Stefan Klein
Department of Radiology and Nuclear Medicine, Erasmus MC Cancer Institute, University Medical Center Rotterdam, Rotterdam, the Netherlands
Longitudinal diffusion MRI analysis using Segis-Net: a single-step deep-learning framework for simultaneous segmentation and registration
Dec 28, 2020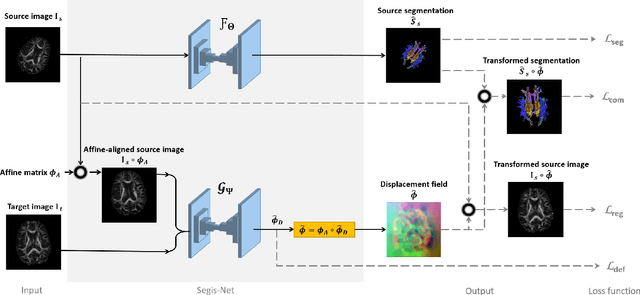
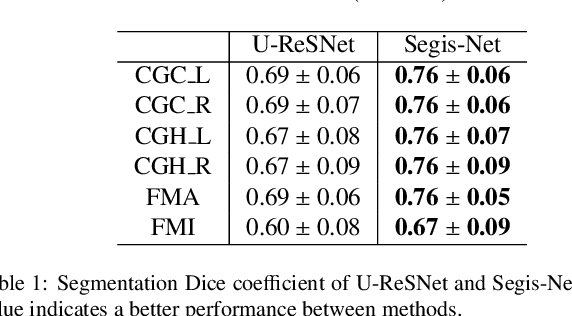
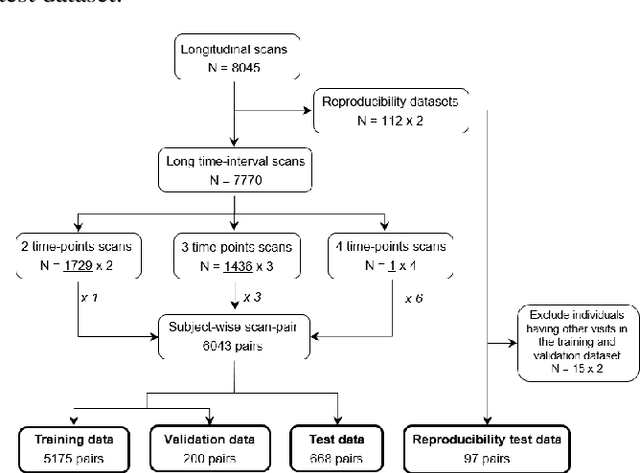
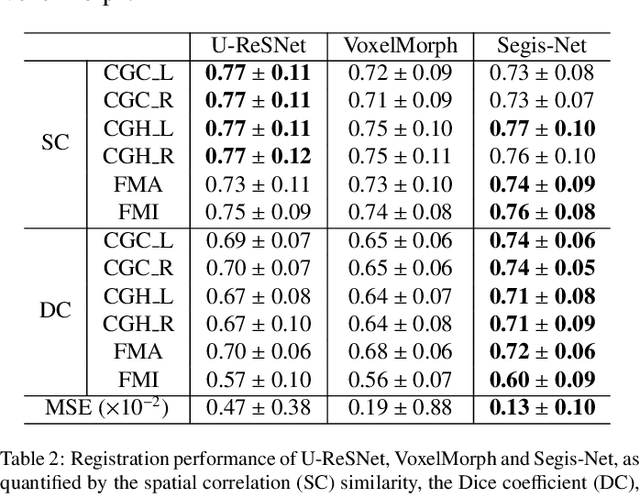
Abstract:This work presents a single-step deep-learning framework for longitudinal image analysis, coined Segis-Net. To optimally exploit information available in longitudinal data, this method concurrently learns a multi-class segmentation and nonlinear registration. Segmentation and registration are modeled using a convolutional neural network and optimized simultaneously for their mutual benefit. An objective function that optimizes spatial correspondence for the segmented structures across time-points is proposed. We applied Segis-Net to the analysis of white matter tracts from N=8045 longitudinal brain MRI datasets of 3249 elderly individuals. Segis-Net approach showed a significant increase in registration accuracy, spatio-temporal segmentation consistency, and reproducibility comparing with two multistage pipelines. This also led to a significant reduction in the sample-size that would be required to achieve the same statistical power in analyzing tract-specific measures. Thus, we expect that Segis-Net can serve as a new reliable tool to support longitudinal imaging studies to investigate macro- and microstructural brain changes over time.
Cross-Cohort Generalizability of Deep and Conventional Machine Learning for MRI-based Diagnosis and Prediction of Alzheimer's Disease
Dec 16, 2020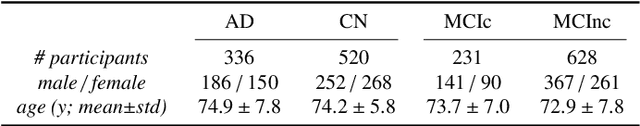
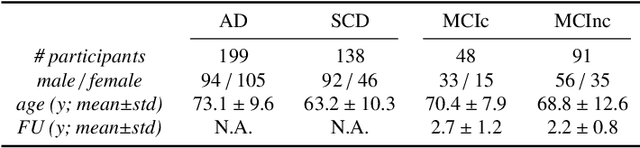
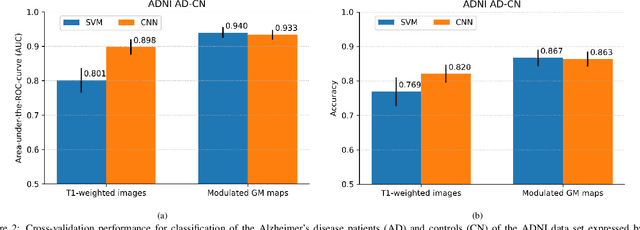
Abstract:This work validates the generalizability of MRI-based classification of Alzheimer's disease (AD) patients and controls (CN) to an external data set and to the task of prediction of conversion to AD in individuals with mild cognitive impairment (MCI). We used a conventional support vector machine (SVM) and a deep convolutional neural network (CNN) approach based on structural MRI scans that underwent either minimal pre-processing or more extensive pre-processing into modulated gray matter (GM) maps. Classifiers were optimized and evaluated using cross-validation in the ADNI (334 AD, 520 CN). Trained classifiers were subsequently applied to predict conversion to AD in ADNI MCI patients (231 converters, 628 non-converters) and in the independent Health-RI Parelsnoer data set. From this multi-center study representing a tertiary memory clinic population, we included 199 AD patients, 139 participants with subjective cognitive decline, 48 MCI patients converting to dementia, and 91 MCI patients who did not convert to dementia. AD-CN classification based on modulated GM maps resulted in a similar AUC for SVM (0.940) and CNN (0.933). Application to conversion prediction in MCI yielded significantly higher performance for SVM (0.756) than for CNN (0.742). In external validation, performance was slightly decreased. For AD-CN, it again gave similar AUCs for SVM (0.896) and CNN (0.876). For prediction in MCI, performances decreased for both SVM (0.665) and CNN (0.702). Both with SVM and CNN, classification based on modulated GM maps significantly outperformed classification based on minimally processed images. Deep and conventional classifiers performed equally well for AD classification and their performance decreased only slightly when applied to the external cohort. We expect that this work on external validation contributes towards translation of machine learning to clinical practice.
Learning unbiased registration and joint segmentation: evaluation on longitudinal diffusion MRI
Nov 03, 2020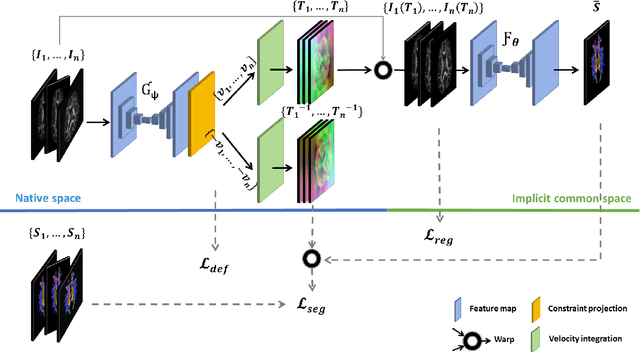
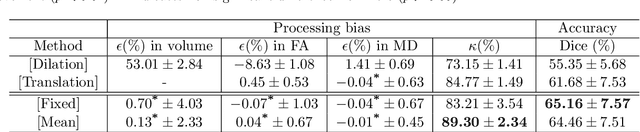
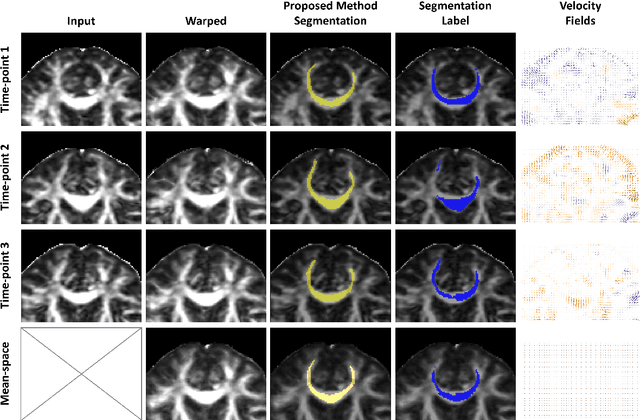
Abstract:Analysis of longitudinal changes in imaging studies often involves both segmentation of structures of interest and registration of multiple timeframes. The accuracy of such analysis could benefit from a tailored framework that jointly optimizes both tasks to fully exploit the information available in the longitudinal data. Most learning-based registration algorithms, including joint optimization approaches, currently suffer from bias due to selection of a fixed reference frame and only support pairwise transformations. We here propose an analytical framework based on an unbiased learning strategy for group-wise registration that simultaneously registers images to the mean space of a group to obtain consistent segmentations. We evaluate the proposed method on longitudinal analysis of a white matter tract in a brain MRI dataset with 2-3 time-points for 3249 individuals, i.e., 8045 images in total. The reproducibility of the method is evaluated on test-retest data from 97 individuals. The results confirm that the implicit reference image is an average of the input image. In addition, the proposed framework leads to consistent segmentations and significantly lower processing bias than that of a pair-wise fixed-reference approach. This processing bias is even smaller than those obtained when translating segmentations by only one voxel, which can be attributed to subtle numerical instabilities and interpolation. Therefore, we postulate that the proposed mean-space learning strategy could be widely applied to learning-based registration tasks. In addition, this group-wise framework introduces a novel way for learning-based longitudinal studies by direct construction of an unbiased within-subject template and allowing reliable and efficient analysis of spatio-temporal imaging biomarkers.
Differential diagnosis and molecular stratification of gastrointestinal stromal tumors on CT images using a radiomics approach
Oct 15, 2020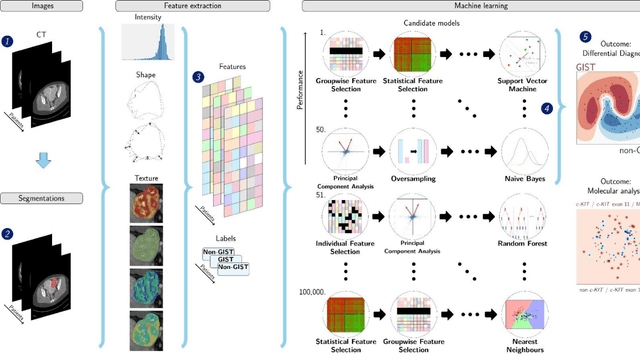
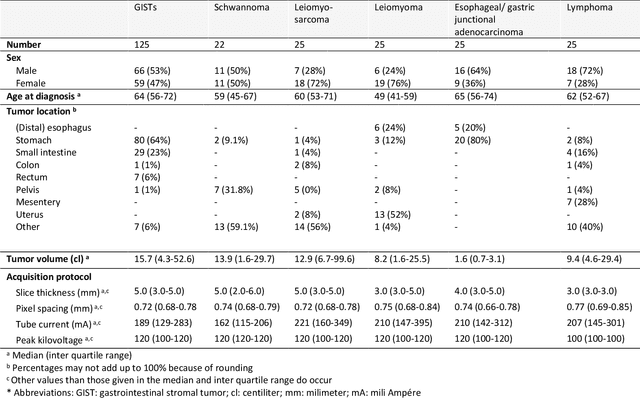
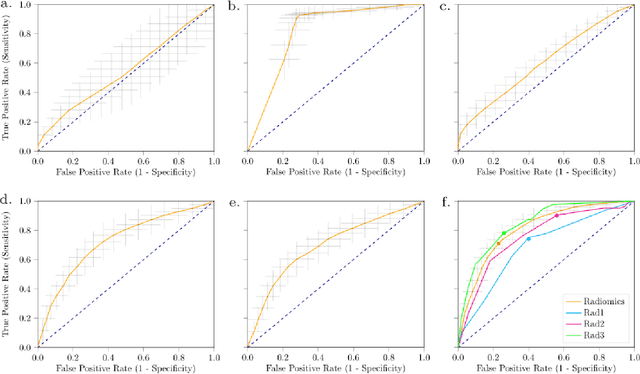
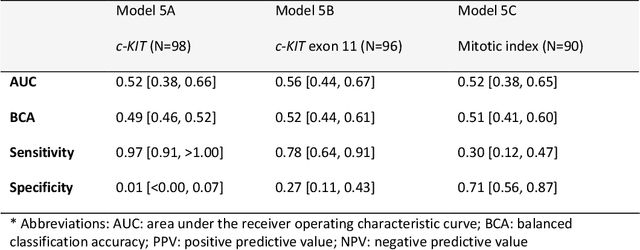
Abstract:Distinguishing gastrointestinal stromal tumors (GISTs) from other intra-abdominal tumors and GISTs molecular analysis is necessary for treatment planning, but challenging due to its rarity. The aim of this study was to evaluate radiomics for distinguishing GISTs from other intra-abdominal tumors, and in GISTs, predict the c-KIT, PDGFRA,BRAF mutational status and mitotic index (MI). All 247 included patients (125 GISTS, 122 non-GISTs) underwent a contrast-enhanced venous phase CT. The GIST vs. non-GIST radiomics model, including imaging, age, sex and location, had a mean area under the curve (AUC) of 0.82. Three radiologists had an AUC of 0.69, 0.76, and 0.84, respectively. The radiomics model had an AUC of 0.52 for c-KIT, 0.56 for c-KIT exon 11, and 0.52 for the MI. Hence, our radiomics model was able to distinguish GIST from non-GISTS with a performance similar to three radiologists, but was not able to predict the c-KIT mutation or MI.
WHO 2016 subtyping and automated segmentation of glioma using multi-task deep learning
Oct 09, 2020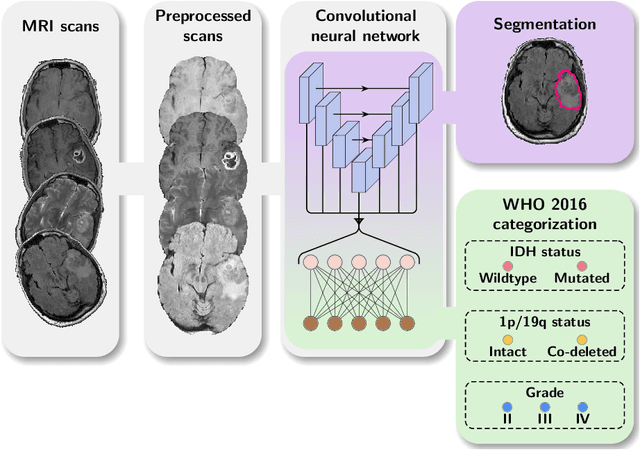
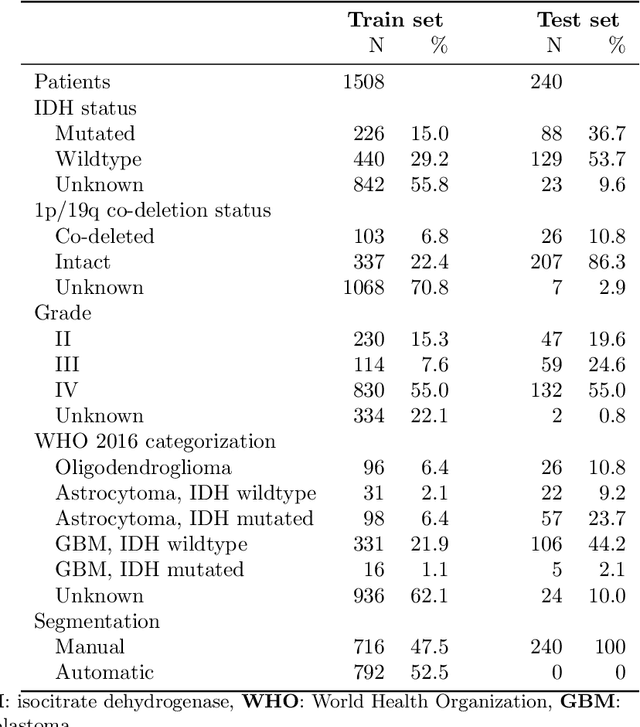
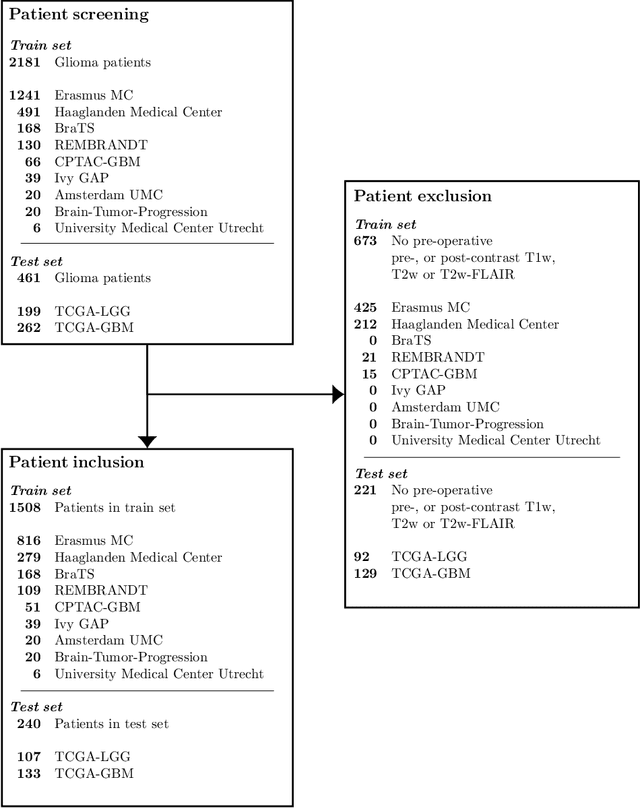
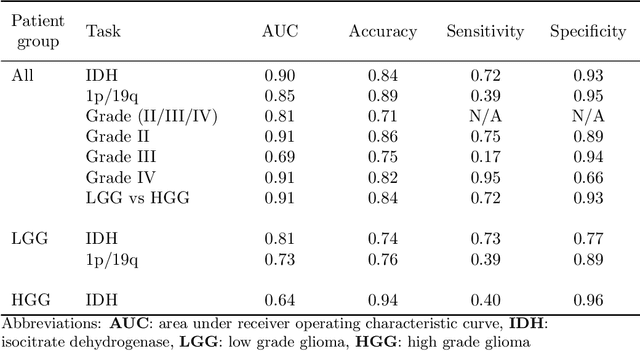
Abstract:Accurate characterization of glioma is crucial for clinical decision making. A delineation of the tumor is also desirable in the initial decision stages but is a time-consuming task. Leveraging the latest GPU capabilities, we developed a single multi-task convolutional neural network that uses the full 3D, structural, pre-operative MRI scans to can predict the IDH mutation status, the 1p/19q co-deletion status, and the grade of a tumor, while simultaneously segmenting the tumor. We trained our method using the largest, most diverse patient cohort to date containing 1508 glioma patients from 16 institutes. We tested our method on an independent dataset of 240 patients from 13 different institutes, and achieved an IDH-AUC of 0.90, 1p/19q-AUC of 0.85, grade-AUC of 0.81, and a mean whole tumor DICE score of 0.84. Thus, our method non-invasively predicts multiple, clinically relevant parameters and generalizes well to the broader clinical population.
Analyzing the effect of APOE on Alzheimer's disease progression using an event-based model for stratified populations
Sep 15, 2020
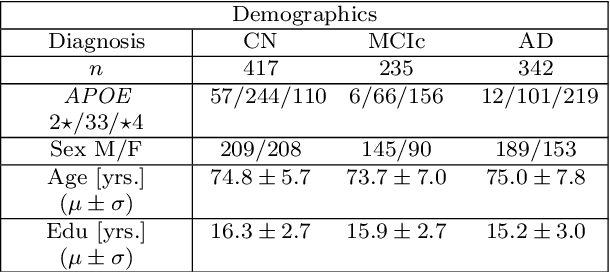
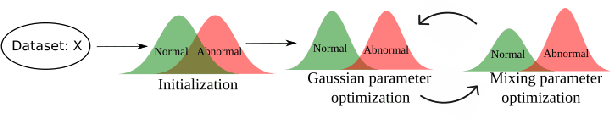
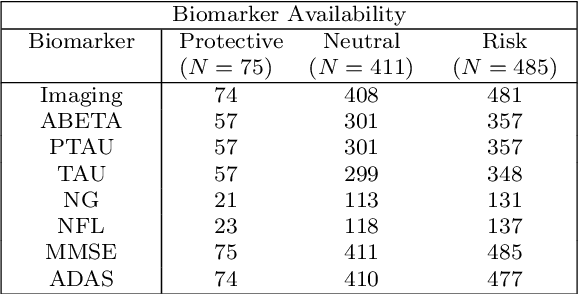
Abstract:Alzheimer's disease (AD) is the most common form of dementia and is phenotypically heterogeneous. APOE is a triallelic gene which correlates with phenotypic heterogeneity in AD. In this work, we determined the effect of APOE alleles on the disease progression timeline of AD using a discriminative event-based model (DEBM). Since DEBM is a data-driven model, stratification into smaller disease subgroups would lead to more inaccurate models as compared to fitting the model on the entire dataset. Hence our secondary aim is to propose and evaluate novel approaches in which we split the different steps of DEBM into group-aspecific and group-specific parts, where the entire dataset is used to train the group-aspecific parts and only the data from a specific group is used to train the group-specific parts of the DEBM. We performed simulation experiments to benchmark the accuracy of the proposed approaches and to select the optimal approach. Subsequently, the chosen approach was applied to the baseline data of 417 cognitively normal, 235 mild cognitively impaired who convert to AD within 3 years, and 342 AD patients from the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset to gain new insights into the effect of APOE carriership on the disease progression timeline of AD. The presented models could aid understanding of the disease, and in selecting homogeneous group of presymptomatic subjects at-risk of developing symptoms for clinical trials.
Towards segmentation and spatial alignment of the human embryonic brain using deep learning for atlas-based registration
May 13, 2020



Abstract:We propose an unsupervised deep learning method for atlas based registration to achieve segmentation and spatial alignment of the embryonic brain in a single framework. Our approach consists of two sequential networks with a specifically designed loss function to address the challenges in 3D first trimester ultrasound. The first part learns the affine transformation and the second part learns the voxelwise nonrigid deformation between the target image and the atlas. We trained this network end-to-end and validated it against a ground truth on synthetic datasets designed to resemble the challenges present in 3D first trimester ultrasound. The method was tested on a dataset of human embryonic ultrasound volumes acquired at 9 weeks gestational age, which showed alignment of the brain in some cases and gave insight in open challenges for the proposed method. We conclude that our method is a promising approach towards fully automated spatial alignment and segmentation of embryonic brains in 3D ultrasound.
Towards continuous learning for glioma segmentation with elastic weight consolidation
Sep 25, 2019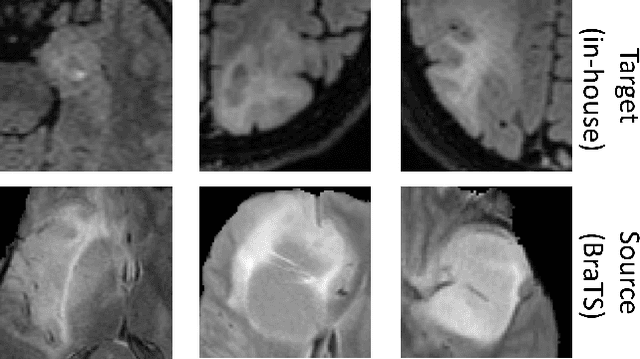
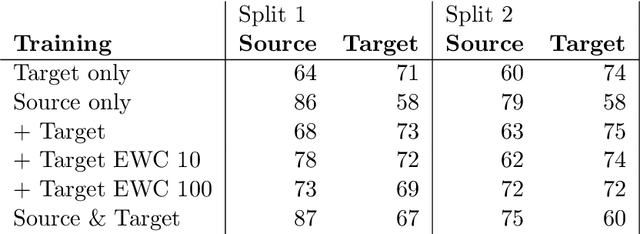
Abstract:When finetuning a convolutional neural network (CNN) on data from a new domain, catastrophic forgetting will reduce performance on the original training data. Elastic Weight Consolidation (EWC) is a recent technique to prevent this, which we evaluated while training and re-training a CNN to segment glioma on two different datasets. The network was trained on the public BraTS dataset and finetuned on an in-house dataset with non-enhancing low-grade glioma. EWC was found to decrease catastrophic forgetting in this case, but was also found to restrict adaptation to the new domain.
Multi-modal segmentation with missing MR sequences using pre-trained fusion networks
Sep 25, 2019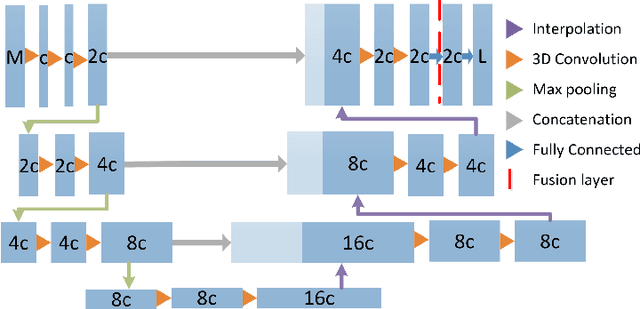
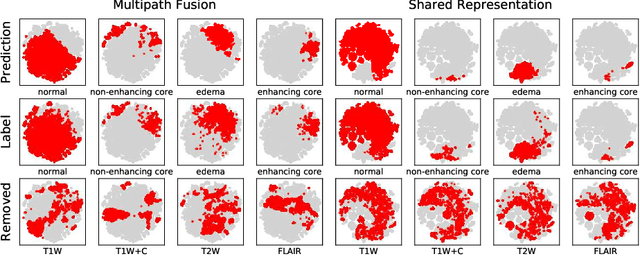
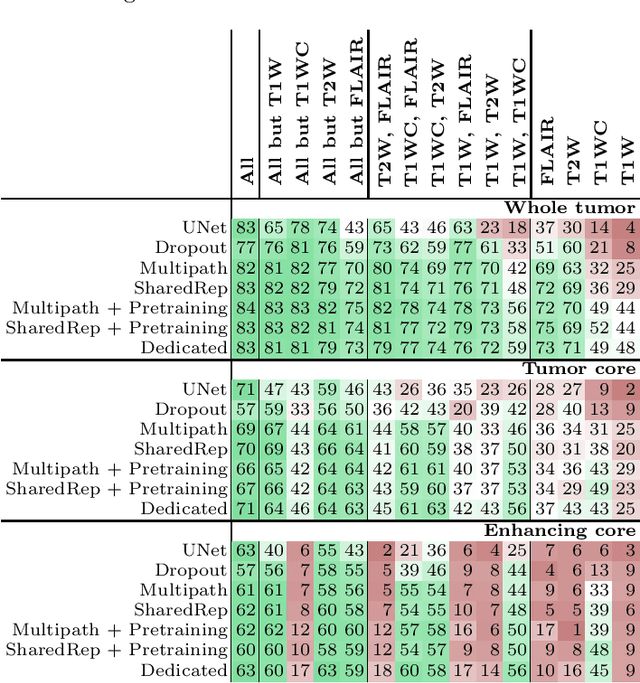

Abstract:Missing data is a common problem in machine learning and in retrospective imaging research it is often encountered in the form of missing imaging modalities. We propose to take into account missing modalities in the design and training of neural networks, to ensure that they are capable of providing the best possible prediction even when multiple images are not available. The proposed network combines three modifications to the standard 3D UNet architecture: a training scheme with dropout of modalities, a multi-pathway architecture with fusion layer in the final stage, and the separate pre-training of these pathways. These modifications are evaluated incrementally in terms of performance on full and missing data, using the BraTS multi-modal segmentation challenge. The final model shows significant improvement with respect to the state of the art on missing data and requires less memory during training.
APIR-Net: Autocalibrated Parallel Imaging Reconstruction using a Neural Network
Sep 19, 2019



Abstract:Deep learning has been successfully demonstrated in MRI reconstruction of accelerated acquisitions. However, its dependence on representative training data limits the application across different contrasts, anatomies, or image sizes. To address this limitation, we propose an unsupervised, auto-calibrated k-space completion method, based on a uniquely designed neural network that reconstructs the full k-space from an undersampled k-space, exploiting the redundancy among the multiple channels in the receive coil in a parallel imaging acquisition. To achieve this, contrary to common convolutional network approaches, the proposed network has a decreasing number of feature maps of constant size. In contrast to conventional parallel imaging methods such as GRAPPA that estimate the prediction kernel from the fully sampled autocalibration signals in a linear way, our method is able to learn nonlinear relations between sampled and unsampled positions in k-space. The proposed method was compared to the start-of-the-art ESPIRiT and RAKI methods in terms of noise amplification and visual image quality in both phantom and in-vivo experiments. The experiments indicate that APIR-Net provides a promising alternative to the conventional parallel imaging methods, and results in improved image quality especially for low SNR acquisitions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge