Tuan Truong
Revisiting Integration of Image and Metadata for DICOM Series Classification: Cross-Attention and Dictionary Learning
Feb 27, 2026Abstract:Automated identification of DICOM image series is essential for large-scale medical image analysis, quality control, protocol harmonization, and reliable downstream processing. However, DICOM series classification remains challenging due to heterogeneous slice content, variable series length, and entirely missing, incomplete or inconsistent DICOM metadata. We propose an end-to-end multimodal framework for DICOM series classification that jointly models image content and acquisition metadata while explicitly accounting for all these challenges. (i) Images and metadata are encoded with modality-aware modules and fused using a bi-directional cross-modal attention mechanism. (ii) Metadata is processed by a sparse, missingness-aware encoder based on learnable feature dictionaries and value-conditioned modulation. By design, the approach does not require any form of imputation. (iii) Variability in series length and image data dimensions is handled via a 2.5D visual encoder and attention operating on equidistantly sampled slices. We evaluate the proposed approach on the publicly available Duke Liver MRI dataset and a large multi-institutional in-house cohort, assessing both in-domain performance and out-of-domain generalization. Across all evaluation settings, the proposed method consistently outperforms relevant image only, metadata-only and multimodal 2D/3D baselines. The results demonstrate that explicitly modeling metadata sparsity and cross-modal interactions improves robustness for DICOM series classification.
DoRAN: Stabilizing Weight-Decomposed Low-Rank Adaptation via Noise Injection and Auxiliary Networks
Oct 05, 2025



Abstract:Parameter-efficient fine-tuning (PEFT) methods have become the standard paradigm for adapting large-scale models. Among these techniques, Weight-Decomposed Low-Rank Adaptation (DoRA) has been shown to improve both the learning capacity and training stability of the vanilla Low-Rank Adaptation (LoRA) method by explicitly decomposing pre-trained weights into magnitude and directional components. In this work, we propose DoRAN, a new variant of DoRA designed to further stabilize training and boost the sample efficiency of DoRA. Our approach includes two key stages: (i) injecting noise into the denominator of DoRA's weight decomposition, which serves as an adaptive regularizer to mitigate instabilities; and (ii) replacing static low-rank matrices with auxiliary networks that generate them dynamically, enabling parameter coupling across layers and yielding better sample efficiency in both theory and practice. Comprehensive experiments on vision and language benchmarks show that DoRAN consistently outperforms LoRA, DoRA, and other PEFT baselines. These results underscore the effectiveness of combining stabilization through noise-based regularization with network-based parameter generation, offering a promising direction for robust and efficient fine-tuning of foundation models.
HoRA: Cross-Head Low-Rank Adaptation with Joint Hypernetworks
Oct 05, 2025Abstract:Low-Rank Adaptation (LoRA) is a parameter-efficient fine-tuning (PEFT) technique that adapts large pre-trained models by adding low-rank matrices to their weight updates. However, in the context of fine-tuning multi-head self-attention (MHA), LoRA has been employed to adapt each attention head separately, thereby overlooking potential synergies across different heads. To mitigate this issue, we propose a novel Hyper-shared Low-Rank Adaptation (HoRA) method, which utilizes joint hypernetworks to generate low-rank matrices across attention heads. By coupling their adaptation through a shared generator, HoRA encourages cross-head information sharing, and thus directly addresses the aforementioned limitation of LoRA. By comparing LoRA and HoRA through the lens of hierarchical mixture of experts, our theoretical findings reveal that the latter achieves superior sample efficiency to the former. Furthermore, through extensive experiments across diverse language and vision benchmarks, we demonstrate that HoRA outperforms LoRA and other PEFT methods while requiring only a marginal increase in the number of trainable parameters.
AstroCompress: A benchmark dataset for multi-purpose compression of astronomical data
Jun 10, 2025



Abstract:The site conditions that make astronomical observatories in space and on the ground so desirable -- cold and dark -- demand a physical remoteness that leads to limited data transmission capabilities. Such transmission limitations directly bottleneck the amount of data acquired and in an era of costly modern observatories, any improvements in lossless data compression has the potential scale to billions of dollars worth of additional science that can be accomplished on the same instrument. Traditional lossless methods for compressing astrophysical data are manually designed. Neural data compression, on the other hand, holds the promise of learning compression algorithms end-to-end from data and outperforming classical techniques by leveraging the unique spatial, temporal, and wavelength structures of astronomical images. This paper introduces AstroCompress: a neural compression challenge for astrophysics data, featuring four new datasets (and one legacy dataset) with 16-bit unsigned integer imaging data in various modes: space-based, ground-based, multi-wavelength, and time-series imaging. We provide code to easily access the data and benchmark seven lossless compression methods (three neural and four non-neural, including all practical state-of-the-art algorithms). Our results on lossless compression indicate that lossless neural compression techniques can enhance data collection at observatories, and provide guidance on the adoption of neural compression in scientific applications. Though the scope of this paper is restricted to lossless compression, we also comment on the potential exploration of lossy compression methods in future studies.
Promoting Ensemble Diversity with Interactive Bayesian Distributional Robustness for Fine-tuning Foundation Models
Jun 08, 2025Abstract:We introduce Interactive Bayesian Distributional Robustness (IBDR), a novel Bayesian inference framework that allows modeling the interactions between particles, thereby enhancing ensemble quality through increased particle diversity. IBDR is grounded in a generalized theoretical framework that connects the distributional population loss with the approximate posterior, motivating a practical dual optimization procedure that enforces distributional robustness while fostering particle diversity. We evaluate IBDR's performance against various baseline methods using the VTAB-1K benchmark and the common reasoning language task. The results consistently show that IBDR outperforms these baselines, underscoring its effectiveness in real-world applications.
RepLoRA: Reparameterizing Low-Rank Adaptation via the Perspective of Mixture of Experts
Feb 05, 2025Abstract:Low-rank adaptation (LoRA) has emerged as a powerful method for fine-tuning large-scale foundation models. Despite its popularity, the theoretical understanding of LoRA has remained limited. This paper presents a theoretical analysis of LoRA by examining its connection to the Mixture of Experts models. Under this framework, we show that simple reparameterizations of the LoRA matrices can notably accelerate the low-rank matrix estimation process. In particular, we prove that reparameterization can reduce the data needed to achieve a desired estimation error from an exponential to a polynomial scale. Motivated by this insight, we propose Reparameterized Low-rank Adaptation (RepLoRA), which incorporates lightweight MLPs to reparameterize the LoRA matrices. Extensive experiments across multiple domains demonstrate that RepLoRA consistently outperforms vanilla LoRA. Notably, with limited data, RepLoRA surpasses LoRA by a margin of up to 40.0% and achieves LoRA's performance with only 30.0% of the training data, highlighting both the theoretical and empirical robustness of our PEFT method.
Exploring AI-based System Design for Pixel-level Protected Health Information Detection in Medical Images
Jan 16, 2025



Abstract:De-identification of medical images is a critical step to ensure privacy during data sharing in research and clinical settings. The initial step in this process involves detecting Protected Health Information (PHI), which can be found in image metadata or imprinted within image pixels. Despite the importance of such systems, there has been limited evaluation of existing AI-based solutions, creating barriers to the development of reliable and robust tools. In this study, we present an AI-based pipeline for PHI detection, comprising three key components: text detection, text extraction, and analysis of PHI content in medical images. By experimenting with exchanging roles of vision and language models within the pipeline, we evaluate the performance and recommend the best setup for the PHI detection task.
Leveraging Hierarchical Taxonomies in Prompt-based Continual Learning
Oct 06, 2024



Abstract:Drawing inspiration from human learning behaviors, this work proposes a novel approach to mitigate catastrophic forgetting in Prompt-based Continual Learning models by exploiting the relationships between continuously emerging class data. We find that applying human habits of organizing and connecting information can serve as an efficient strategy when training deep learning models. Specifically, by building a hierarchical tree structure based on the expanding set of labels, we gain fresh insights into the data, identifying groups of similar classes could easily cause confusion. Additionally, we delve deeper into the hidden connections between classes by exploring the original pretrained model's behavior through an optimal transport-based approach. From these insights, we propose a novel regularization loss function that encourages models to focus more on challenging knowledge areas, thereby enhancing overall performance. Experimentally, our method demonstrated significant superiority over the most robust state-of-the-art models on various benchmarks.
Improving Generalization with Flat Hilbert Bayesian Inference
Oct 05, 2024
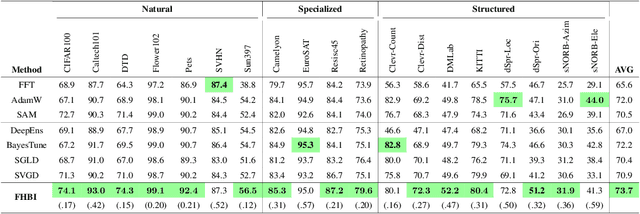
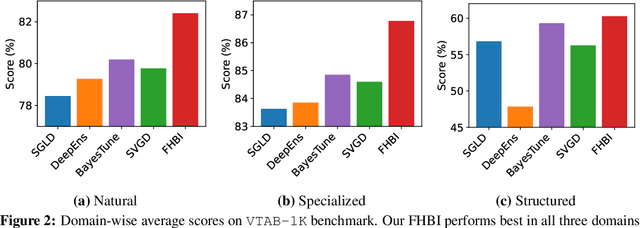

Abstract:We introduce Flat Hilbert Bayesian Inference (FHBI), an algorithm designed to enhance generalization in Bayesian inference. Our approach involves an iterative two-step procedure with an adversarial functional perturbation step and a functional descent step within the reproducing kernel Hilbert spaces. This methodology is supported by a theoretical analysis that extends previous findings on generalization ability from finite-dimensional Euclidean spaces to infinite-dimensional functional spaces. To evaluate the effectiveness of FHBI, we conduct comprehensive comparisons against seven baseline methods on the VTAB-1K benchmark, which encompasses 19 diverse datasets across various domains with diverse semantics. Empirical results demonstrate that FHBI consistently outperforms the baselines by notable margins, highlighting its practical efficacy.
Five Pitfalls When Assessing Synthetic Medical Images with Reference Metrics
Aug 12, 2024Abstract:Reference metrics have been developed to objectively and quantitatively compare two images. Especially for evaluating the quality of reconstructed or compressed images, these metrics have shown very useful. Extensive tests of such metrics on benchmarks of artificially distorted natural images have revealed which metric best correlate with human perception of quality. Direct transfer of these metrics to the evaluation of generative models in medical imaging, however, can easily lead to pitfalls, because assumptions about image content, image data format and image interpretation are often very different. Also, the correlation of reference metrics and human perception of quality can vary strongly for different kinds of distortions and commonly used metrics, such as SSIM, PSNR and MAE are not the best choice for all situations. We selected five pitfalls that showcase unexpected and probably undesired reference metric scores and discuss strategies to avoid them.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge