Wei-Hung Weng
Computer Science and Artificial Intelligence Laboratory, MIT
Explainable Deep Learning in Healthcare: A Methodological Survey from an Attribution View
Dec 05, 2021

Abstract:The increasing availability of large collections of electronic health record (EHR) data and unprecedented technical advances in deep learning (DL) have sparked a surge of research interest in developing DL based clinical decision support systems for diagnosis, prognosis, and treatment. Despite the recognition of the value of deep learning in healthcare, impediments to further adoption in real healthcare settings remain due to the black-box nature of DL. Therefore, there is an emerging need for interpretable DL, which allows end users to evaluate the model decision making to know whether to accept or reject predictions and recommendations before an action is taken. In this review, we focus on the interpretability of the DL models in healthcare. We start by introducing the methods for interpretability in depth and comprehensively as a methodological reference for future researchers or clinical practitioners in this field. Besides the methods' details, we also include a discussion of advantages and disadvantages of these methods and which scenarios each of them is suitable for, so that interested readers can know how to compare and choose among them for use. Moreover, we discuss how these methods, originally developed for solving general-domain problems, have been adapted and applied to healthcare problems and how they can help physicians better understand these data-driven technologies. Overall, we hope this survey can help researchers and practitioners in both artificial intelligence (AI) and clinical fields understand what methods we have for enhancing the interpretability of their DL models and choose the optimal one accordingly.
Addressing the Real-world Class Imbalance Problem in Dermatology
Oct 09, 2020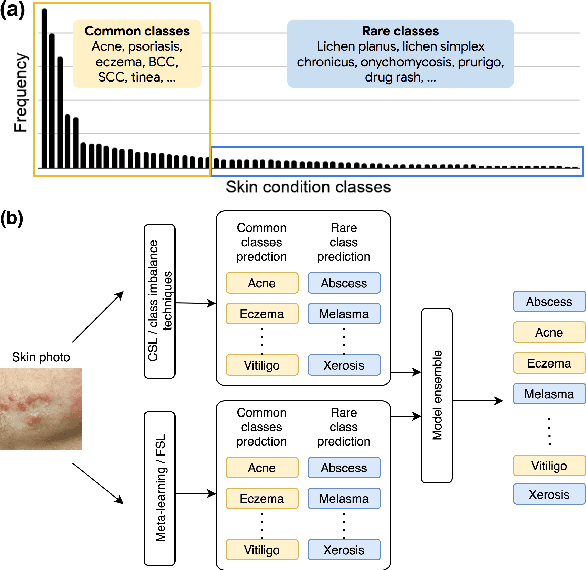
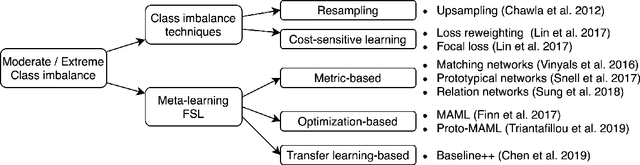
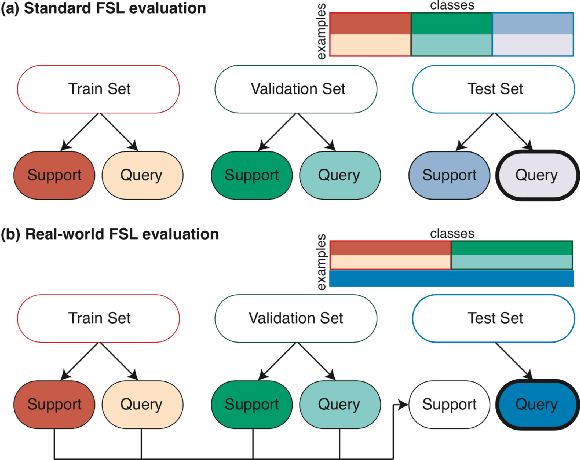
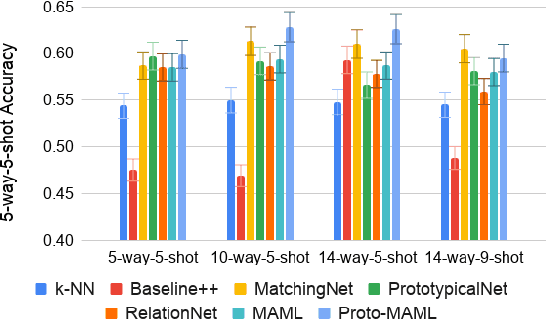
Abstract:Class imbalance is a common problem in medical diagnosis, causing a standard classifier to be biased towards the common classes and perform poorly on the rare classes. This is especially true for dermatology, a specialty with thousands of skin conditions but many of which have rare prevalence in the real world. Motivated by recent advances, we explore few-shot learning methods as well as conventional class imbalance techniques for the skin condition recognition problem and propose an evaluation setup to fairly assess the real-world utility of such approaches. When compared to conventional class imbalance techniques, we find that few-shot learning methods are not as performant as those conventional methods, but combining the two approaches using a novel ensemble leads to improvement in model performance, especially for rare classes. We conclude that the ensemble can be useful to address the class imbalance problem, yet progress here can further be accelerated by the use of real-world evaluation setups for benchmarking new methods.
What Disease does this Patient Have? A Large-scale Open Domain Question Answering Dataset from Medical Exams
Sep 28, 2020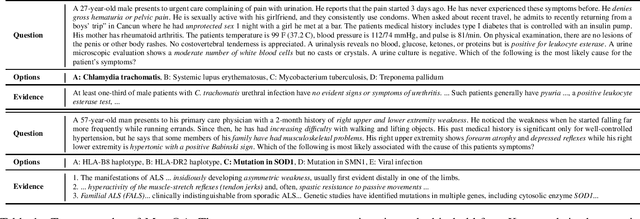
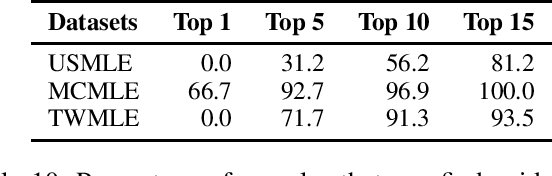

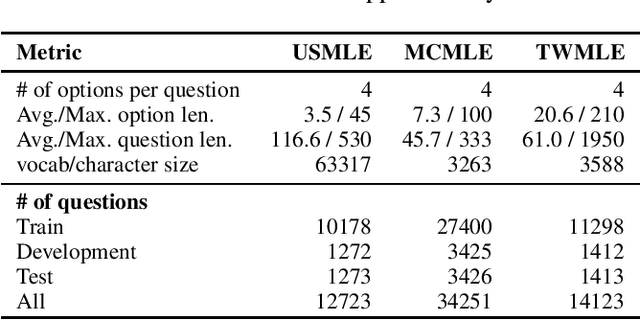
Abstract:Open domain question answering (OpenQA) tasks have been recently attracting more and more attention from the natural language processing (NLP) community. In this work, we present the first free-form multiple-choice OpenQA dataset for solving medical problems, MedQA, collected from the professional medical board exams. It covers three languages: English, simplified Chinese, and traditional Chinese, and contains 12,723, 34,251, and 14,123 questions for the three languages, respectively. We implement both rule-based and popular neural methods by sequentially combining a document retriever and a machine comprehension model. Through experiments, we find that even the current best method can only achieve 36.7\%, 42.0\%, and 70.1\% of test accuracy on the English, traditional Chinese, and simplified Chinese questions, respectively. We expect MedQA to present great challenges to existing OpenQA systems and hope that it can serve as a platform to promote much stronger OpenQA models from the NLP community in the future.
CheXpert++: Approximating the CheXpert labeler for Speed,Differentiability, and Probabilistic Output
Jun 26, 2020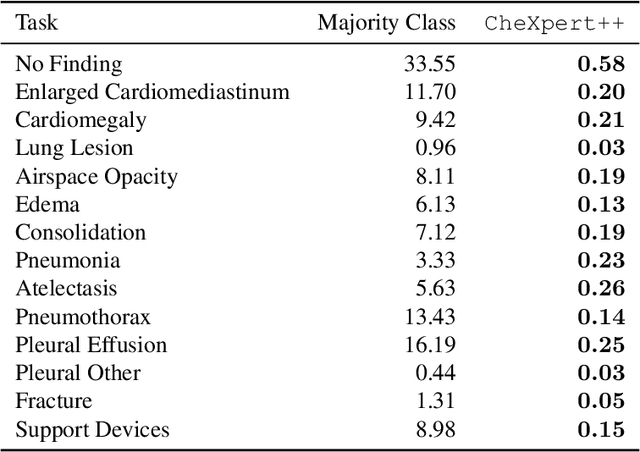
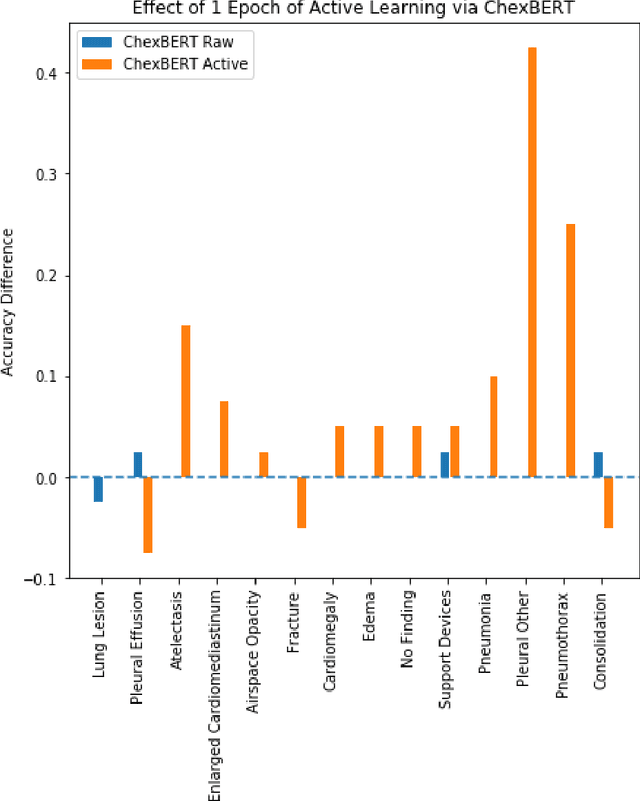
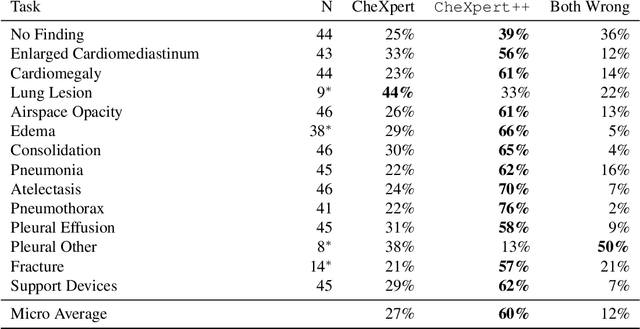
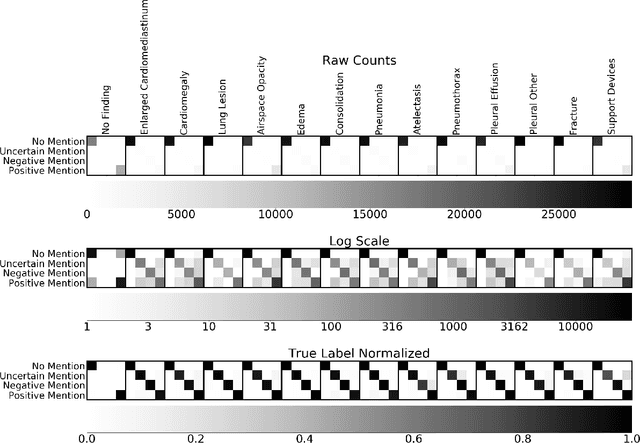
Abstract:It is often infeasible or impossible to obtain ground truth labels for medical data. To circumvent this, one may build rule-based or other expert-knowledge driven labelers to ingest data and yield silver labels absent any ground-truth training data. One popular such labeler is CheXpert, a labeler that produces diagnostic labels for chest X-ray radiology reports. CheXpert is very useful, but is relatively computationally slow, especially when integrated with end-to-end neural pipelines, is non-differentiable so can't be used in any applications that require gradients to flow through the labeler, and does not yield probabilistic outputs, which limits our ability to improve the quality of the silver labeler through techniques such as active learning. In this work, we solve all three of these problems with $\texttt{CheXpert++}$, a BERT-based, high-fidelity approximation to CheXpert. $\texttt{CheXpert++}$ achieves 99.81\% parity with CheXpert, which means it can be reliably used as a drop-in replacement for CheXpert, all while being significantly faster, fully differentiable, and probabilistic in output. Error analysis of $\texttt{CheXpert++}$ also demonstrates that $\texttt{CheXpert++}$ has a tendency to actually correct errors in the CheXpert labels, with $\texttt{CheXpert++}$ labels being more often preferred by a clinician over CheXpert labels (when they disagree) on all but one disease task. To further demonstrate the utility of these advantages in this model, we conduct a proof-of-concept active learning study, demonstrating we can improve accuracy on an expert labeled random subset of report sentences by approximately 8\% over raw, unaltered CheXpert by using one-iteration of active-learning inspired re-training. These findings suggest that simple techniques in co-learning and active learning can yield high-quality labelers under minimal, and controllable human labeling demands.
Entity-Enriched Neural Models for Clinical Question Answering
May 13, 2020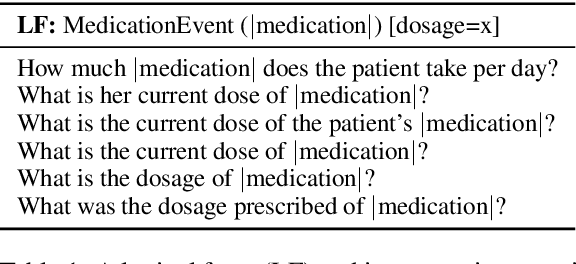
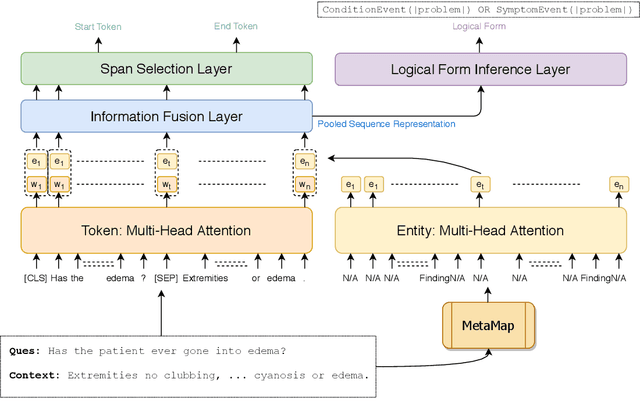
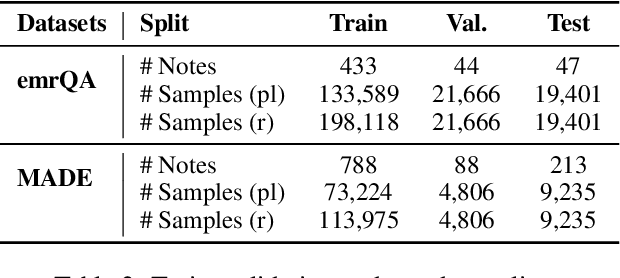
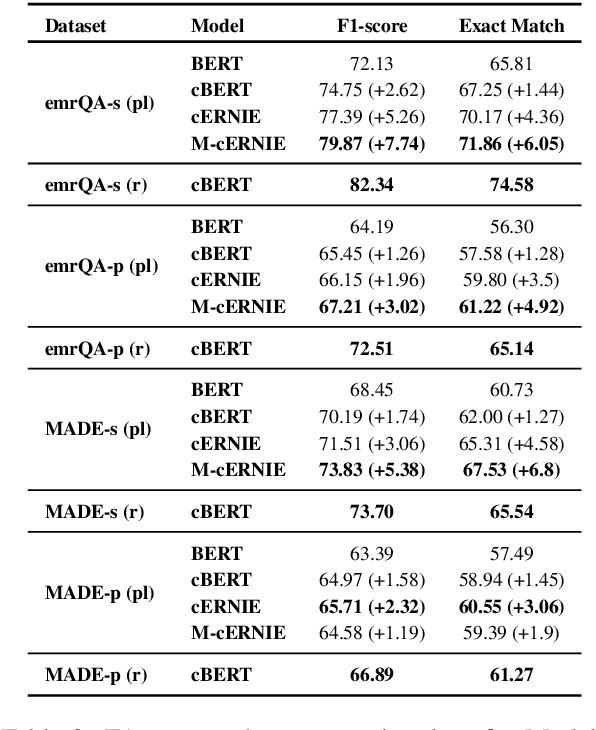
Abstract:We explore state-of-the-art neural models for question answering on electronic medical records and improve their ability to generalize better on previously unseen (paraphrased) questions at test time. We enable this by learning to predict logical forms as an auxiliary task along with the main task of answer span detection. The predicted logical forms also serve as a rationale for the answer. Further, we also incorporate medical entity information in these models via the ERNIE architecture. We train our models on the large-scale emrQA dataset and observe that our multi-task entity-enriched models generalize to paraphrased questions ~5% better than the baseline BERT model.
Weakly Supervised Context Encoder using DICOM metadata in Ultrasound Imaging
Mar 20, 2020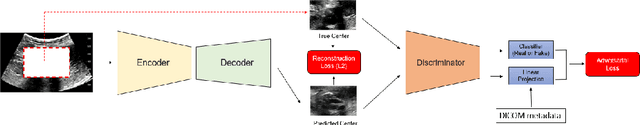
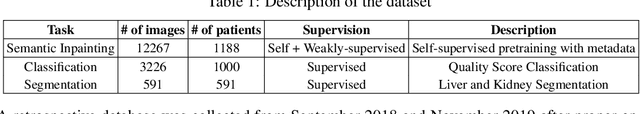
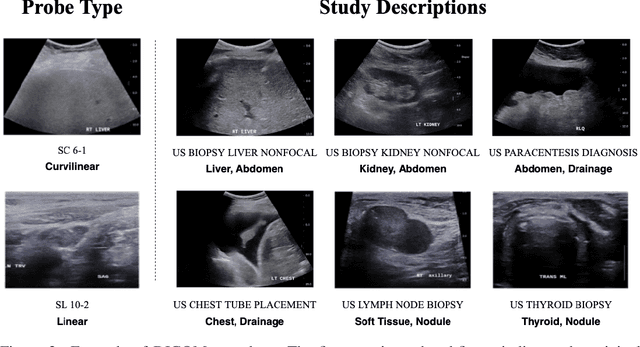
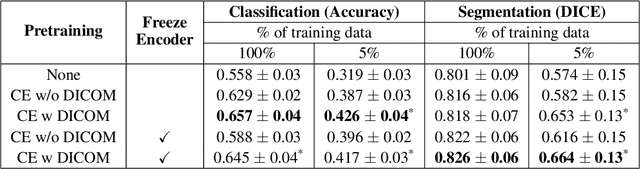
Abstract:Modern deep learning algorithms geared towards clinical adaption rely on a significant amount of high fidelity labeled data. Low-resource settings pose challenges like acquiring high fidelity data and becomes the bottleneck for developing artificial intelligence applications. Ultrasound images, stored in Digital Imaging and Communication in Medicine (DICOM) format, have additional metadata data corresponding to ultrasound image parameters and medical exams. In this work, we leverage DICOM metadata from ultrasound images to help learn representations of the ultrasound image. We demonstrate that the proposed method outperforms the non-metadata based approaches across different downstream tasks.
Clinical Text Summarization with Syntax-Based Negation and Semantic Concept Identification
Feb 29, 2020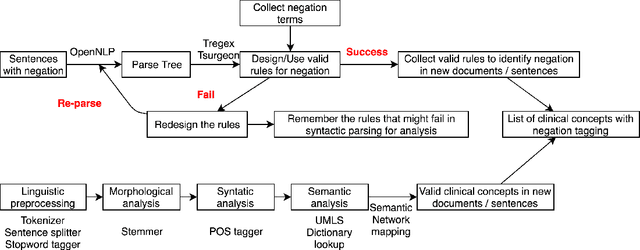
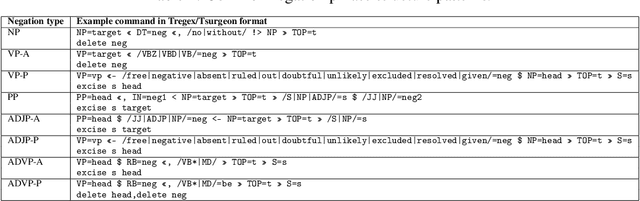


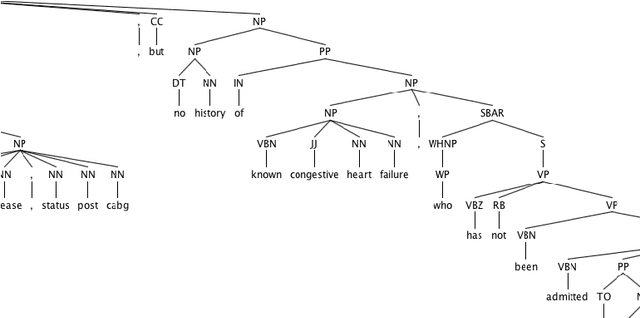
Abstract:In the era of clinical information explosion, a good strategy for clinical text summarization is helpful to improve the clinical workflow. The ideal summarization strategy can preserve important information in the informative but less organized, ill-structured clinical narrative texts. Instead of using pure statistical learning approaches, which are difficult to interpret and explain, we utilized knowledge of computational linguistics with human experts-curated biomedical knowledge base to achieve the interpretable and meaningful clinical text summarization. Our research objective is to use the biomedical ontology with semantic information, and take the advantage from the language hierarchical structure, the constituency tree, in order to identify the correct clinical concepts and the corresponding negation information, which is critical for summarizing clinical concepts from narrative text. We achieved the clinically acceptable performance for both negation detection and concept identification, and the clinical concepts with common negated patterns can be identified and negated by the proposed method.
Human-centric Metric for Accelerating Pathology Reports Annotation
Nov 12, 2019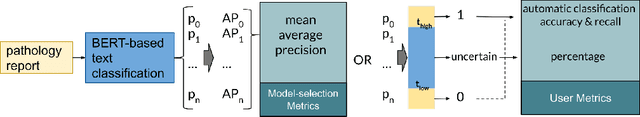
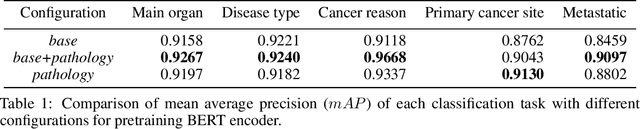
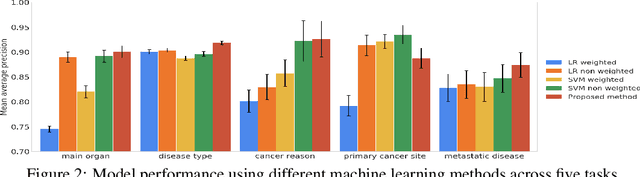
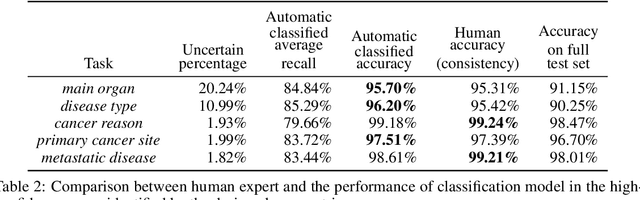
Abstract:Pathology reports contain useful information such as the main involved organ, diagnosis, etc. These information can be identified from the free text reports and used for large-scale statistical analysis or serve as annotation for other modalities such as pathology slides images. However, manual classification for a huge number of reports on multiple tasks is labor-intensive. In this paper, we have developed an automatic text classifier based on BERT and we propose a human-centric metric to evaluate the model. According to the model confidence, we identify low-confidence cases that require further expert annotation and high-confidence cases that are automatically classified. We report the percentage of low-confidence cases and the performance of automatically classified cases. On the high-confidence cases, the model achieves classification accuracy comparable to pathologists. This leads a potential of reducing 80% to 98% of the manual annotation workload.
Representation Learning for Electronic Health Records
Sep 19, 2019Abstract:Information in electronic health records (EHR), such as clinical narratives, examination reports, lab measurements, demographics, and other patient encounter entries, can be transformed into appropriate data representations that can be used for downstream clinical machine learning tasks using representation learning. Learning better representations is critical to improve the performance of downstream tasks. Due to the advances in machine learning, we now can learn better and meaningful representations from EHR through disentangling the underlying factors inside data and distilling large amounts of information and knowledge from heterogeneous EHR sources. In this chapter, we first introduce the background of learning representations and reasons why we need good EHR representations in machine learning for medicine and healthcare in Section 1. Next, we explain the commonly-used machine learning and evaluation methods for representation learning using a deep learning approach in Section 2. Following that, we review recent related studies of learning patient state representation from EHR for clinical machine learning tasks in Section 3. Finally, in Section 4 we discuss more techniques, studies, and challenges for learning natural language representations when free texts, such as clinical notes, examination reports, or biomedical literature are used. We also discuss challenges and opportunities in these rapidly growing research fields.
Machine Learning for Clinical Predictive Analytics
Sep 19, 2019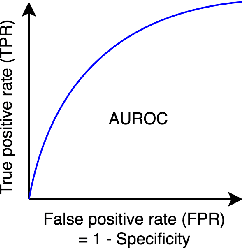
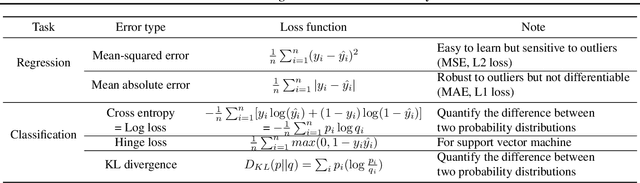

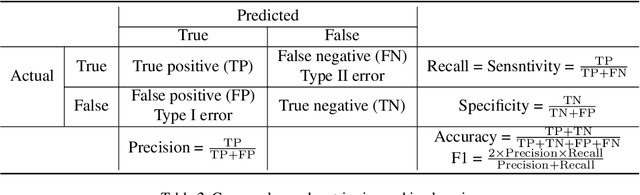
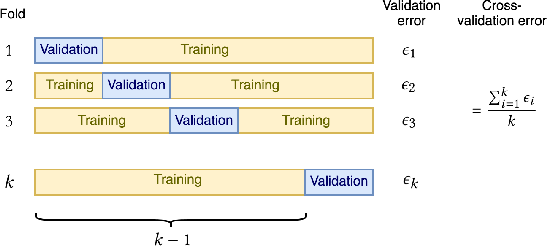
Abstract:In this chapter, we provide a brief overview of applying machine learning techniques for clinical prediction tasks. We begin with a quick introduction to the concepts of machine learning and outline some of the most common machine learning algorithms. Next, we demonstrate how to apply the algorithms with appropriate toolkits to conduct machine learning experiments for clinical prediction tasks. The objectives of this chapter are to (1) understand the basics of machine learning techniques and the reasons behind why they are useful for solving clinical prediction problems, (2) understand the intuition behind some machine learning models, including regression, decision trees, and support vector machines, and (3) understand how to apply these models to clinical prediction problems using publicly available datasets via case studies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge