image registration (DIR) is an active research topic in biomedical imaging. There is a growing interest in developing DIR methods based on deep learning (DL). A traditional DL approach to DIR is based on training a convolutional neural network (CNN) to estimate the registration field between two input images. While conceptually simple, this approach comes with a limitation that it exclusively relies on a pre-trained CNN without explicitly enforcing fidelity between the registered image and the reference. We present plug-and-play image registration network (PIRATE) as a new DIR method that addresses this issue by integrating an explicit data-fidelity penalty and a CNN prior. PIRATE pre-trains a CNN denoiser on the registration field and "plugs" it into an iterative method as a regularizer. We additionally present PIRATE+ that fine-tunes the CNN prior in PIRATE using deep equilibrium models (DEQ). PIRATE+ interprets the fixed-point iteration of PIRATE as a network with effectively infinite layers and then trains the resulting network end-to-end, enabling it to learn more task-specific information and boosting its performance. Our numerical results on OASIS and CANDI datasets show that our methods achieve state-of-the-art performance on DIR.
Block Coordinate Plug-and-Play Methods for Blind Inverse Problems
May 22, 2023Abstract:Plug-and-play (PnP) prior is a well-known class of methods for solving imaging inverse problems by computing fixed-points of operators combining physical measurement models and learned image denoisers. While PnP methods have been extensively used for image recovery with known measurement operators, there is little work on PnP for solving blind inverse problems. We address this gap by presenting a new block-coordinate PnP (BC-PnP) method that efficiently solves this joint estimation problem by introducing learned denoisers as priors on both the unknown image and the unknown measurement operator. We present a new convergence theory for BC-PnP compatible with blind inverse problems by considering nonconvex data-fidelity terms and expansive denoisers. Our theory analyzes the convergence of BC-PnP to a stationary point of an implicit function associated with an approximate minimum mean-squared error (MMSE) denoiser. We numerically validate our method on two blind inverse problems: automatic coil sensitivity estimation in magnetic resonance imaging (MRI) and blind image deblurring. Our results show that BC-PnP provides an efficient and principled framework for using denoisers as PnP priors for jointly estimating measurement operators and images.
SINCO: A Novel structural regularizer for image compression using implicit neural representations
Oct 26, 2022Abstract:Implicit neural representations (INR) have been recently proposed as deep learning (DL) based solutions for image compression. An image can be compressed by training an INR model with fewer weights than the number of image pixels to map the coordinates of the image to corresponding pixel values. While traditional training approaches for INRs are based on enforcing pixel-wise image consistency, we propose to further improve image quality by using a new structural regularizer. We present structural regularization for INR compression (SINCO) as a novel INR method for image compression. SINCO imposes structural consistency of the compressed images to the groundtruth by using a segmentation network to penalize the discrepancy of segmentation masks predicted from compressed images. We validate SINCO on brain MRI images by showing that it can achieve better performance than some recent INR methods.
CoRRECT: A Deep Unfolding Framework for Motion-Corrected Quantitative R2* Mapping
Oct 12, 2022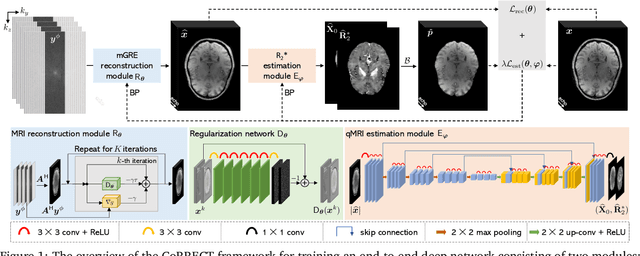
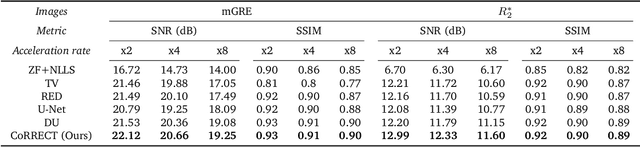
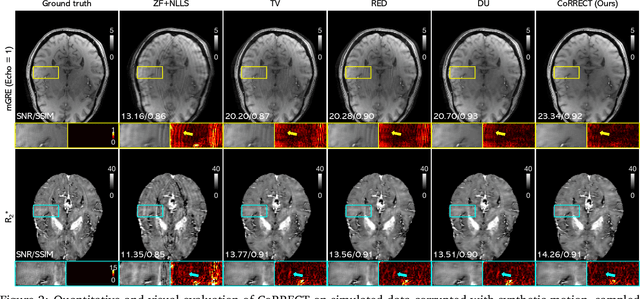
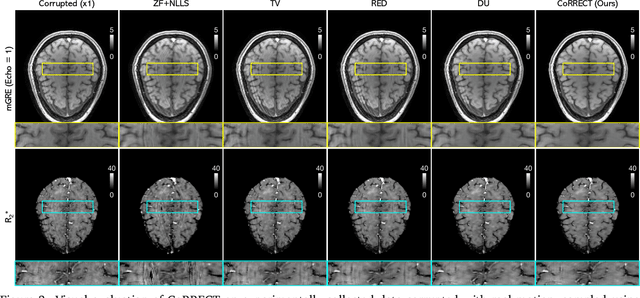
Abstract:Quantitative MRI (qMRI) refers to a class of MRI methods for quantifying the spatial distribution of biological tissue parameters. Traditional qMRI methods usually deal separately with artifacts arising from accelerated data acquisition, involuntary physical motion, and magnetic-field inhomogeneities, leading to suboptimal end-to-end performance. This paper presents CoRRECT, a unified deep unfolding (DU) framework for qMRI consisting of a model-based end-to-end neural network, a method for motion-artifact reduction, and a self-supervised learning scheme. The network is trained to produce R2* maps whose k-space data matches the real data by also accounting for motion and field inhomogeneities. When deployed, CoRRECT only uses the k-space data without any pre-computed parameters for motion or inhomogeneity correction. Our results on experimentally collected multi-Gradient-Recalled Echo (mGRE) MRI data show that CoRRECT recovers motion and inhomogeneity artifact-free R2* maps in highly accelerated acquisition settings. This work opens the door to DU methods that can integrate physical measurement models, biophysical signal models, and learned prior models for high-quality qMRI.
Self-Supervised Deep Equilibrium Models for Inverse Problems with Theoretical Guarantees
Oct 07, 2022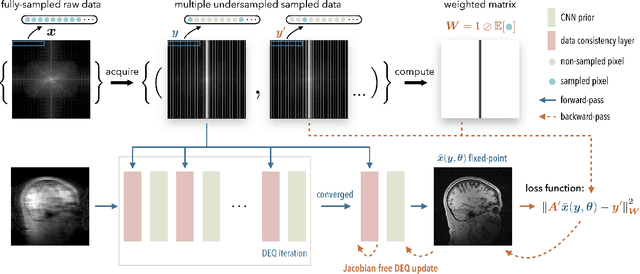
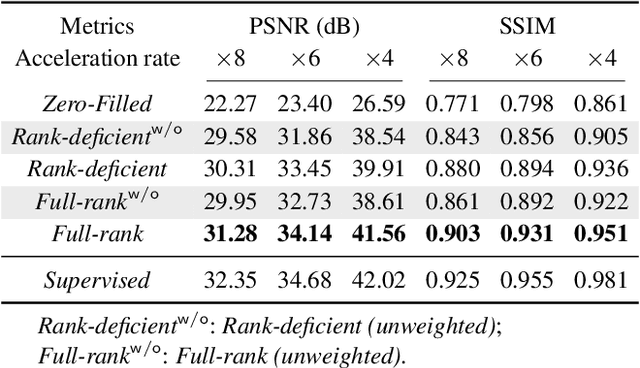
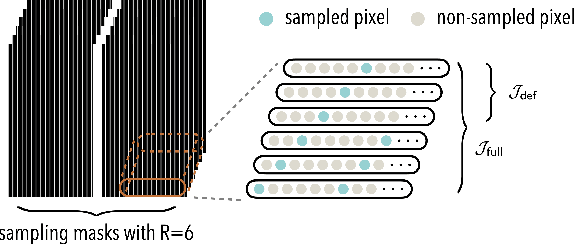
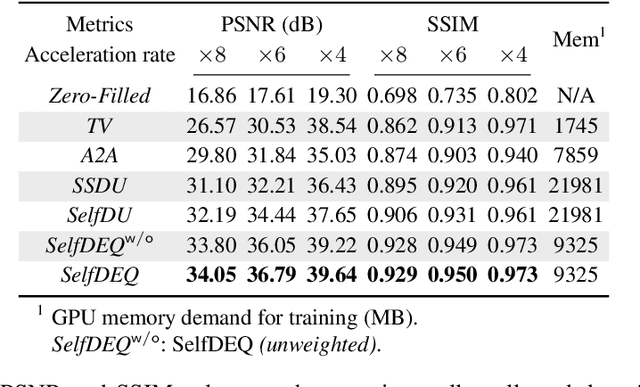
Abstract:Deep equilibrium models (DEQ) have emerged as a powerful alternative to deep unfolding (DU) for image reconstruction. DEQ models-implicit neural networks with effectively infinite number of layers-were shown to achieve state-of-the-art image reconstruction without the memory complexity associated with DU. While the performance of DEQ has been widely investigated, the existing work has primarily focused on the settings where groundtruth data is available for training. We present self-supervised deep equilibrium model (SelfDEQ) as the first self-supervised reconstruction framework for training model-based implicit networks from undersampled and noisy MRI measurements. Our theoretical results show that SelfDEQ can compensate for unbalanced sampling across multiple acquisitions and match the performance of fully supervised DEQ. Our numerical results on in-vivo MRI data show that SelfDEQ leads to state-of-the-art performance using only undersampled and noisy training data.
SPICE: Self-Supervised Learning for MRI with Automatic Coil Sensitivity Estimation
Oct 05, 2022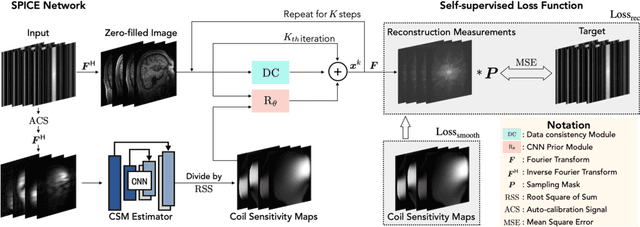
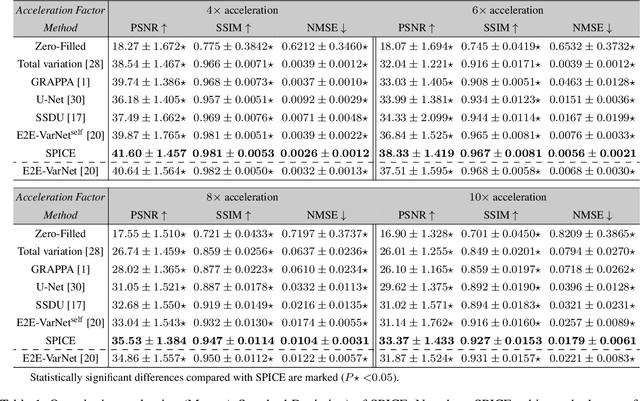
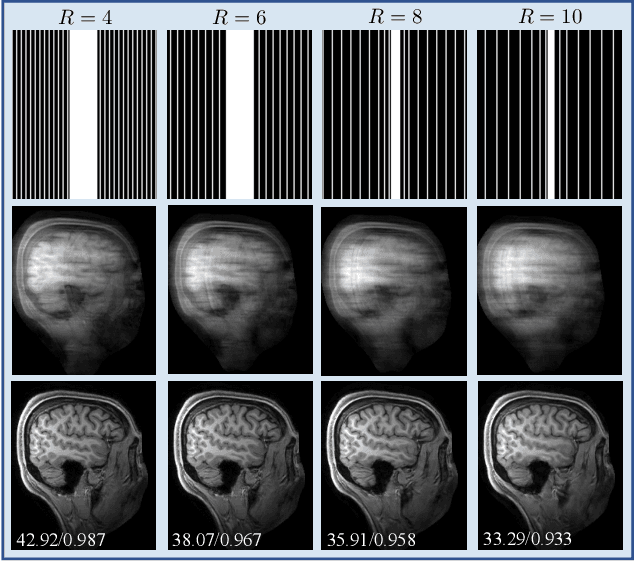
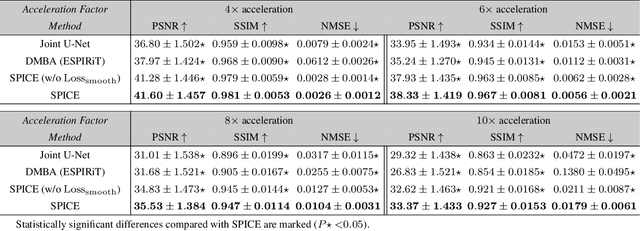
Abstract:Deep model-based architectures (DMBAs) integrating physical measurement models and learned image regularizers are widely used in parallel magnetic resonance imaging (PMRI). Traditional DMBAs for PMRI rely on pre-estimated coil sensitivity maps (CSMs) as a component of the measurement model. However, estimation of accurate CSMs is a challenging problem when measurements are highly undersampled. Additionally, traditional training of DMBAs requires high-quality groundtruth images, limiting their use in applications where groundtruth is difficult to obtain. This paper addresses these issues by presenting SPICE as a new method that integrates self-supervised learning and automatic coil sensitivity estimation. Instead of using pre-estimated CSMs, SPICE simultaneously reconstructs accurate MR images and estimates high-quality CSMs. SPICE also enables learning from undersampled noisy measurements without any groundtruth. We validate SPICE on experimentally collected data, showing that it can achieve state-of-the-art performance in highly accelerated data acquisition settings (up to 10x).
Online Deep Equilibrium Learning for Regularization by Denoising
May 25, 2022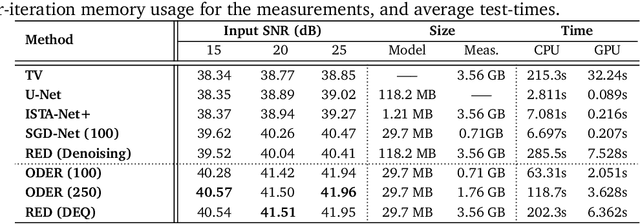
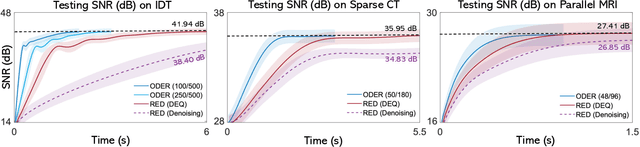
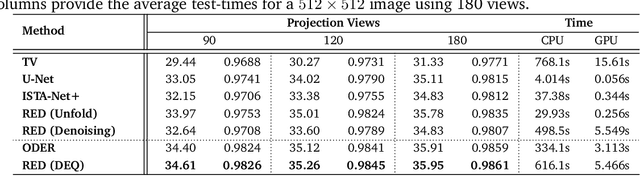
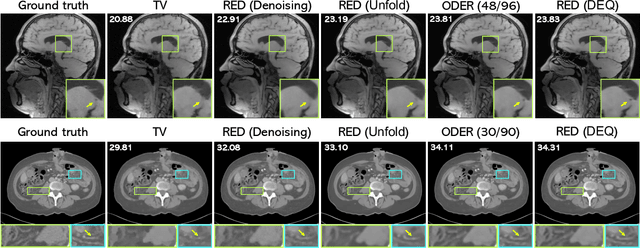
Abstract:Plug-and-Play Priors (PnP) and Regularization by Denoising (RED) are widely-used frameworks for solving imaging inverse problems by computing fixed-points of operators combining physical measurement models and learned image priors. While traditional PnP/RED formulations have focused on priors specified using image denoisers, there is a growing interest in learning PnP/RED priors that are end-to-end optimal. The recent Deep Equilibrium Models (DEQ) framework has enabled memory-efficient end-to-end learning of PnP/RED priors by implicitly differentiating through the fixed-point equations without storing intermediate activation values. However, the dependence of the computational/memory complexity of the measurement models in PnP/RED on the total number of measurements leaves DEQ impractical for many imaging applications. We propose ODER as a new strategy for improving the efficiency of DEQ through stochastic approximations of the measurement models. We theoretically analyze ODER giving insights into its convergence and ability to approximate the traditional DEQ approach. Our numerical results suggest the potential improvements in training/testing complexity due to ODER on three distinct imaging applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge